Beta hairpin

The beta hairpin (sometimes also called beta-ribbon or beta-beta unit) is a simple
Researchers such as
Classification
Beta hairpins were originally categorized solely by the number of amino acid residues in their loop sequences, such that they were named one-residue, two-residue, etc.[2] This system, however, is somewhat ambiguous as it does not take into account whether the residues that signal the end of the hairpin are singly or doubly hydrogen bonded to one another. An improved means of classification has since been proposed by Milner-White and Poet.[3] Beta hairpins are broken into four distinct classes as depicted in the publication's Figure 1. Each class begins with the smallest possible number of loop residues and progressively increases the loop size by removing hydrogen bonds in the beta sheet. The primary hairpin of class 1 is a one-residue loop where the bound residues share two hydrogen bonds. One hydrogen bond is then removed to create a three-residue loop, which is the secondary hairpin of class 1. Singly bound residues are counted in the loop sequence but also signal the end of the loop, thus defining this hairpin as a three-residue loop. This single hydrogen bond is then removed to create the tertiary hairpin; a five-residue loop with doubly bound residues. This pattern continues indefinitely and defines all beta hairpins within the class. Class 2 follows the same pattern beginning with a two-residue loop with terminating residues that share two hydrogen bonds. Class 3 begins with a three-residue, and class 4 with a four-residue. Class 5 does not exist as that primary hairpin is already defined in class 1. Pi This classification scheme not only accounts for various degrees of hydrogen bonding, but also says something about the biological behavior of the hairpin. Single amino acid replacements may destroy a particular hydrogen bond, but will not unfold the hairpin or change its class. On the other hand, amino acid insertions and deletions will have to unfold and reform the entire
Folding and binding dynamics

Understanding the mechanism through which micro-domains fold can help to shed light onto the folding patterns of whole
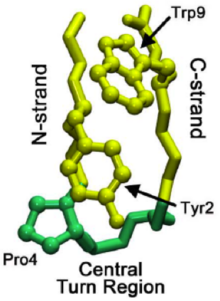
In the folding of overall proteins, the turn may originate not in the native turn region but in the C-strand of the beta-hairpin. This turn then propagates through the C-strand (the beta strand leading to C-terminus) until it reaches the native turn region. Sometimes the residue interactions leading up to the native turn region are too strong, causing reverse propagation. However, once the native turn does form, interactions between prolines and tryptophan residues (seen in image at right) in the region help to stabilize the turn, preventing "roll back" or dissolution.
Researchers believe that turns do not originate in the N-strand, due to increased rigidity (often caused by a proline leading up to the native turn region) and less conformational options. The initial turn formation takes place in about 1 μs. Once the initial turn has been established, two mechanisms have been proposed as to how the rest of the beta-hairpin folds: a hydrophobic collapse with side-chain level rearrangements, or the more accepted zipper-like mechanism.[4]
The β-hairpin loop motif can be found in many macromolecular proteins. However, small and simple β-hairpins can exist on their own as well. To see this clearly, the
Proteins that are β-sheet rich, also called
This enzyme binds its ligand through
It is also common to find proline residues within the actual loop portion of the β-hairpin, since this amino acid is rigid and contributes to the "turn" formation. These proline residues can be seen as red side chains in the image of the Pin1 WW domain below (left).
 |
 |
Artificially designed beta-hairpin
The design of peptides that adopt β-hairpin structure (without relying on metal binding, unusual amino acids, or disulfide crosslinks) has made significant progress and yielded insights into protein dynamics. Unlike

The synthesis of trpzip β-hairpin peptides has incorporated photoswitches that facilitate precise control over folding. Several amino acids in the turn are replaced by azobenzene, which can be induced to switch from the trans to the cis conformation by light at 360 nm. When the azobenzene moiety is in the cis conformation, the amino acid residues align correctly to adopt a β-hairpin formation. However, the trans conformation does not have proper turn geometry for the β-hairpin.[8] This phenomenon can be used to investigate peptide conformational dynamics with femtosecond absorption spectroscopy.[8]
References
- S2CID 35065527.
- ^ Sibanda, B.L.; Blundell, T.L.; Thorton, J.M. (1985). "Conformations of Beta-Hairpins in Protein Structures". Nature(London) 316 170–174.
- ^ a b Milner-White, J.; Poet, R. (1986). "Four Classes of Beta-Hairpins in Proteins". Biochemical Journal 240 289–292.
- ^ PMID 22970341.
- PMID 18844292.
- ^ Kay, B.K.; Williamson, M.P.; Sudol, M. The Importance of Being Proline: the interaction of proline-rich motifs in signaling proteins with their cognate domains. The FASEB Journal. 2000, 14, 231–241.
- PMID 11331745.
- ^ PMID 16294349.
