Bacteriophage MS2
| Emesvirus zinderi | |
|---|---|
| |
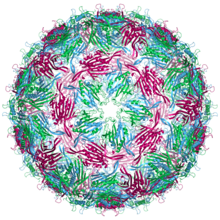
| Bacteriophage MS2 conformers are labelled blue (chain a), green (chain b) and magenta (chain c)
| |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Lenarviricota |
| Class: | Leviviricetes |
| Order: | Norzivirales |
| Family: | Fiersviridae |
| Genus: | Emesvirus |
| Species: | Emesvirus zinderi
|
Bacteriophage MS2 (Emesvirus zinderi), commonly called MS2, is an icosahedral,
It is small and contains a maturation protein, coat protein, and genomic RNA. It also has one of the smallest known genomes, encoding four proteins.
The MS2 lifecycle involves infecting bacteria with the fertility factor, enabling the virus to attach to the pilus, though the mechanism by which the virus's RNA enters the bacterium remains unknown. Once inside, the viral RNA starts functioning as a messenger RNA to produce viral proteins. MS2 replicates its plus-strand genome by creating a minus strand RNA as a template. The virus then assembles, and the bacterial cell lyses, releasing new viruses.
The virus was isolated in 1961 and its genome was the first to be fully sequenced, in 1976, providing a crucial understanding of genetic codes. In practical applications, MS2's structural components have been used to detect RNA in living cells. The virus is also under research for potential uses in drug delivery, tumor imaging, and light harvesting. Furthermore, because of its structural similarities to
Virology
Genome

| Gene | Size | Gene product | aa |
|---|---|---|---|
| mat
(MS2g1) |
1487 nt | maturation | 393 |
| cp
(MS2g2) |
510 nt | coat protein | 130 |
| lys
(MS2g3) |
295 nt | lysis protein | 75 |
| rep
(MS2g4) |
2055 nt | RNA replicase, | 545 |
The MS2 genome is one of the smallest known, consisting of 3569 nucleotides of single-stranded RNA.
Structure

An MS2
The structure of the coat protein is a five-stranded β-sheet with two α-helices and a hairpin. When the capsid is assembled, the helices and hairpin face the exterior of the particle, while the β-sheet faces the interior.[7]
Life cycle
MS2 infects enteric bacteria carrying the
Once the viral RNA has entered the cell, it begins to function as a messenger RNA for the production of phage proteins. The gene for the most abundant protein, the coat protein, can be immediately translated. The translation start of the replicase gene is normally hidden within RNA secondary structure, but can be transiently opened as ribosomes pass through the coat protein gene. Replicase translation is also shut down once large amounts of coat protein have been made; coat protein dimers bind and stabilize the RNA "operator hairpin", blocking the replicase start. The start of the maturation protein gene is accessible in RNA being replicated but hidden within RNA secondary structure in the completed MS2 RNA; this ensures translation of only a very few copies of maturation protein per RNA. Finally, the lysis protein gene can only be initiated by ribosomes that have completed translation of the coat protein gene and "slip back" to the start of the lysis protein gene, at about a 5% frequency.[1]

Replication of the plus-strand MS2 genome requires synthesis of the complementary minus strand RNA, which can then be used as a template for synthesis of a new plus strand RNA. MS2 replication has been much less well studied than replication of the highly related
The formation of the virion is thought to be initiated by binding of maturation protein to the MS2 RNA; in fact, the complex of maturation protein and RNA is infectious. The assembly of the icosahedral shell or capsid from coat proteins can occur in the absence of RNA; however, capsid assembly is nucleated by coat protein dimer binding to the operator hairpin, and assembly occurs at much lower concentrations of coat protein when MS2 RNA is present.[1]
Bacterial lysis and release of newly formed virions occurs when sufficient lysis protein has accumulated. Lysis (L) protein forms pores in the cytoplasmic membrane, which leads to loss of
MS2 in History of Science and Use
In 1961, MS2 was isolated by Alvin John Clark and recognized as an RNA-containing phage very similar to bacteriophage f2.[10]
In 1976, the MS2 genome was the first genome to be completely sequenced.[3] This was accomplished by Walter Fiers and his team, building upon their earlier milestone in 1972 of the first gene to be completely sequenced, the MS2 coat protein.[11] These sequences were determined at the RNA level.[12] The first effort at a statistical analysis of the MS2 genome was a search for patterns in the nucleotide sequence. Several non-coding sequences were identified, however at the time of this investigation (1979), the functions of the non-coding patterns were unknown.[13]
Since 1998,
MS2, due to its structural similarities to noroviruses, its similar optimum proliferation conditions, and non-pathogenicity to humans, has been used as substitute for noroviruses in studies of disease transmission.[16]
See also
- bacteriophage
- bacteriophage f2
- bacteriophage Qβ
- phi-X174 phage
References
- ^ ISBN 978-0195148503.
- PMID 8976557.
- ^ S2CID 4289674.
- PMID 13978804.
- S2CID 2803228.
- PMID 9464373.
- PMID 8254664.
- PMID 28396351.
- PMID 28691656.
- PMID 17795773.
- S2CID 4153893.
- S2CID 4206886.
- S2CID 85199492.
- PMID 9809065.
- S2CID 6212583.
- ^ Fox M (8 September 2014). "Viruses spread 'like crazy' in an office, study finds". The Today Show.
External links
- Complete genome (also isolates R17, DL16, and J20)
