Origin recognition complex
| Origin recognition complex subunit 2 | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | ORC2 | ||||||||
| Pfam | PF04084 | ||||||||
| InterPro | IPR007220 | ||||||||
| |||||||||
| Origin recognition complex (ORC) subunit 3 N-terminus | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | ORC3_N | ||||||||
| Pfam | PF07034 | ||||||||
| InterPro | IPR010748 | ||||||||
| |||||||||
| Origin recognition complex subunit 6 (ORC6) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | ORC6 | ||||||||
| Pfam | PF05460 | ||||||||
| InterPro | IPR008721 | ||||||||
| |||||||||
In
ORC directs
The ORC is present throughout the cell cycle bound to replication origins, but is only active in late mitosis and early G1.
In yeast, ORC also plays a role in the establishment of silencing at the
Both Orc1 and Orc5 bind ATP, though only Orc1 has
Proteins
The following proteins are present in the ORC:
| S. cerevisiae | S. pombe | D. melanogaster | Vertebrates |
|---|---|---|---|
| ORC 1-6 | ORC 1-6 | ORC 1-6 | ORC 1-6 |
| Cdc6 | Cdc18 | Cdc6 | Cdc6 |
| Cdt1/Tah11/Sid2 | Cdt1 | DUP | Cdt1/RLF-B |
| Mcm2 | Mcm2/Cdc19/Nda1 | Mcm2 | Mcm2 |
| Mcm3 | Mcm3 | Mcm3 | Mcm3 |
| Cdc54/Mcm4 | Cdc21 | DPA | Mcm4 |
| Cdc46/Mcm5 | Mcm5/Nda4 | Mcm5 | Mcm5 |
| Mcm6 | Mcm6/Mis5 | Mcm6 | Mcm6 |
| Cdc47/Mcm7 | Mcm7 | Mcm7 | mcm7 |
Archaea feature a simplified version of the ORC, Mcm, and as a consequence the combined pre-RC. Instead of using six different mcm proteins to form a pseudo-symmetrical heterohexamer, all six subunits in the archaeal MCM are the same. They usually have multiple proteins that are homologous to both Cdc6 and Orc1, some of which perform the function of both. Unlike eukaryotic Orc, they do not always form a complex. In fact, they have divergent complex structures when these do form. Sulfolobus islandicus also uses a Cdt1 homologue to recognize one of its replication origins.[28]
Autonomously replicating sequences
Budding yeast
Autonomously Replicating Sequences (ARS), first discovered in budding yeast, are integral to the success of the ORC. These 100-200bp sequences facilitate replication activity during S phase. ARSs can be placed at any novel location of the chromosomes of budding yeast and will facilitate replication from those sites. A highly conserved sequence of 11bp (known as the A element) is thought to be essential for origin function in budding yeast.[27] The ORC was originally identified by its ability to bind to the A element of the ARS in budding yeast.
Animals
Animal cells contain a much more cryptic version of an ARS, with no
Role in pre-RC assembly
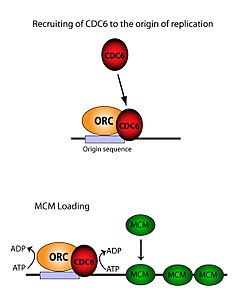
The ORC is essential for the loading of MCM complexes (
Origin binding activity
Although the ORC is composed of six discrete subunits, only one of these has been found to be significant - ORC1. In vivo studies have shown that Lys-263 and Arg-367 are the basic residues responsible for faithful ORC loading. These molecules represent the above-mentioned ARS.[33] ORC1 interacts with ATP and these basic residues in order to bind the ORC to origin DNA. It has been established that this occurs far before replication, and that the ORC itself is already bound to Origin DNA by the time any Mcm2-7 loading occurs.[31] When Mcm2-7 is first loaded it completely encircles the DNA and helicase activity is inhibited. In S phase, the Mcm2-7 complex interacts with helicase cofactors Cdc45 and GINS to isolate a single DNA strand, unwind the origin, and begin replication down the chromosome. In order to have bidirectional replication, this process happens twice at an origin. Both loading events are mediated by one ORC via an identical process as the first.[34]
See also
- Cyclin dependant kinases (CDK)
- Cyclins
- DNA helicase
- DnaA
- Pre-replication complex
References
- ^ Origin+Recognition+Complex at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- PMID 9442876.
- PMID 17241905.
- ^ PMID 17825065.
- ^ S2CID 4346767.
- ^ PMID 7585959.
- ^ S2CID 22439595.
- PMID 12110182.
- PMID 7892251.
- PMID 7781615.
- ^ PMID 16228006.
- PMID 10966477.
- PMID 12045100.
- S2CID 33220937.
- PMID 11572976.
- S2CID 4393812.
- PMID 16024805.
- S2CID 4309206.
- PMID 9171055.
- PMID 9038340.
- PMID 11459976.
- PMID 15610739.
- PMID 16387651.
- PMID 17053779.
- PMID 33992866.
- PMID 32160540.
- ^ ISBN 978-0878935086.
- PMID 28146124.
- PMID 23603117.
- PMID 16387651.
- ^ PMID 16228006.
- PMID 28191893.
- PMID 26456755.
- PMID 25910200.
Further reading
- PMID 12045100.
A comprehensive review of molecular DNA replication