DNA methyltransferase
| N-6 DNA Methylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
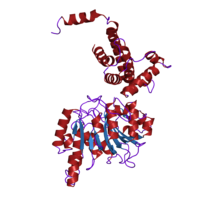 crystal structure of type i restriction enzyme ecoki m protein (ec 2.1.1.72) (m.ecoki) | |||||||||
| Identifiers | |||||||||
| Symbol | N6_Mtase | ||||||||
| Pfam | PF02384 | ||||||||
| Pfam clan | CL0063 | ||||||||
| InterPro | IPR003356 | ||||||||
| PROSITE | PDOC00087 | ||||||||
| |||||||||
| HsdM N-terminal domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | HsdM_N | ||||||||
| Pfam | PF12161 | ||||||||
| |||||||||
| C-5 cytosine-specific DNA methylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| CDD | cd00315 | ||||||||
| |||||||||
| DNA methylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
In
Classification
Substrate
MTases can be divided into three different groups on the basis of the chemical reactions they catalyze:
- m6A - those that generate
- m4C - those that generate
- m5C - those that generate
m6A and m4C methyltransferases are found primarily in prokaryotes (although recent evidence has suggested that m6A is abundant in eukaryotes
The m6A methyltransferases (N-6 adenine-specific DNA methylase) (A-Mtase) are
Among the m6A methyltransferases (N-6 adenine-specific DNA methylase) there is a group of orphan MTases that do not participate in the bacterial restriction/methylation system.
m4C methyltransferases (N-4 cytosine-specific DNA methylases) are
m5C methyltransferases (C-5 cytosine-specific DNA methylase) (C5 Mtase) are enzymes that specifically methylate the C-5
Highly conserved DNA methyltransferases of the m4C, m5C, and m6A types have been reported,[17] which appear as promising targets for the development of novel epigenetic inhibitors to fight bacterial virulence, antibiotic resistance, among other biomedical applications.
De novo vs. maintenance
De novo methyltransferases recognize something in the DNA that allows them to newly methylate cytosines. These are expressed mainly in early embryo development and they set up the pattern of methylation. De novo methyltransferases are also active when a signal-responsive cell, such as a neuron, needs to alter protein expression.[18] As an example, when fear conditioning creates a new memory in a rat, 9.17% of the genes in the rat hippocampus neuron genome are differentially methylated.[19]
Maintenance methyltransferases add methylation to DNA when one strand is already methylated. These work throughout the life of the organism to maintain the methylation pattern that had been established by the de novo methyltransferases.[citation needed]
Mammalian
At least four differently active DNA methyltransferases have been identified in mammals. They are named

Manzo et al.[24] observed differences in genomic binding of DNMT3a1, DNMT3a2 and DNMT3b. They found 3,970 regions exclusively enriched for DNMT3a1, 3,838 exclusively enriched for DNMT3a2 and 3,432 exclusively enriched for DNMT3b.
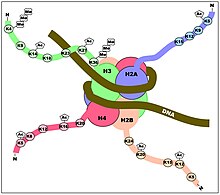
The DNMT enzymes are not only regulated in their methylating locations on the genome by where they bind to DNA,[24] but they are also regulated by the post-translational modifications on the histone proteins of the nucleosomes around which the genomic DNA is wrapped (see Figures). Rose and Klose[25] reviewed the relationship between DNA methylation and histone lysine methylation. For example, they indicated that H3K4me3 appears to block DNA methylation while H3K9me3 plays a role in promoting DNA methylation.
DNMT3L[26] is a protein closely related to DNMT3a and DNMT3b in structure and critical for DNA methylation, but appears to be inactive on its own.
DNMT1
DNMT1 is the most abundant DNA methyltransferase in mammalian cells, and considered to be the key maintenance methyltransferase in mammals. It predominantly methylates hemimethylated CpG di-nucleotides in the mammalian genome. The recognition motif for the human enzyme involves only three of the bases in the CpG dinucleotide pair: a C on one strand and CpG on the other. This relaxed substrate specificity requirement allows it to methylate unusual structures like DNA slippage intermediates at de novo rates that equal its maintenance rate.[27] Like other DNA cytosine-5 methyltransferases the human enzyme recognizes flipped out cytosines in double stranded DNA and operates by the nucleophilic attack mechanism.[28] In human cancer cells DNMT1 is responsible for both de novo and maintenance methylation of tumor suppressor genes.[29][30] The enzyme is about 1,620 amino acids long. The first 1,100 amino acids constitute the regulatory domain of the enzyme, and the remaining residues constitute the catalytic domain. These are joined by Gly-Lys repeats. Both domains are required for the catalytic function of DNMT1.[citation needed]
DNMT1 has several
DNMT1 null mutant embryonic stem cells were viable and contained a small percentage of methylated DNA and methyltransferase activity. Mouse embryos homozygous for a deletion in Dnmt1 die at 10–11 days gestation.[31]
TRDMT1
Although this enzyme has strong sequence similarities with 5-methylcytosine methyltransferases of both prokaryotes and eukaryotes, in 2006, the enzyme was shown to methylate position 38 in aspartic acid transfer RNA and does not methylate DNA.[32] The name for this methyltransferase has been changed from DNMT2 to TRDMT1 (tRNA aspartic acid methyltransferase 1) to better reflect its biological function.[33] TRDMT1 is the first RNA cytosine methyltransferase to be identified in human cells.
DNMT3
DNMT3 is a family of DNA methyltransferases that could methylate hemimethylated and unmethylated CpG at the same rate. The architecture of DNMT3 enzymes is similar to that of DNMT1, with a regulatory region attached to a catalytic domain. There are at least five members of the DNMT3 family: DNMT3a1, DNMT3a2, 3b, 3c and 3L.[citation needed]
DNMT3a1, DNMT3a2 and DNMT3b can mediate methylation of CpG sites in gene promoters, resulting in gene repression. These DNA methyltransferases can also methylate CpG sites within the coding regions of genes, where such methylation can increase gene transcription.[34] Work with DNMT3a1 showed it preferentially localized to CpG islands bivalently marked by H3K4me3 (a transcription promoting mark) and H3K27me3 (a transcription repressive mark), coinciding with the promoters of many transcription factors. Work with DNMT3a2, in neurons, found that the DNA methylation changes caused by DNMT3a2 predominantly occur in intergenic and intronic regions. These intergenic and intronic DNA methylations were thought to likely regulate enhancer activity, alternative splicing or the expression of non-coding RNAs.[35]
DNMT3a1 can co-localize with heterochromatin protein (HP1) and methyl-CpG-binding protein (MeCBP), among a number of other factors.[36] They can also interact with DNMT1, which might be a co-operative event during DNA methylation. DNMT3a prefers CpG methylation to CpA, CpT, and CpC methylation, though there appears to be some sequence preference of methylation for DNMT3a and DNMT3b. DNMT3a methylates CpG sites at a rate much slower than DNMT1, but greater than DNMT3b.
The expression of DNMT3a2 differs from DNMT3a1 and DNMT3b because DNMT3a2 expression occurs in the pattern of an immediate early gene. DNMT3a2 is induced to express in neurons, for instance, by new neuronal activity.[37][35] This may be of importance in establishing long-term memory.[38] In a rat, high levels of new DNA methylations in neurons of the hippocampus occur after a memorable event is imposed on a rat, such as contextual fear conditioning.[19] Bayraktar and Kreutz[39] found that DNMT inhibitors, applied in the brain, prevented long-term memories from forming.
DNMT3L contains DNA methyltransferase
Clinical significance
DNMT inhibitors
Because of the
- guadecitabine, an experimental drug under development by Astex Pharmaceuticals and Otsuka Pharmaceutical. It failed to meet primary endpoints in a 2018 Phase III AML trial.
See also
References
- PMID 26660621.
- PMID 3323532.
- PMID 3248728.
- PMID 2541254.
- ^ PMID 7607512.
- PMID 7971991.
- PMID 1833555.
- PMID 27476077.
- PMID 19766657.
- PMID 26475175.
- PMID 31768029.
- PMID 3248729.
- ^ PMID 8127644.
- PMID 2716049.
- PMID 7773746.
- S2CID 54238106.
- PMID 32417228.
- S2CID 18241370.
- ^ PMID 28620075.
- ^ "DNMT1". Gene Symbol Report. HUGO Gene Nomenclature Committee. Archived from the original on 2012-10-06. Retrieved 2012-09-27.
- PMID 12138111.
- ^ "DNMT3B". Gene Symbol Report. HUGO Gene Nomenclature Committee. Archived from the original on 2012-11-20. Retrieved 2012-09-27.
- S2CID 30907442.
- ^ PMID 29074627.
- PMID 24560929.
- ^ "DNMT3L". Gene Symbol Report. HUGO Gene Nomenclature Committee. Retrieved 2012-09-27.
- PMID 9451440.
- PMID 1584813.
- PMID 16423997.
- PMID 16424002.
- S2CID 19879601.
- S2CID 39089541.
- ^ "TRDMT1 tRNA aspartic acid methyltransferase 1 (Homo sapiens)". Entrez Gene. NCBI. 2010-11-01. Retrieved 2010-11-07.
- PMID 25263941.
- ^ PMID 32005851.
- PMID 33604794.
- S2CID 10590208.
- PMID 35098021.
- PMID 29875631.
- S2CID 11480326.
- ^ "EC Approves Marketing Authorization Of DACOGEN For Acute Myeloid Leukemia". 2012-09-28. Retrieved 28 September 2012.
Further reading
- Smith SS (1994). Biological implications of the mechanism of action of human DNA (cytosine-5)methyltransferase. Progress in Nucleic Acid Research and Molecular Biology. Vol. 49. pp. 65–111. PMID 7863011.
- Pradhan S, Esteve PO (October 2003). "Mammalian DNA (cytosine-5) methyltransferases and their expression". Clinical Immunology. 109 (1): 6–16. PMID 14585271.
- Goll MG, Bestor TH (2005). "Eukaryotic cytosine methyltransferases". Annual Review of Biochemistry. 74: 481–514. S2CID 32123961.
External links
- Information about DNA methyltransferases and DNA methylation at epigeneticstation.com
- Data for a DNA methyltransferase (DNMT) Antibody
- DNA+Modification+Methyltransferases at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
