Exoenzyme

An exoenzyme, or extracellular enzyme, is an
History
Very limited information is available about the original discovery of exoenzymes. According to Merriam-Webster dictionary, the term "exoenzyme" was first recognized in the English language in 1908.[6] The book "Intracellular Enzymes: A Course of Lectures Given in the Physiological," by Horace Vernon is thought to be the first publication using this word in that year.[7] Based on the book, it can be assumed that the first known exoenzymes were pepsin and trypsin, as both are mentioned by Vernon to have been discovered by scientists Briike and Kiihne before 1908.[8]
Function
In
Many exoenzymes are also used as
In
Examples of exoenzymes as virulence factors
Source:[3]

Necrotizing enzymes
Coagulase
By binding to

Kinases
The opposite of coagulase, kinases can dissolve clots. S. aureus can also produce staphylokinase, allowing them to dissolve the clots they form, to rapidly diffuse into the host at the correct time.[14]
Hyaluronidase
Similar to collagenase, hyaluronidase enables a pathogen to penetrate deep into tissues. Bacteria such as Clostridium do so by using the enzyme to dissolve collagen and hyaluronic acid, the protein and saccharides, respectively, that hold tissues together.
Hemolysins
Examples of digestive exoenzymes
Amylases
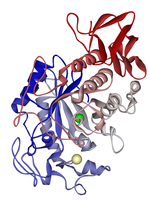
Lipoprotein lipase
Pectinase
the pectic elements found in plants and plant-derived products.Pepsin
Discovered in 1836,
Trypsin
Also one of the first exoenzymes to be discovered,
Bacterial assays
The production of a particular digestive exoenzyme by a bacterial cell can be assessed using plate assays. Bacteria are streaked across the agar, and are left to incubate. The release of the enzyme into the surroundings of the cell cause the breakdown of the macromolecule on the plate. If a reaction does not occur, this means that the bacteria does not create an exoenzyme capable of interacting with the surroundings. If a reaction does occur, it becomes clear that the bacteria does possess an exoenzyme, and which macromolecule is hydrolyzed determines its identity.[2]
Amylase
Amylase breaks down carbohydrates into mono- and disaccharides, so a starch agar must be used for this assay. Once the bacteria is streaked on the agar, the plate is flooded with iodine. Since iodine binds to starch but not its digested by-products, a clear area will appear where the amylase reaction has occurred. Bacillus subtilis is a bacterium that results in a positive assay as shown in the picture.[2]
Lipase
Lipase assays are done using a
Biotechnological and industrial applications

Bioremediation applications


References
- PMID 18577009.
- ^ a b c d Roberts, K. "Exoenzymes". Prince George's Community College. Archived from the original on 13 June 2013. Retrieved 8 December 2013.
- ^ ISBN 9781605476735.)
{{cite book}}: CS1 maint: multiple names: authors list (link - ^ ISBN 9781402092121.)
{{cite book}}:|first=has generic name (help)CS1 maint: multiple names: authors list (link - PMID 21329211.
- ^ "Merriam-Webster". Retrieved 2013-10-26.
- ^ "Lexic.us". Retrieved 2013-10-26.
- ^ a b Vernon, Horace. "Intracellular Enzymes: A Course of Lectures Given in the Physiological". Retrieved 2013-10-26.
- ^ Kaiser, Gary. "Lab 8: Identification of Bacteria Through Biochemical Testing". Biol 230 Lab Manual. Archived from the original on 11 December 2013. Retrieved 9 December 2013.
- PMID 20926516.
- PMID 10377131.
- ISBN 978-0716776017.
- ^ Andrews, Lary. "Supplemental Enzymes for Digestion". Health and Healing Research. Archived from the original on 27 July 2013. Retrieved 9 December 2013.
- ^ Todar, Kenneth. "Mechanisms of Bacterial Pathogenicity". Todar's Online Textbook of Bacteriology. Kenneth Todar, PhD. Retrieved 12 December 2013.
- S2CID 253807898.
- .
- PMID 10744959.
- PMID 21634067. Retrieved 25 November 2013.
- ^ S2CID 40089672.
- PMID 2677048.
- .
- .
- .
- ^ a b c "Encyclopædia Britannica". Retrieved November 14, 2013.
- PMID 6785873.
- ^ a b Worthington, Krystal. "Trypsin". Worthington Biochemical Corporation. Retrieved 26 November 2013.
- ^ "Trypsin". Free Dictionary. Retrieved 26 November 2013.
- ^ "Trypsin Product Information". Worthington Biochemical Corporation. Retrieved 26 November 2013.
- ^ S2CID 7785046.
- PMID 22138412.
- ^ PMID 12323363.
- PMID 22426735.
- ^ PMID 14538100.
- ^ "A Citizen's Guide to Bioremediation". United States Environmental Protection Agency. September 2012. Retrieved 5 December 2013.
- PMID 21912739.
- ^ S2CID 24676340.
- ^ .
- S2CID 253773148.


