Uracil-DNA glycosylase
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 12: 109.1 – 109.11 Mb | Chr 5: 114.27 – 114.28 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Uracil-DNA glycosylase (also known as UNG or UDG) is an enzyme. Its most important function is to prevent mutagenesis by eliminating uracil from DNA molecules by cleaving the N-glycosidic bond and initiating the base-excision repair (BER) pathway.
Function
The human gene encodes one of several uracil-DNA glycosylases. Alternative promoter usage and splicing of this gene leads to two different isoforms: the mitochondrial UNG1 and the nuclear UNG2.
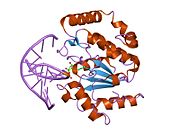
Structure
UDG is made of a four-stranded parallel
- Water-activating loop: 63-QDPYH-67[12]
- Pro-rich loop: 165-PPPPS-169[10]
- Uracil-binding motif: 199-GVLLLN-204[10][11]
- Gly-Ser loop: 246-GS-247[10]
- Minor groove intercalation loop: 268-HPSPLS-273[10][11]
Mechanism
Glycosidic bond cleavage follows a “pinch-push-pull” mechanism using the five conserved motifs.[10]
Pinch: UDG scans DNA for uracil by nonspecifically binding to the strand and creating a kink in the backbone, thereby positioning the selected base for detection. The Pro-rich and Gly-Ser loops form polar contacts with the 3’ and 5’ phosphates flanking the examined base.[11] This compression of the DNA backbone, or “pinch,” allows for close contact between UDG and base of interest.[10]
Push: To fully assess the nucleotide identity, the intercalation loop penetrates, or pushes into, the DNA minor groove and induces a conformational change to flip the nucleotide out of the helix.[13] Backbone compression favors eversion of the now extrahelical nucleotide, which is positioned for recognition by the uracil-binding motif.[10] The coupling of intercalation and eversion helps compensate for the disruption of favorable base stacking interactions within the DNA helix. Leu272 fills the void left by the flipped nucleotide to create dispersion interactions with neighboring bases and restore stacking stability.[11]
Pull: Now accessible to the active site, the nucleotide interacts with the uracil binding motif. The active site shape complements the everted uracil structure, allowing for high substrate specificity.



The position of the residues that activate the water nucleophile and protonate the uracil leaving group are widely debated, though the most commonly followed mechanism employs the water activating loop detailed in the enzyme structure.[12][14] Regardless of position, the identities of the aspartic acid and histidine residues are consistent across catalytic studies.[10][11][12][14][15]
Laboratory use
Uracil N-glycosylase (UNG) is utilized to eliminate carryover polymerase chain reaction (PCR) products in PCR. This method modifies PCR products such that in a new reaction, any residual products from previous PCR amplifications will be digested and prevented from amplifying, but the true DNA templates will be unaffected.[16] PCR synthesizes abundant amplification products each round, but contamination of further rounds of PCR with trace amounts of these products, called carry-over contamination, yields false positive results. Carry-over contamination from some previous PCR can be a significant problem, due both to the abundance of PCR products, and to the ideal structure of the contaminant material for re-amplification. However carry-over contamination can be controlled by the following two steps: (i) incorporating dUTP in all PCR products (by substituting dUTP for dTTP, or by incorporating uracil during synthesis of primers; and (ii) treating all subsequent fully preassembled starting reactions with uracil DNA glycosylase (UDG), followed by thermal inactivation of UDG. UDG cleaves the uracil base from the phosphodiester backbone of uracil-containing DNA, but has no effect on natural (i.e., thymine-containing) DNA. The resulting apyrimidinic sites block replication by DNA polymerases, and are very labile to acid/base hydrolysis. Because UDG does not react with dTTP, and is also inactivated by heat denaturation prior to the actual PCR, carry-over contamination of PCRs can be controlled effectively if the contaminants contain uracils in place of thymines.[6]
Uracil N-glycosylase was also used in a study to detect evidence of ongoing low-level metabolic activity and
Uracil N glycosylase has also been used in a method for cloning of PCR amplified DNA fragments. In this method the primers used in PCR are synthesized with uracil residues instead of thymine. When these primers are incorporated into PCR amplified fragments the primer sequence becomes susceptible to digestion with Uracil N Glycosylase and produce 3' protruding ends that can be annealed to an appropriately prepared vector DNA. The resulting chimeric molecules can be transformed into competent cells with high efficiency, without the need for in vitro ligation.[18]
Interactions
Uracil-DNA glycosylase has been shown to
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000076248 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000029591 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: UNG uracil-DNA glycosylase". Ncbi.nlm.nih.gov. Retrieved 29 December 2017.
- ^ PMID 2227421.
- ^ PMID 10946227.
- S2CID 4310250.
- ^ PMID 10767624.
- ^ PMID 10946228.
- ^ PMID 19909758.
- ^ PMID 12855717.
- S2CID 14851787.
- ^ PMID 17605817.
- S2CID 4315434.
- ^ "Support Centers - Thermo Fisher Scientific". Abcommunity.thermofisher.com. Retrieved 29 December 2017.
- ^ PMID 17728401.
- ^ .Analytical biochemistry 1992 ;206(1):91-7.
- PMID 9045683.