Biomolecular structure
This article may be confusing or unclear to readers. (February 2016) |
 |
 |
Biomolecular structure is the intricate folded, three-dimensional shape that is formed by a
Primary structure
The primary structure of a
The
Secondary structure
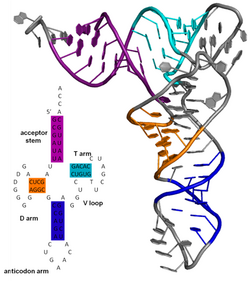
The secondary structure of a protein is the pattern of hydrogen bonds in a biopolymer. These determine the general three-dimensional form of local segments of the biopolymers, but does not describe the global structure of specific atomic positions in three-dimensional space, which are considered to be tertiary structure. Secondary structure is formally defined by the hydrogen bonds of the biopolymer, as observed in an atomic-resolution structure. In proteins, the secondary structure is defined by patterns of hydrogen bonds between backbone amine and carboxyl groups (sidechain–mainchain and sidechain–sidechain hydrogen bonds are irrelevant), where the DSSP definition of a hydrogen bond is used.
The secondary structure of a nucleic acid is defined by the hydrogen bonding between the nitrogenous bases.
For proteins, however, the hydrogen bonding is correlated with other structural features, which has given rise to less formal definitions of secondary structure. For example, helices can adopt backbone dihedral angles in some regions of the Ramachandran plot; thus, a segment of residues with such dihedral angles is often called a helix, regardless of whether it has the correct hydrogen bonds. Many other less formal definitions have been proposed, often applying concepts from the differential geometry of curves, such as curvature and torsion. Structural biologists solving a new atomic-resolution structure will sometimes assign its secondary structure by eye and record their assignments in the corresponding Protein Data Bank (PDB) file.
The
Tertiary structure
The
Quaternary structure
The protein quaternary structure [a] refers to the number and arrangement of multiple protein molecules in a multi-subunit complex.
For nucleic acids, the term is less common, but can refer to the higher-level organization of DNA in chromatin,[7] including its interactions with histones, or to the interactions between separate RNA units in the ribosome[8][9] or spliceosome.
Structure determination
Structure probing is the process by which biochemical techniques are used to determine biomolecular structure.[10] This analysis can be used to define the patterns that can be used to infer the molecular structure, experimental analysis of molecular structure and function, and further understanding on development of smaller molecules for further biological research.[11] Structure probing analysis can be done through many different methods, which include chemical probing, hydroxyl radical probing, nucleotide analog interference mapping (NAIM), and in-line probing.[10]
and the structure is not tractable using only the standard analysis.In contrast, the standard analysis, involving only Fourier transforms of Bessel functions[19] and DNA molecular models, is still routinely used to analyze A-DNA and Z-DNA X-ray diffraction patterns.[20]
Structure prediction

Biomolecular structure prediction is the prediction of the three-dimensional structure of a protein from its amino acid sequence, or of a nucleic acid from its nucleobase (base) sequence. In other words, it is the prediction of secondary and tertiary structure from its primary structure. Structure prediction is the inverse of biomolecular design, as in rational design, protein design, nucleic acid design, and biomolecular engineering.
Protein structure prediction is one of the most important goals pursued by bioinformatics and theoretical chemistry. Protein structure prediction is of high importance in medicine (for example, in drug design) and biotechnology (for example, in the design of novel enzymes). Every two years, the performance of current methods is assessed in the Critical Assessment of protein Structure Prediction (CASP) experiment.
There has also been a significant amount of
Secondary structure of small nucleic acid molecules is determined largely by strong, local interactions such as
Sequence covariation methods rely on the existence of a data set composed of multiple homologous RNA sequences with related but dissimilar sequences. These methods analyze the covariation of individual base sites in evolution; maintenance at two widely separated sites of a pair of base-pairing nucleotides indicates the presence of a structurally required hydrogen bond between those positions. The general problem of pseudoknot prediction has been shown to be NP-complete.[23]
Design
Biomolecular design can be considered the inverse of structure prediction. In structure prediction, the structure is determined from a known sequence, whereas, in protein or nucleic acid design, a sequence that will form a desired structure is generated.
Other biomolecules
This section needs expansion. You can help by adding to it. (April 2010) |
Other biomolecules, such as polysaccharides, polyphenols and lipids, can also have higher-order structure of biological consequence.
See also
- Biomolecular
- Comparison of nucleic acid simulation software
- Gene structure
- List of RNA structure prediction software
- Non-coding RNA
Notes
- distributive numbers, and follows binary and ternary; while quartary is derived from Latin ordinal numbers, and follows secondary and tertiary. However, quaternary is standard in biology.
References
- PMID 9649444.
- PMID 9106664.
- S2CID 4162567.
- PMID 3313277.
- S2CID 9982829.
- S2CID 35930758.
- PMID 6206780.
- PMID 11296253.
- ^ ISBN 978-90-901323-4-1.
- ISBN 978-0-87969-589-7.
- .
- S2CID 4268222.
- S2CID 4280080.
- PMID 7441761.
- S2CID 189888972.
- ^ Hosemann R, Bagchi RN (1962). Direct analysis of diffraction by matter. Amsterdam/New York: North-Holland.
- .
- ^ "Bessel functions and diffraction by helical structures". planetphysics.org.[permanent dead link]
- ^ "X-Ray Diffraction Patterns of Double-Helical Deoxyribonucleic Acid (DNA) Crystals". planetphysics.org. Archived from the original on 24 July 2009.
- ^ PMID 16500677.
- S2CID 189885784.
- PMID 11108471.

