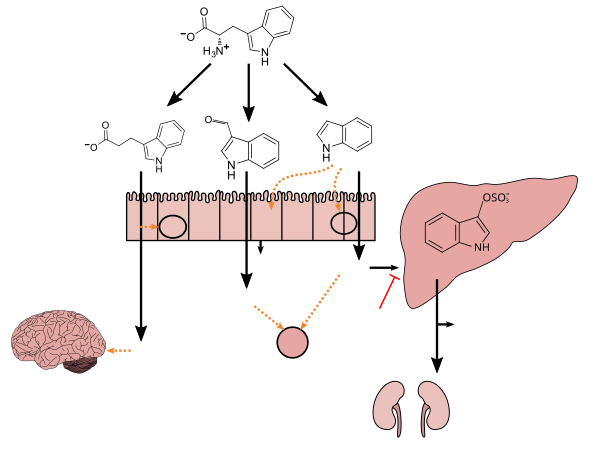
Template:Tryptophan metabolism by human microbiota
Tryptophan metabolism by
human gastrointestinal microbiota ( )activated charcoal), an intestinal sorbent that is taken by mouth, adsorbs indole, in turn decreasing the concentration of indoxyl sulfate in blood plasma.[1] |
To
transclude
this template as is, use:{{Tryptophan metabolism by human microbiota}}To transclude this template with a header or with a different alignment, use:
{{Tryptophan metabolism by human microbiota | align=(left/right/center) | header=(desired header) | header align=(left/right/center) | header background=(color hex code)}}
The image's
alternative text
(i.e. |alt= parameter) is "Tryptophan metabolism diagram".
Reflist
- ^ PMID 27102537.
Lactobacillus spp. convert tryptophan to indole-3-aldehyde (I3A) through unidentified enzymes [125]. Clostridium sporogenes convert tryptophan to IPA [6], likely via a tryptophan deaminase. ... IPA also potently scavenges hydroxyl radicals
Table 2: Microbial metabolites: their synthesis, mechanisms of action, and effects on health and disease
Figure 1: Molecular mechanisms of action of indole and its metabolites on host physiology and disease - PMID 19234110.
Production of IPA was shown to be completely dependent on the presence of gut microflora and could be established by colonization with the bacterium Clostridium sporogenes.
IPA metabolism diagram - ^ "3-Indolepropionic acid". Human Metabolome Database. University of Alberta. Retrieved 12 June 2018.
- S2CID 6630247.
[Indole-3-propionic acid (IPA)] has previously been identified in the plasma and cerebrospinal fluid of humans, but its functions are not known. ... In kinetic competition experiments using free radical-trapping agents, the capacity of IPA to scavenge hydroxyl radicals exceeded that of melatonin, an indoleamine considered to be the most potent naturally occurring scavenger of free radicals. In contrast with other antioxidants, IPA was not converted to reactive intermediates with pro-oxidant activity.