Cdc6
| Cell division protein Cdc6/18 | |
|---|---|
| Identifiers | |
| Symbol | Cdc6 |
| InterPro | IPR016314 |
| Orc1/Cdc6-type DNA replication protein, archaea | |
|---|---|
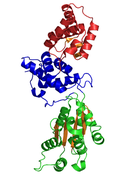 Crystal Structure of CDC6 from P. aerophilum (PDB: 1FNN). Green and blue domains are AAA domains; red domain is the Cdc6 C-terminal HTH domain (infobox below).[1] | |
| Identifiers | |
| Symbol | Orc1/Cdc6_arc |
| InterPro | IPR014277 |
| Cdc6/Orc1, C-terminal | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Cdc6_C | ||||||||
| Pfam | PF09079 | ||||||||
| InterPro | IPR015163 | ||||||||
| CDD | cd08768 | ||||||||
| |||||||||
Cdc6, or cell division cycle 6, is a
Function

CDC6 is an
Regulation
CDC6 is normally present at high levels during the G1 phase of the
Two states can be distinguished. In the first state (during G1 phase) Cdk2-activity is low, CDC6 can accumulate, hence the pre-RC can be formed but not activated. In the second state Cdk2-activity is high, CDC6 becomes inactivated, hence the pre-RC is activated but not formed. This change assures that DNA replication is performed only once per cell cycle. It has been shown that overexpression of CDC6 does not induce re-replication in cognate cells, probably due to inhibition through CDK that resets the cell cycle clock to G1. Nevertheless, it has been suggested that regulation of CDC6 is one of several redundant mechanisms that prevent re-replication of the DNA in eukaryotic cells.[10]
Structure

The crystallographic structure of a Cdc6/Orc1-related protein from the archaeon Pyrobaculum aerophilum has been solved and three structural domains have been identified.[1] Domain I and II form the ATP binding/hydrolysis site and are similar to other AAA+ ATPases. Domain III is structurally related to a winged-helix domain, thus may interact with origin DNA. From studies with E. coli γ clamp loading complex, it was suggested that domain III mediates protein-protein interactions with other AAA+ ATPases in the pre-RC, thus suggesting that the CDC6 builds a homodimer in its native form. The domains I and II form a cashew-shaped molecule that bind ATP in the cleft and additionally build the sensor motif for ATP/ADP recognition. These domains are also thought to mediate subsequent conformational changes. Nevertheless, the exact functional roles of these domains remain unclear.[8]
Disease
It has been shown CDC6 shows
