L-ribulose-5-phosphate 4-epimerase
| L-ribulose-phosphate 4-epimerase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
In
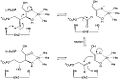
enzymology, a L-ribulose-5-phosphate 4-epimerase (EC 5.1.3.4) is an enzyme that catalyzes the interconversion of ribulose 5-phosphate and xylulose 5-phosphate in the oxidative phase of the Pentose phosphate pathway.[1]
- L-ribulose 5-phosphate D-xylulose 5-phosphate
This enzyme has a molecular mass of 102
pentose and glucuronate interconversions and ascorbate and aldarate metabolism
.
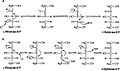
Enzyme Mechanism
-
Mechanism of ribulose 5-phosphate 4-epimerase in active site
-
Aldol and dehydration mechanisms
L-Ribulose 5-phosphate 4-epimerase catalyzes the epimerization of L-
hydroxyl group on C-4, followed by cleavage of the bond between C-3 and C-4 to give a metal-stabilized acetone enediolate and a glycolaldehyde phosphate fragment. The C–C bond of glycolaldehyde phosphate is then rotated 180°, and the C–C bond between C-3 and C-4 is regenerated to give inversion of stereochemistry at C-4.[3]
This mechanism is contested by a possible alternative
rate-limiting step in this mechanism and substituting the C-3 hydrogen with deuterium would significantly alter the rate. At the same time there are significantly large 13C isotope effects, suggesting rate-limiting C–C bond breakage, as expected with the aldol mechanism.[3]
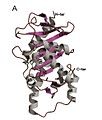
Structure
-
Homo-tetrameric structure of L-Ru5P
-
L-Ru5P monomer
-
L-Ru5p active site
The structure is homo-tetrameric and displays C4 symmetry.aldolases that catalyze carbon-carbon bond cleavage reactions via a metal-stabilized enolate intermediate.
Biological Function
Ribulose 5-phosphate 4-epimerase is found on the well studied
intracellular arabinose in three steps catalyzed by the products of the araB, araA, araD genes to D-xylulose-5-phosphate.[5]
| Gene | Protein |
|---|---|
| AraA | Isomerase |
| AraB | Ribulokinase |
| AraC | Regulatory |
| AraD | Epimerase |
| AraE | Uptake |
| AraF | Uptake |
| AraG | Uptake |
| AraH | Uptake |
Evolution
L-Ribulose-5-phosphate 4-epimerase and L-fuculose-1-phosphate (L-Fuc1P) aldolase are evolutionarily related enzymes that display 26% sequence identity and a very high degree of structural similarity.substrates differing greatly in the position of the phosphate (C-5 vs C-1).[6]
References
Further reading
- BURMA DP, HORECKER BL (1958). "Pentose Fermentation by Lactobacillus Plantarum: IV. L-Ribulose-5-phosphate 4-Epimerase". J. Biol. Chem. 231 (2): 1053–64. PMID 13539036.
- Deupree JD; Wood WA (1970). "L-Ribulose 5-phosphate 4-epimerase of Aerobacter aerogenes. Evidence for nicotinamide adenine dinucleotide-independent 4-epimerization by the crystalline enzyme". J. Biol. Chem. 245 (15): 3988–95. PMID 4395381.
- Lee N, Patrick JW, Masson M (1968). "Crystalline L-ribulose 5-phosphate 4-epimerase from Escherichia coli". J. Biol. Chem. 243 (18): 4700–5. PMID 4879898.
- WOLIN MJ, SIMPSON FJ, WOOD WA (1958). "Degradation of L-arabinose by Aerobacter aerogenes. III Identification and properties of L-ribulose-5-phosphate 4-epimerase". J. Biol. Chem. 232 (1): 559–75. PMID 13549442.