DNA base flipping
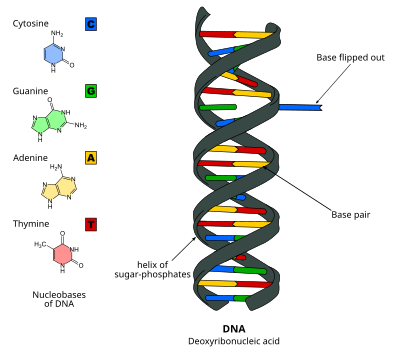
DNA base flipping, or nucleotide flipping, is a mechanism in which a single
DNA base flipping occurs by breaking the
Discovery
Base flipping was first observed in 1994 when researchers Klimasauskas, Kumar, Roberts, and Cheng used X-ray crystallography to view an intermediate step in the chemical reaction of a methyltransferase bound to DNA.[3] The methyltransferase they used was the C5-cytosine methyltransferase from Haemophilus haemolyticus (M. HhaI). This enzyme recognizes a specific sequence of the DNA (5'-GCGC-3') and methylates the first cytosine base of the sequence at its C5 location.[3] Upon crystallization of the M. HhaI-DNA complex, they saw the target cytosine base was rotated completely out of the double helix and was positioned in the active site of the M. HhaI. It was held in place by numerous interactions between the M. HhaI and DNA.[3]
The authors theorized that base flipping was a mechanism used by many other enzymes, such as
Mechanism
There are two mechanisms of DNA base flipping: active and passive.[13] In the active mechanism, an enzyme binds to the DNA and then actively rotates the base, while in the passive mechanism a damaged base rotates out spontaneously first, then is recognized and bound by the enzyme.[8] Research has demonstrated both mechanisms: uracil-DNA glycosylase follows the passive mechanism[8] and Tn10 transposase follows the active mechanism.[14]
Furthermore, studies have shown that DNA base flipping is used by many different enzymes in a variety biological processes such as
Biological processes
DNA modification and repair

DNA can have
Replication, transcription and recombination
Base flipping occurs during latter stages of recombination.[21] RecA is a protein that promotes strand invasion[15] during homologous recombination. Base flipping has been proposed as the mechanism by which RecA can enable a single strand to recognize homology in duplex DNA.[22] Other studies indicate that it is also involved in V(D)J Recombination.[23]
DNA methylation

DNA methylation is the process in which a methyl group is added to either a cytosine or adenine.[24] This process causes the activation or inactivation of gene expression, thereby resulting in gene regulation in eukaryotic cells. DNA methylation process is also known to be involved in certain types of cancer formation.[25][26][27] In order for this chemical modification to occur, it is necessary that the target base flips out of the DNA double helix to allow the methyltransferases to catalyze the reaction.[5]
Target recognition by restriction endonucleases
Restriction endonucleases, also known as
Experimental approaches for detection
X-ray crystallography

- Purification
- Crystallization
- Data Collection
- Structure determination and refinement
During purification, Haemophilus haemolyticus
NMR spectroscopy
Fluorescence spectroscopy
Fluorescence spectroscopy is a technique that is used to assay a sample using a fluorescent probe. DNA nucleotides themselves are not good candidates for this technique because they do not readily re-emit light upon light excitation.[39] A fluorescent marker is needed to detect base flipping. 2-Aminopurine is a base that is structurally similar to adenine, but is very fluorescent when flipped out from the DNA duplex.[40] It is commonly used to detect base flipping and has an excitation at 305‑320 nm and emission at 370 nm so that it well separated from the excitations of proteins and DNA. Other fluorescent probes used to study DNA base flipping are 6MAP (4‑amino‑6‑methyl‑7(8H)‑pteridone)[41] and Pyrrolo‑C (3-[β-D-2-ribofuranosyl]-6-methylpyrrolo[2,3-d]pyrimidin-2(3H)-one).[42][43] Time-resolved fluorescence spectroscopy is also employed to provide a more detailed picture of the extent of base flipping as well as the conformational dynamics occurring during base flipping.[44]
Hybridization probing
Hybridization probes can be used to detect base flipping. This technique uses a molecule that has a complementary sequence to the sequence you would like to detect such that it binds to a single-strand of the DNA or RNA. Several hybridization probes have been used to detect base flipping. Potassium permanganate is used to detect thymine residues that have been flipped out by cytosine-C5 and adenine-N6 methyltransferases.[45] Chloroacetaldehyde is used to detect cytosine residues flipped out by the HhaI DNA cytosine-5 methyltransferase (M. HhaI).[46]

See also
- DNA repair
- Base excision repair
- DNA replication
- RNA transcription
- DNA methylation
- DNA methyltransferase
- Genetic recombination
- Homologous recombination
- DNA
- Epigenetics
- Epigenomics
References
- PMID 9759487.
- S2CID 25391616.
- ^ S2CID 23161543.
- ^ a b Brown, Tom. "Nucleic Acids Book". ATDBio. Retrieved 26 February 2014.
- ^ PMID 12506195.
- ^ a b Grubmüller, Helmut. "DNA Base Flipping". Archived from the original on 4 February 2017. Retrieved 26 February 2014.
- PMID 17496048. Archived from the original(PDF) on 2017-08-09. Retrieved 2014-03-15.
- ^ PMID 15178685.
- ISBN 978-1-58706-329-9. Archived from the original on 2014-04-07. Retrieved 2014-03-10.)
{{cite book}}:|first=has generic name (help - ^ PMID 12506195.
- PMID 12595551.
- PMID 12506195.
- ]
- PMID 19593448.
- ^ ISBN 978-0-321-76243-6.)
{{cite book}}: CS1 maint: multiple names: authors list (link - PMID 12483510.
- S2CID 4426014.
- PMID 12417196.
- S2CID 16277925.
- PMID 11734629.
- PMID 15383274.
- PMID 15383285.
- PMID 19720743.
- PMID 16403636.
- PMID 11707319.
- PMID 11896451.
- S2CID 35380651.
- ^ "Biology and Activity of Restriction Endonucleases". Archived from the original on 2014-04-18. Retrieved 2014-04-03.
- PMID 16473850.
- ^ X-ray crystallography
- PMID 1390649.
- ^ S2CID 54238106.
- ^ Brunger A.T. (1992)"X-PLOR, Version 3.1 : A system for x-ray crystallography and NMR"(New Haven, Connecticut: Yale University Press)
- .
- NMR spectroscopy
- ^ Gueron, M., and J. L. Leroy. 1995. Studies of basepair kinetics by NMR measurement of proton exchange. In Nuclear Magnetic Resonance And Nucleic Acids. Academic Press, San Diego, CA.
- PMID 1331987.
- ^ Klimasaukas, Salius, and Zita Liutkeviciute. "Experimental Approaches to Study DNA Base Flipping." DNA and RNA Modification Enzymes: Structure, Mechanism, Function and Evolution. Landes Bioscience, 2009. 37-50. Web. 16 Mar. 2014. <https://www.landesbioscience.com/pdf/04GrosjeanKlimasauskas.pdf Archived 2014-04-07 at the Wayback Machine>.
- )
- PMID 9461471.
- S2CID 4998287.
- S2CID 44506405.
- .
- PMID 19740769.
- PMID 9671807.
- PMID 18450817.
