De novo gene birth
De novo gene birth is the process by which new genes evolve from non-coding DNA.[1][3] De novo genes represent a subset of novel genes, and may be protein-coding or instead act as RNA genes.[4] The processes that govern de novo gene birth are not well understood, although several models exist that describe possible mechanisms by which de novo gene birth may occur.
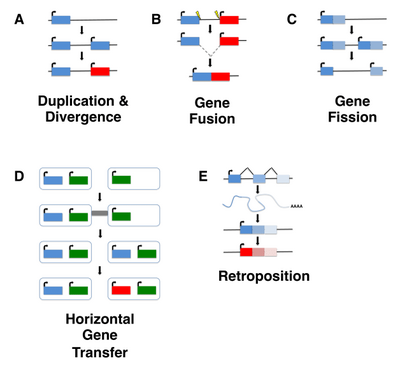
Although de novo gene birth may have occurred at any point in an organism's evolutionary history, ancient de novo gene birth events are difficult to detect. Most studies of de novo genes to date have thus focused on young genes, typically taxonomically restricted genes (TRGs) that are present in a single species or lineage, including so-called orphan genes, defined as genes that lack any identifiable homolog. It is important to note, however, that not all orphan genes arise de novo, and instead may emerge through fairly well characterized mechanisms such as gene duplication (including retroposition) or horizontal gene transfer followed by sequence divergence, or by gene fission/fusion.[5][6]
Although de novo gene birth was once viewed as a highly unlikely occurrence,[7] several unequivocal examples have now been described,[8] and some researchers speculate that de novo gene birth could play a major role in evolutionary innovation, morphological specification, and adaptation,[9][10] probably promoted by their low level of pleiotropy.
History
As early as the 1930s, J. B. S. Haldane and others suggested that copies of existing genes may lead to new genes with novel functions.[6] In 1970, Susumu Ohno published the seminal text Evolution by Gene Duplication.[11] For some time subsequently, the consensus view was that virtually all genes were derived from ancestral genes,[12] with François Jacob famously remarking in a 1977 essay that "the probability that a functional protein would appear de novo by random association of amino acids is practically zero."[7]
In the same year, however, Pierre-Paul Grassé coined the term "overprinting" to describe the emergence of genes through the expression of alternative open reading frames (ORFs) that overlap preexisting genes.[13] These new ORFs may be out of frame with or antisense to the preexisting gene. They may also be in frame with the existing ORF, creating a truncated version of the original gene, or represent 3’ extensions of an existing ORF into a nearby ORF. The first two types of overprinting may be thought of as a particular subtype of de novo gene birth; although overlapping with a previously coding region of the genome, the primary amino-acid sequence of the new protein is entirely novel and derived from a frame that did not previously contain a gene. The first examples of this phenomenon in bacteriophages were reported in a series of studies from 1976 to 1978,[14][15][16] and since then numerous other examples have been identified in viruses, bacteria, and several eukaryotic species.[17][18][19][20][21][22]
The phenomenon of exonization also represents a special case of de novo gene birth, in which, for example, often-repetitive intronic sequences acquire splice sites through mutation, leading to de novo exons. This was first described in 1994 in the context of Alu sequences found in the coding regions of primate mRNAs.[23] Interestingly, such de novo exons are frequently found in minor splice variants, which may allow the evolutionary “testing” of novel sequences while retaining the functionality of the major splice variant(s).[24]
Still, it was thought by some that most or all eukaryotic proteins were constructed from a constrained pool of “starter type” exons.[25] Using the sequence data available at the time, a 1991 review estimated the number of unique, ancestral eukaryotic exons to be < 60,000,[25] while in 1992 a piece was published estimating that the vast majority of proteins belonged to no more than 1,000 families.[26] Around the same time, however, the sequence of chromosome III of the budding yeast Saccharomyces cerevisiae was released,[27] representing the first time an entire chromosome from any eukaryotic organism had been sequenced. Sequencing of the entire yeast nuclear genome was then completed by early 1996 through a massive, collaborative international effort.[28] In his review of the yeast genome project, Bernard Dujon noted that the unexpected abundance of genes lacking any known homologs was perhaps the most striking finding of the entire project.[28]
In 2006 and 2007, a series of studies provided arguably the first documented examples of de novo gene birth that did not involve overprinting.[29][30][31] These studies were conducted using the accessory gland transcriptomes of Drosophila yakuba and Drosophila erecta and they identified 20 putative lineage-restricted genes that appeared unlikely to have resulted from gene duplication.[31] Levine and colleagues identified and confirmed five de novo candidate genes specific to Drosophila melanogaster and/or the closely related Drosophila simulans through a rigorous approach that combined bioinformatic and experimental techniques.[30]
Since these initial studies, many groups have identified specific cases of de novo gene birth events in diverse organisms.[32] The first de novo gene identified in yeast, BSC4 gene was identified in S. cerevisiae in 2008. This gene shows evidence of purifying selection, is expressed at both the mRNA and protein levels, and when deleted is synthetically lethal with two other yeast genes, all of which indicate a functional role for the BSC4 gene product.[33] Historically, one argument against the notion of widespread de novo gene birth is the evolved complexity of protein folding. Interestingly, Bsc4 was later shown to adopt a partially folded state that combines properties of native and non-native protein folding.[34] In plants, the first de novo gene to be functionally characterized was QQS, an Arabidopsis thaliana gene identified in 2009 that regulates carbon and nitrogen metabolism.[35] The first functionally characterized de novo gene identified in mice, a noncoding RNA gene, was also described in 2009.[36] In primates, a 2008 informatic analysis estimated that 15/270 primate orphan genes had been formed de novo.[37] A 2009 report identified the first three de novo human genes, one of which is a therapeutic target in chronic lymphocytic leukemia.[38] Since this time, a plethora of genome-level studies have identified large numbers of orphan genes in many organisms, although the extent to which they arose de novo, and the degree to which they can be deemed functional, remain debated.
Identification
Identification of de novo emerging sequences
There are two major approaches to the systematic identification of novel genes: genomic phylostratigraphy[39] and synteny-based methods.[40] Both approaches are widely used, individually or in a complementary fashion.
Genomic phylostratigraphy
Genomic phylostratigraphy involves examining each gene in a focal, or reference, species and inferring the presence or absence of ancestral homologs through the use of the BLAST sequence alignment algorithms[41] or related tools. Each gene in the focal species can be assigned an age (aka “conservation level” or “genomic phylostratum”) that is based on a predetermined phylogeny, with the age corresponding to the most distantly related species in which a homolog is detected.[39] When a gene lacks any detectable homolog outside of its own genome, or close relatives, it is said to be a novel, taxonomically restricted or orphan gene.
Phylostratigraphy is limited by the set of closely related genomes that are available, and results are dependent on BLAST search criteria.[42] In addition, it is often difficult to determine based on lack of observed sequence similarity whether a novel gene has emerged de novo or has diverged from an ancestral gene beyond recognition, for instance following a duplication event. This was pointed out by a study that simulated the evolution of genes of equal age and found that distant orthologs can be undetectable for rapidly evolving genes.[43] On the other hand, when accounting for changes in the rate of evolution in young regions of genes, a phylostratigraphic approach was more accurate at assigning gene ages in simulated data.[44] Subsequent studies using simulated evolution found that phylostratigraphy failed to detect an ortholog in the most distantly related species for 13.9% of D. melanogaster genes and 11.4% of S. cerevisiae genes.[45][46] However, a reanalysis of studies that used phylostratigraphy in yeast, fruit flies and humans found that even when accounting for such error rates and excluding difficult-to-stratify genes from the analyses, the qualitative conclusions were unaffected.[47] The impact of phylostratigraphic bias on studies examining various features of de novo genes remains debated.
Synteny-based approaches
Synteny-based approaches use order and relative positioning of genes (or other features) to identify the potential ancestors of candidate de novo genes.[10][42] Syntenic alignments are anchored by conserved “markers.” Genes are the most common marker in defining syntenic blocks, although k-mers and exons are also used.[48][40] Confirmation that the syntenic region lacks coding potential in outgroup species allows a de novo origin to be asserted with higher confidence.[42] The strongest possible evidence for de novo emergence is the inference of the specific "enabling" mutation(s) that created coding potential, typically through the analysis of smaller sequence regions, termed microsyntenic regions, of closely related species.
One challenge in applying synteny-based methods is that synteny can be difficult to detect across longer timescales. To address this, various optimization techniques have been created, such as using exons clustered irrespective of their specific order to define syntenic blocks[40] or algorithms that use well-conserved genomic regions to expand microsyntenic blocks.[49] There are also difficulties associated with applying synteny-based approaches to genome assemblies that are fragmented[50] or in lineages with high rates of chromosomal rearrangements, as is common in insects.[51] Synteny-based approaches can be applied to genome-wide surveys of de novo genes[37][38][52][53][54][55][56][57] and represent a promising area of algorithmic development for gene birth dating. Some have used synteny-based approaches in combination with similarity searches in an attempt to develop standardized, stringent pipelines[58] that can be applied to any group of genomes in an attempt to address discrepancies in the various lists of de novo genes that have been generated.
Determination of status
Even when the evolutionary origin of a particular coding sequence has been established, there is still a lack of consensus about what constitutes a genuine de novo gene birth event. One reason for this is a lack of agreement on whether or not the entirety of the sequence must be non-genic in origin. For protein-coding de novo genes, it has been proposed that de novo genes be divided into subtypes based on the proportion of the ORF in question that was derived from a previously noncoding sequence.[42] Furthermore, for de novo gene birth to occur, the sequence in question must be a gene which has led to a questioning of what constitutes a gene, with some models establishing a strict dichotomy between genic and non-genic sequences, and others proposing a more fluid continuum.[59]
All definitions of genes are linked to the notion of function, as it is generally agreed that a genuine gene should encode a functional product, be it RNA or protein. There are, however, different views of what constitutes function, depending whether a given sequence is assessed using genetic, biochemical, or evolutionary approaches.[42][60][61][62] The ambiguity of the concept of ‘function’ is especially problematic for the de novo gene birth field, where the objects of study are often rapidly evolving.[62] To address these challenges, the Pittsburgh Model of Function deconstructs ‘function’ into five meanings to describe the different properties that are acquired by a locus undergoing de novo gene birth: Expression, Capacities, Interactions, Physiological Implications, and Evolutionary Implications.[62]
It is generally accepted that a genuine de novo gene is expressed in at least some context,
Genetic approaches to detect a specific phenotype or change in fitness upon disruption of a particular sequence, are useful to infer function.[61] Other experimental approaches, including screens for protein-protein and/or genetic interactions, may also be employed to confirm a biological effect for a particular de novo ORF.
Evolutionary approaches may be employed to infer the existence of a molecular function from computationally derived signatures of selection. In the case of TRGs, one common signature of selection is the ratio of nonsynonymous to synonymous substitutions (
Prevalence
Estimates of numbers
Frequency and number estimates of de novo genes in various lineages vary widely and are highly dependent on methodology. Studies may identify de novo genes by phylostratigraphy/BLAST-based methods alone, or may employ a combination of computational techniques, and may or may not assess experimental evidence for expression and/or biological role.[10] Furthermore, genome-scale analyses may consider all or most ORFs in the genome,[59] or may instead limit their analysis to previously annotated genes.
The D. melanogaster lineage is illustrative of these differing approaches. An early survey using a combination of BLAST searches performed on cDNA sequences along with manual searches and synteny information identified 72 new genes specific to D. melanogaster and 59 new genes specific to three of the four species in the D. melanogaster species complex. This report found that only 2/72 (~2.8%) of D. melanogaster-specific new genes and 7/59 (~11.9%) of new genes specific to the species complex were derived de novo,[56] with the remainder arising via duplication/retroposition. Similarly, an analysis of 195 young (<35 million years old) D. melanogaster genes identified from syntenic alignments found that only 16 had arisen de novo.[54] In contrast, an analysis focused on transcriptomic data from the testes of six D. melanogaster strains identified 106 fixed and 142 segregating de novo genes.[55] For many of these, ancestral ORFs were identified but were not expressed. A newer study found that up to 39 % of orphan genes in the Drosophila clade may have emerged de novo, as they overlap with non-coding regions of the genome.[67] Highlighting the differences between inter- and intra-species comparisons, a study in natural Saccharomyces paradoxus populations found that the number of de novo polypeptides identified more than doubled when considering intra-species diversity.[68] In primates, one early study identified 270 orphan genes (unique to humans, chimpanzees, and macaques), of which 15 were thought to have originated de novo.[37] Later reports identified many more de novo genes in humans alone that are supported by transcriptional and proteomic evidence.[57][69] Studies in other lineages/organisms have also reached different conclusions with respect to the number of de novo genes present in each organism, as well as the specific sets of genes identified. A sample of these large-scale studies is described in the table below.
Generally speaking, it remains debated whether duplication and divergence or de novo gene birth represent the dominant mechanism for the emergence of new genes,[54][56][59][70][71][72] in part because de novo genes are likely to both emerge and be lost more frequently than other young genes. In a study on the origin of orphan genes in 3 different eukaryotic lineages, authors found that on average only around 30% of orphan genes can be explained by sequence divergence.[72]
Dynamics
It is important to distinguish between the frequency of de novo gene birth and the number of de novo genes in a given lineage. If de novo gene birth is frequent, it might be expected that genomes would tend to grow in their gene content over time; however, the gene content of genomes is usually relatively stable.[10] This implies that a frequent gene death process must balance de novo gene birth, and indeed, de novo genes are distinguished by their rapid turnover relative to established genes. In support of this notion, recently emerged Drosophila genes are much more likely to be lost, primarily through pseudogenization, with the youngest orphans being lost at the highest rate;[73] this is despite the fact that some Drosophila orphan genes have been shown to rapidly become essential.[54] A similar trend of frequent loss among young gene families was observed in the nematode genus Pristionchus.[74] Similarly, an analysis of five mammalian transcriptomes found that most ORFs in mice were either very old or species specific, implying frequent birth and death of de novo transcripts.[71] A comparable trend could be shown by further analyses of six primate transcriptomes.[69] In wild S. paradoxus populations, de novo ORFs emerge and are lost at similar rates.[68] Nevertheless, there remains a positive correlation between the number of species-specific genes in a genome and the evolutionary distance from its most recent ancestor.[75][67] A rapid gain and loss of de novo genes was also found on a population level by analyzing nine natural three-spined stickleback populations.[76] In addition to the birth and death of de novo genes at the level of the ORF, mutational and other processes also subject genomes to constant “transcriptional turnover”. One study in murines found that while all regions of the ancestral genome were transcribed at some point in at least one descendant, the portion of the genome under active transcription in a given strain or subspecies is subject to rapid change.[77] The transcriptional turnover of noncoding RNA genes is particularly fast compared to coding genes.[78]
Examples de novo genes
| Organism/Lineage | Gene | Evidence of
de novo origin |
Evidence of selection | Phenotypic evidence | Year discovered | Notes | Ref. |
|---|---|---|---|---|---|---|---|
| Arabidopsis thaliana | QQS | N/A | Excess leaf starch in RNAi knockdowns | 2009 | [35] | ||
| Drosophila | CG9284 | Syntenic alignments of 12 Drosophila species | RNAi knockdown is lethal | 2010 | [54] | ||
| Drosophila | CG30395 | Syntenic alignments of 12 Drosophila species | RNAi knockdown is lethal | 2010 | [54] | ||
| Drosophila | CG31882 | Syntenic alignments of 12 Drosophila species | RNAi knockdown is lethal | 2010 | [54] | ||
| Drosophila | CG31406 | tBLASTn of protein-coding regions to all 12 Drosophila genomes and comparison of BLASTZ alignments | dN/dS <1 indicates purifying selection | RNAi knockdown inhibits fertility | 2013 | [79] | |
| Drosophila | CG32582 | tBLASTn of protein-coding regions to all 12 Drosophila genomes and comparison of BLASTZ alignments | Possible positive selection but not statistically significant | RNAi knockdown inhibits fertility | 2013 | [79] | |
| Drosophila | CG33235 | tBLASTn of protein-coding regions to all 12 Drosophila genomes and comparison of BLASTZ alignments | dN/dS <1 indicates purifying selection | RNAi knockdown inhibits fertility | 2013 | [79] | |
| Drosophila | CG34434 | tBLASTn of protein-coding regions to all 12 Drosophila genomes and comparison of BLASTZ alignments | dN/dS <1 indicates purifying selection | RNAi knockdown inhibits fertility | 2013 | [79] | |
| Drosophila melanogaster | goddard | Genome-wide tblastn searches and LASTZ- and Exonerate-based analyses of the syntenic regions | essential for individualization of elongated spermatids;
RNAi knockdown experiments in male flies |
2017 | Structure prediction: half disordered, half alpha-helical | [80][81] | |
| Drosophila simulans and Drosophila sechellia | Dsim_GD19764 and Dsec_GM10790 | Exonerate-based analyses of the syntenic regions | Conservation across two sister species | Testes expression | 2020 | Born inside intron of another gene, contains conserved intron present at time of birth (length not multiple of 3). Structure prediction: contains a transmembrane alpha helix | [82] |
| Gadidae | AFGP | Examination of Gadid phylogeny | Gene multiplied in Gadid species in colder habitats but decayed in species not under threat of freezing | Inhibit ice growth formation | Function is similar to other antifreeze proteins that evolved independently | [83][84] | |
| Mus | Gm13030 | Combined phylostratigraphy and synteny approach | ORF only retained in M. m. musculus and M. m. castaneus populations; no evidence of positive selection | Knockout mutant has irregular pregnancy cycles | 2019 | [85] | |
| Mus | Poldi | Homologous region not expressed in closely related and outgroup species | Evidence of recent selective sweep in M. m. musculus | Knockout mutant has reduced sperm motility and testis weight | 2009 | RNA gene | [36] |
| Placental Mammals | ORF-Y | PhyloCSF of POLG gene in Homo sapiens, synonymous site conservation across mammals, and tBLASTN of mammals, sauropsids, amphibians, and teleost fish | Disappearance of enhanced synonymous site conservation within the POLG ORF after the ORF-Y’s stop codon and high conservation of the initiation context of the start codon indicate purifying selection | 41 Clinvar variants that affect the ORF-Y peptide but not the amino acid sequence of POLG | 2020 | [22] | |
| Saccharomyces
cerevisiae |
BSC4 | tBLASTN and syntenic alignments of closely related species | Under negative selection based on population data | Has two synthetic lethal partners | 2008 | Adopts a partially specific three-dimensional structure | [33][34] |
| Saccharomyces
cerevisiae |
MDF1 | Only identified putative homologs are truncated, non-expressed, non-functional ORFs | Fixed in 39 diverse strains, no frameshift or nonsense mutations | Decreases mating efficiency by binding MATα2; promotes growth through an interaction with Snf1 | 2010 | Expression is suppressed by its antisense gene | [86][87] |
Features
General Features
Recently emerged de novo genes differ from established genes in a number of ways. Across a broad range of species, young and/or taxonomically restricted genes have been reported to be shorter in length than established genes, more positively charged, faster evolving,[88] and to be less expressed.[37][59][73][74][89][90][91][92][93][94][95][96][71][69][67][76][excessive citations] Although these trends could be a result of homology detection bias, a reanalysis of several studies that accounted for this bias found that the qualitative conclusions reached were unaffected.[47] Another feature includes the tendency for young genes to have their hydrophobic amino acids more clustered near one another along the primary sequence.[97][98]
The expression of young genes has also been found to be more tissue- or condition-specific than that of established genes.[29][31][37][55][57][59][94][99][100][101][67][76] In particular, relatively high expression of de novo genes was observed in male reproductive tissues in Drosophila, stickleback, mice, and humans, and, in the human brain.[57][102][67][76] In animals with adaptive immune systems, higher expression in the brain and testes may be a function of the immune-privileged nature of these tissues. An analysis in mice found specific expression of intergenic transcripts in the thymus and spleen (in addition to the brain and testes). It has been proposed that in vertebrates de novo transcripts must first be expressed in tissues lacking immune cells before they can be expressed in tissues that have immune surveillance.[101]
Evolutionary rate
For sequence evolution, dN/dS analysis studies often indicate that de novo genes evolve at a higher rate compared to other genes.[103][88] For expression evolution and structural evolution, quantitative studies across different evolutionary ages or phylostratigraphic branches are very few.
Features that promote de novo gene birth
Its also of interest to compare features of recently emerged de novo genes to the pool of non-genic ORFs from which they emerge. Theoretical modeling has shown that such differences are the product both of selection for features that increase the likelihood of functionalization, and of neutral evolutionary forces that influence allelic turnover.[104] Experiments in S. cerevisiae showed that predicted transmembrane domains were strongly associated with beneficial fitness effects when young ORFs were overexpressed, but not when established (older) ORFs were overexpressed.[105] Experiments in E. coli showed that random peptides tended to have more benign effects when they were enriched for amino acids that were small, and that promoted intrinsic structural disorder.[106]
Lineage-dependent features
Features of de novo genes can depend on the species or lineage being examined. This appears to partly be a result of varying GC content in genomes and that young genes bear more similarity to non-genic sequences from the genome in which they arose than do established genes.[107] Features in the resulting protein, such as the percentage of transmembrane residues and the relative frequency of various predicted secondary structural features show a strong GC dependency in orphan genes, whereas in more ancient genes these features are only weakly influenced by GC content.[107]
The relationship between gene age and the amount of predicted intrinsic structural disorder (ISD) in the encoded proteins has been subject to considerable debate. It has been claimed that ISD is also a lineage-dependent feature, exemplified by the fact that in organisms with relatively high GC content, ranging from D. melanogaster to the parasite Leishmania major, young genes have high ISD,[108][109] while in a low GC genome such as budding yeast, several studies have shown that young genes have low ISD.[59][89][96][107] However, a study that excluded young genes with dubious evidence for functionality, defined in binary terms as being under selection for gene retention, found that the remaining young yeast genes have high ISD, suggesting that the yeast result may be due to contamination of the set of young genes with ORFs that do not meet this definition, and hence are more likely to have properties that reflect GC content and other non-genic features of the genome.[110] Beyond the very youngest orphans, this study found that ISD tends to decrease with increasing gene age, and that this is primarily due to amino acid composition rather than GC content.[110] Within shorter time scales ,using de novo genes that have the most validation suggests that younger genes are more disordered in Lachancea, but less disordered in Saccharomyces.[96] Intrinsic structural disorder and aggregation propensity did not show significant differences with age in some studies of mammals [71] and primates,[69] but did in other studies of mammals.[110] One large study of the entire Pfam protein domain database showed enrichment of younger protein domain for disorder-promoting amino acids across animals, but enrichment on the basis of amino acid availability in plants.[98]
Role of epigenetic modifications
An examination of de novo genes in A. thaliana found that they are both hypermethylated and generally depleted of histone modifications.[53] In agreement with either the proto-gene model or contamination with non-genes, methylation levels of de novo genes were intermediate between established genes and intergenic regions. The methylation patterns of these de novo genes are stably inherited, and methylation levels were highest, and most similar to established genes, in de novo genes with verified protein-coding ability.[53] In the pathogenic fungus Magnaporthe oryzae, less conserved genes tend to have methylation patterns associated with low levels of transcription.[111] A study in yeasts also found that de novo genes are enriched at recombination hotspots, which tend to be nucleosome-free regions.[96]
In Pristionchus pacificus, orphan genes with confirmed expression display chromatin states that differ from those of similarly expressed established genes.[95] Orphan gene start sites have epigenetic signatures that are characteristic of enhancers, in contrast to conserved genes that exhibit classical promoters.[95] Many unexpressed orphan genes are decorated with repressive histone modifications, while a lack of such modifications facilitates transcription of an expressed subset of orphans, supporting the notion that open chromatin promotes the formation of novel genes.[95]
Structural evolution
De novo proteins typically exhibit less well-defined secondary and three-dimensional structures, often lacking rigid folding but having extensive disordered regions.[103][110] Quantitative analyses are still lacking on the evolution of secondary structural elements and tertiary structures over time. As structure is usually more conserved than sequence, comparing structures between orthologs could provide deeper insides into de novo gene emergence and evolution and help to confirm these genes as true de novo genes.[112] Nevertheless, so far only very few de novo proteins have been structurally and functionally characterized, especially due to problems with protein purification and subsequent stability. Progresses have been made using different purification tags, cell types and chaperones.[113]
The ‘antifreeze glycoprotein’ (AFGP) in Arctic codfishes prevents their blood from freezing in arctic waters.[84][83] Bsc4, a short non-essential de novo protein in yeast,[33] has been shown to be built mainly by β-sheets and has a hydrophobic core.[34] It is associated to DNA repair under nutrient-deficient conditions.[114] The Drosophila de novo protein Goddard has been characterized for the first time in 2017. Knockdown Drosophila melanogaster male flies were not able to produce sperm.[80] Recently, it could be shown that this lack was due to failure of individualization of elongated spermatids. By using computational phylogenomic and structure predictions, experimental structural analyses, and cell biological assays, it was proposed that half of Goddard's structure is disordered and the other half is composed by alpha-helical amino acids. These analyses also indicated that Goddard's orthologs show similar results. Goddard's structure therefore appears to have been mainly conserved since its emergence.[81]
Mechanisms
Pervasive expression
With the development of technologies such as RNA-seq and Ribo-seq, eukaryotic genomes are now known to be pervasively transcribed[115][116][117][118] and translated.[119] Many ORFs that are either unannotated, or annotated as long non-coding RNAs (lncRNAs), are translated at some level, either in a condition or tissue-specific manner.[59][119][120][121][122][123] Though infrequent, these translation events expose non-genic sequence to selection. This pervasive expression forms the basis for several models describing de novo gene birth.
It has been speculated that the epigenetic landscape of de novo genes in the early stages of formation may be particularly variable between and among populations, resulting in variable gene expression thereby allowing young genes to explore the “expression landscape.”[124] The QQS gene in A. thaliana is one example of this phenomenon; its expression is negatively regulated by DNA methylation that, while heritable for several generations, varies widely in its levels both among natural accessions and within wild populations.[124] Epigenetics are also largely responsible for the permissive transcriptional environment in the testes, particularly through the incorporation into nucleosomes of non-canonical histone variants that are replaced by histone-like protamines during spermatogenesis.[125]
Intergenic ORFs as elementary structural modules
Analysis of the fold potential diversity shows that the majority of the amino acid sequences encoded by the intergenic ORFs of S. cerevisiae are predicted to be foldable.[126] More importantly, these amino acid sequences with folding potential can serve as elementary building blocks for de novo genes or integrate into pre-existing genes.[126]
Order of events
For birth of a de novo protein-coding gene to occur, a non-genic sequence must both be transcribed and acquire an ORF before becoming translated. These events could occur in either order, and there is evidence supporting both an “ORF first” and a “transcription first” model.[5][127] An analysis of de novo genes that are segregating in D. melanogaster found that sequences that are transcribed had similar coding potential to the orthologous sequences from lines lacking evidence of transcription.[55] This finding supports the notion that many ORFs can exist prior to being transcribed. The antifreeze glycoprotein gene AFGP, which emerged de novo in Arctic codfishes, provides a more definitive example in which the de novo emergence of the ORF was shown to precede the promoter region.[83] Furthermore, putatively non-genic ORFs long enough to encode functional peptides are numerous in eukaryotic genomes, and expected to occur at high frequency by chance.[55][59] Through tracing the evolution history of ORF sequences and transcription activation of human de novo genes, a study showed that some ORFs were ready to confer biological significance upon their birth.[127] At the same time, transcription of eukaryotic genomes is far more extensive than previously thought, and there are documented examples of genomic regions that were transcribed prior to the appearance of an ORF that became a de novo gene.[79] The proportion of de novo genes that are protein-coding is unknown, but the appearance of “transcription first” has led some to posit that protein-coding de novo genes may first exist as RNA gene intermediates. The case of bifunctional RNAs, which are both translated and function as RNA genes, shows that such a mechanism is plausible.[128]
The two events may occur simultaneously when chromosomal rearrangement is the event that precipitates gene birth.[129]
Models
Several theoretical models and possible mechanisms of de novo gene birth have been described. The models are generally not mutually exclusive, and it is possible that multiple mechanisms may give rise to de novo genes.[42] An example is the type III antifreeze protein gene, which originates from an old sialic acid synthase (SAS) gene, in an Antarctic zoarcid fish.
“Out of Testis” hypothesis
An early case study of de novo gene birth, which identified five de novo genes in D. melanogaster, noted preferential expression of these genes in the testes,[30] and several additional de novo genes were identified using transcriptomic data derived from the testes and male accessory glands of D. yakuba and D. erecta.[29][31] This is in agreement with other studies that showed there is rapid evolution of genes related to reproduction across a range of lineages,[130][131][132] suggesting that sexual selection may play a key role in adaptive evolution and de novo gene birth. A subsequent large-scale analysis of six D. melanogaster strains identified 248 testis-expressed de novo genes, of which ~57% were not fixed.[55] A recent study on twelve Drosophila species additionally identified a higher proportion of de novo genes with testis-biased expression compared to annotated proteome.[67] It has been suggested that the large number of de novo genes with male-specific expression identified in Drosophila is likely due to the fact that such genes are preferentially retained relative to other de novo genes, for reasons that are not entirely clear.[73] Interestingly, two putative de novo genes in Drosophila (Goddard and Saturn) were shown to be required for normal male fertility.[80][81] A genetic screen of over 40 putative de novo genes with testis-enriched expression in Drosophila melanogaster revealed that one of the de novo genes, atlas, was required for proper chromatin condensation during the final stages of spermatogenesis in male. atlas evolved from the fusion of a protein-coding gene that arose at the base of Drosophila genus and a conserved non-coding RNA.[133] Comparative analysis of the transcriptomes of testis and accessory glands, a somatic tissue of males that is important for fertility, of D. melanogaster suggests that de novo genes make greater contribution to the transcriptomic complexity of testis as compared to accessory glands.[134] Single-cell RNA-seq of D. melanogaster testis revealed that the expression pattern of de novo genes was biased toward early spermatogenesis.[135]
In humans, a study that identified 60 human-specific de novo genes found that their average expression, as measured by RNA-seq, was highest in the testes.[57] Another study looking at mammalian-specific genes more generally also found enriched expression in the testes.[136] Transcription in mammalian testes is thought to be particularly promiscuous, due in part to elevated expression of the transcription machinery[137][138] and an open chromatin environment.[139] Along with the immune-privileged nature of the testes, this promiscuous transcription is thought to create the ideal conditions for the expression of non-genic sequences required for de novo gene birth. Testes-specific expression seems to be a general feature of all novel genes, as an analysis of Drosophila and vertebrate species found that young genes showed testes-biased expression regardless of their mechanism of origination.[99]
Preadaptation model
The preadaptation model of de novo gene birth uses mathematical modeling to show that when sequences that are normally hidden are exposed to weak or shielded selection, the resulting pool of “cryptic” sequences (i.e. proto-genes) can be purged of “self-evidently deleterious” variants, such as those prone to lead to protein aggregation, and thus enriched in potential adaptations relative to a completely non-expressed and unpurged set of sequences.[140] This revealing and purging of cryptic deleterious non-genic sequences is a byproduct of pervasive transcription and translation of intergenic sequences, and is expected to facilitate the birth of functional de novo protein-coding genes.[122] This is because by eliminating the most deleterious variants, what is left is, by a process of elimination, more likely to be adaptive than expected from random sequences. Using the evolutionary definition of function (i.e. that a gene is by definition under purifying selection against loss), the preadaptation model assumes that “gene birth is a sudden transition to functionality”[110] that occurs as soon as an ORF acquires a net beneficial effect. In order to avoid being deleterious, newborn genes are expected to display exaggerated versions of genic features associated with the avoidance of harm. This is in contrast to the proto-gene model, which expects newborn genes to have features intermediate between old genes and non-genes.[110]
The mathematics of the preadaptation model assume that the distribution of fitness effects is bimodal, with new sequences of mutations tending to break something or tinker, but rarely in between.[140][141] Following this logic, populations may either evolve local solutions, in which selection operates on each individual locus and a relatively high error rate is maintained, or a global solution with a low error rate which permits the accumulation of deleterious cryptic sequences.[140] De novo gene birth is thought to be favored in populations that evolve local solutions, as the relatively high error rate will result in a pool of cryptic variation that is “preadapted” through the purging of deleterious sequences. Local solutions are more likely in populations with a high effective population size.
In support of the preadaptation model, an analysis of ISD in mice and yeast found that young genes have higher ISD than old genes, while random non-genic sequences tend to show the lowest levels of ISD.[110] Although the observed trend may have partly resulted from a subset of young genes derived by overprinting,[142] higher ISD in young genes is also seen among overlapping viral gene pairs.[143] With respect to other predicted structural features such as β-strand content and aggregation propensity, the peptides encoded by proto-genes are similar to non-genic sequences and categorically distinct from canonical genes.[144]
Proto-gene model
This proto-gene model agrees with the preadaptation model about the importance of pervasive expression, and refers to the set of pervasively expressed sequences that do not meet all definitions of a gene as “proto-genes”.[59] In contrast to the preadaptation model, the proto-gene model, suggests newborn genes have features intermediate between old genes and non-genes.[110] Specifically this model envisages a more gradual process under selection from non-genic to genic state, rejecting the binary classification of gene and non-gene.
In an extension of the proto-gene model, it has been proposed that as proto-genes become more gene-like, their potential for adaptive change gives way to selected effects; thus, the predicted impact of mutations on fitness is dependent on the evolutionary status of the ORF.[105] This notion is supported by the fact that overexpression of established ORFs in S. cerevisiae tends to be less beneficial (and more harmful) than does overexpression of emerging ORFs.[105]
Several features of ORFs correlate with ORF age as determined by phylostratigraphic analysis, with young ORFs having properties intermediate between old ORFs and non-genes; this has been taken as evidence in favor of the proto-gene model, in which proto-gene state is a continuum .[59] This evidence has been criticized, because the same apparent trends are also expected under a model in which identity as a gene is a binary. Under this model, when each age group contains a different ratio of genes vs. non-genes, Simpson's paradox can generate correlations in the wrong direction.[110]
Grow slow and moult model
The “grow slow and moult” model describes a potential mechanism of de novo gene birth, particular to protein-coding genes. In this scenario, existing protein-coding ORFs expand at their ends, especially their 3’ ends, leading to the creation of novel N- and C-terminal domains.[145][146][147][148][149] Novel C-terminal domains may first evolve under weak selection via occasional expression through read-through translation, as in the preadaptation model, only later becoming constitutively expressed through a mutation that disrupts the stop codon.[140][146] Genes experiencing high translational readthrough tend to have intrinsically disordered C-termini.[150] Furthermore, existing genes are often close to repetitive sequences that encode disordered domains. These novel, disordered domains may initially confer some non-specific binding capability that becomes gradually refined by selection. Sequences encoding these novel domains may occasionally separate from their parent ORF, leading or contributing to the creation of a de novo gene.[146] Interestingly, an analysis of 32 insect genomes found that novel domains (i.e. those unique to insects) tend to evolve fairly neutrally, with only a few sites under positive selection, while their host proteins remain under purifying selection, suggesting that new functional domains emerge gradually and somewhat stochastically.[151]
Escape from adaptive conflict
The evolutionary model escape from adaptive conflict (EAC) proposes a possible way for new gene duplication to be fixed: conflict due to contrasting function within a single gene drives the fixation of new duplication.[152][153]
Pleiotropy-barrier model
The 'pleiotropy-barrier' model suggests that newly evolved genes, including de novo genes and duplication-related genes, could facilitate evolutionary innovation or evolution of specific functions due to their low (or no) pleiotropic effect, when facing new selective force, based on observations from human gene-disease data.
Human health
In addition to its significance for the field of evolutionary biology, de novo gene birth has implications for human health. It has been speculated that novel genes, including de novo genes, may play an outsized role in species-specific traits;[6][10][32][154] however, many species-specific genes lack functional annotation.[136] Nevertheless, there is evidence to suggest that human-specific de novo genes are involved in diseases such as cancer. NYCM, a de novo gene unique to humans and chimpanzees, regulates the pathogenesis of neuroblastomas in mouse models,[155] and the primate-specific PART1, an lncRNA gene, has been identified as both a tumor suppressor and an oncogene in different contexts.[37][156][157] Several other human- or primate-specific de novo genes, including PBOV1,[158] GR6,[159][160] MYEOV,[161] ELFN1-AS1,[162] and CLLU1,[38] are also linked to cancer. Some have even suggested considering tumor-specifically expressed, evolutionary novel genes as their own class of genetic elements, noting that many such genes are under positive selection and may be neofunctionalized in the context of tumors.[162]
The specific expression of many de novo genes in the human brain[57] also raises the intriguing possibility that de novo genes influence human cognitive traits. One such example is FLJ33706, a de novo gene that was identified in GWAS and linkage analyses for nicotine addiction and shows elevated expression in the brains of Alzheimer's patients.[163] Generally speaking, expression of young, primate-specific genes is enriched in the fetal human brain relative to the expression of similarly young genes in the mouse brain.[164] Most of these young genes, several of which originated de novo, are expressed in the neocortex, which is thought to be responsible for many aspects of human-specific cognition. Many of these young genes show signatures of positive selection, and functional annotations indicate that they are involved in diverse molecular processes, but are enriched for transcription factors.[164]
In addition to their roles in cancer processes, de novo originated human genes have been implicated in the maintenance of pluripotency[165] and in immune function.[37][136][166] The preferential expression of de novo genes in the testes is also suggestive of a role in reproduction. Given that the function of many de novo human genes remains uncharacterized, it seems likely that an appreciation of their contribution to human health and development will continue to grow.
Genome-scale studies of orphan and de novo genes in various lineages.
| Organism/Lineage | Homology Detection Method(s) | Evidence of Expression? | Evidence of Selection? | Evidence of Physiological Role? | # Orphan/De Novo Genes | Notes | Ref. |
|---|---|---|---|---|---|---|---|
| Arthropods | BLASTP for all 30 species against each other, TBLASTN for Formicidae only, searched by synteny for unannotated orthologs in Formicidae only | ESTs, RNA-seq; RT-PCR on select candidates | 37 Formicidae-restricted orthologs appear under positive selection (M1a to M2a and M7 to M8 models using likelihood ratio tests); as a group, Formicidae-restricted orthologs have a significantly higher Ka/Ks rate than non-restricted orthologs | Prediction of signal peptides and subcellular localization for subset of orphans | ~65,000 orphan genes across 30 species | Abundance of orphan genes dependent on time since emergence from common ancestor; >40% of orphans from intergenic matches indicating possible de novo origin | [75] |
| Arabidopsis thaliana | BLASTP against 62 species, PSI-BLAST against NCBI nonredundant protein database, TBLASTN against PlantGDB-assembled unique transcripts database; searched syntenic region of two closely related species | Transcriptomic and translatomic data from multiple sources | Allele frequencies of de novo genes correlated with their DNA methylation levels | None | 782 de novo genes | Also assessed DNA methylation and histone modifications | [53] |
| Bombyx mori | BLASTP against four lepidopterans, TBLASTN against lepidopteran EST sequences, BLASTP against NCBI nonredundant protein database | Microarray, RT-PCR | None | RNAi on five de novo genes produced no visible phenotypes | 738 orphan genes | Five orphans identified as de novo genes | [93] |
| Brassicaceae | BLASTP against NCBI nonredundant protein database, TBLASTN against NCBI nucleotide database, TBLASTN against NCBI EST database, PSI-BLAST against NCBI nonredundant protein database, InterProScan[167] | Microarray | None | TRGs enriched for expression changes in response to abiotic stresses compared to other genes | 1761 nuclear TRGs; 28 mitochondrial TRGs | ~2% of TRGs thought to be de novo genes | [94] |
| Drosophila melanogaster | BLASTN of query cDNAs against D. melanogaster, D. simulans and D. yakuba genomes; also performed check of syntenic region in sister species | cDNA/ expressed sequence tags (ESTs) | Ka/Ks ratios calculated between retained new genes and their parental genes are significantly >1, indicating most new genes are functionally constrained | List includes several genes with characterized molecular roles | 72 orphan genes; 2 de novo genes | Gene duplication dominant mechanism for new genes; 7/59 orphans specific to D. melanogaster species complex identified as de novo | [56] |
| Drosophila melanogaster | Presence or absence of orthologs in other Drosophila species inferred by synteny based on UCSC genome alignments and FlyBase protein-based synteny; TBLASTN against Drosophila subgroup | Indirect (RNAi) | Youngest essential genes show signatures of positive selection (α=0.25 as a group) | Knockdown with constitutive RNAi lethal for 59 TRGs | 195 “young” (>35myo) TRGs; 16 de novo genes | Gene duplication dominant mechanism for new genes | [54] |
| Drosophila melanogaster | RNA-seq in D. melanogaster and close relatives; syntenic alignments with D. simulans and D. yakuba; BLASTP against NCBI nonredundant protein database | RNA-seq | Nucleotide diversity lower in non-expressing relatives; Hudson-Kreitman-Aguade-like statistic lower in fixed de novo genes than in intergenic regions | Structural features of de novo genes (e.g. enrichment of long ORFs) suggestive of function | 106 fixed and 142 segregating de novo genes | Specifically expressed in testes | [55] |
| Drosophila | All-vs-all BLASTP; phylostratigraphic analysis | Annotated proteomes of twelve Drosophila species | Pairwise dN/dS values for all single exon focal ORFs | None | 6297 orphan genes; 2467 de novo genes | [67] | |
Homo sapiens
|
BLASTP against other primates; BLAT against chimpanzee and orangutan genomes, manual check of syntenic regions in chimpanzee and orangutan | RNA-seq | Substitution rate provides some evidence for weak selection; 59/60 de novo genes are fixed | None | 60 de novo genes | Enabling mutations identified; highest expression seen in brain and testes | [57] |
| Homo sapiens | BLASTP against chimpanzee, BLAT and Search of syntenic region in chimpanzee, manual check of syntenic regions in chimpanzee and macaque | EST/cDNA | No evidence of selective constraint seen by nucleotide divergence | One of the genes identified has a known role in leukemia | 3 de novo genes | Estimated that human genome contains ~ 18 human-specific de novo genes | [38] |
| Homo sapiens and five other primates | BLAST against each of the six primate transcriptomes; analysis of syntenic regions | Transcriptome data from up to six tissue types | Pairwise dN/dS ratios for human-chimpanzee homologs indicate mostly neutral or mild purifying selection | None | 29.751 transcribed novel human ORFs in total: 2,749 human-restricted, 5,378 primate-restricted | Novel ORF gain and loss found to be mainly stochastic rather than shaped by selection | [69] |
| Lachancea and Saccharomyces | BLASTP of all focal species against each other, BLASTP against NCBI nonredundant protein database, PSI-BLAST against NCBI nonredundant protein database, HMM Profile-Profile of TRG families against each other; families then merged and searched against four profile databases | Mass Spectrometry (MS) | Ka/Ks ratios across Saccharomyces indicate that candidates are under weak selection that increases with gene age; in Lachancea species with multiple strains, pN/pS ratios are lower for de novo candidates than for "spurious TRGs" | None | 288 candidate de novo TRGs in Saccharomyces, 415 in Lachancea | MS evidence of translation for 25 candidates | [96] |
| Mus musculus and Rattus norvegicus | BLASTP of rat and mouse against each other, BLASTP against Ensembl compara database; searched syntenic regions in rat and mouse | UniGene Database | Subset of genes shows low nucleotide diversity and high ORF conservation across 17 strains | Two mouse genes cause morbidity when knocked out | 69 de novo genes in mouse and 6 "de novo" genes in rat | Enabling mutations identified for 9 mouse genes | [168] |
| Mus musculus | BLASTP against NCBI nonredundant protein database | Microarray | None | None | 781 orphan genes | Age-dependent features of genes compatible with de novo emergence of many orphans | [70] |
| Mus musculus and four other mammals | DIAMOND and BLASTP; phylostratigraphy analysis | High coverage transcriptomes | None | None | ~60,000 transcribed novel ORFs | [71] | |
| Oryza | Protein-to-protein and nucleotide-to-nucleotide BLAT against eight Oryza species and two outgroup species; searched syntenic regions of these species for coding potential | RNA-seq (all de novo TRGs); Ribosome Profiling and targeted MS (some de novo TRGs) | 22 de novo candidates appear under negative selection, and six under positive selection, as measured by Ka/Ks rate | Expression of de novo TRGs is tissue-specific | 175 de novo TRGs | ~57% of de novo genes have translational evidence; transcription predates coding potential in most cases | [169] |
| Primates | BLASTP against 15 eukaryotes, BLASTN against human genome, analysis of syntenic regions | ESTs | Ka/Ks ratios for TRGs below one but higher than established genes; coding scores consistent with translated proteins | Several genes have well-characterized cellular roles | 270 TRGs | ~5.5% of TRGs estimated to have originated de novo | [37] |
| Pristionchus pacificus | BLASTP and tBLASTN, syntenic analysis | RNA-Seq | 2 cases complete de novo gene origination | 27 other high-confidence orphans whose methods of origin included annotation artifacts, chimeric origin, alternative reading frame usage, and gene splitting with subsequent gain of de novo exons | [170] | ||
| Rodentia | BLASTP against NCBI nonredundant protein database | None | Mouse genes share 50% identity with rat ortholog | None | 84 TRGs | Species-specific genes excluded from analysis; results robust to evolutionary rate | [110] |
| Saccharomyces cerevisiae | BLASTP and PSI-BLAST against 18 fungal species, HMMER and HHpred against several databases, TBLASTN against three close relatives | None | None | Majority of orphans have characterized fitness effects | 188 orphan genes | Ages of genes determined at level of individual residues | [89] |
| Saccharomyces cerevisiae | BLASTP, TBLASTX, and TBLASTN against 14 other yeast species, BLASTP against NCBI nonredundant protein database | Ribosome Profiling | All 25 de novo genes, 115 proto-genes under purifying selection (pN/pS < 1) | None | 25 de novo genes; 1,891 “proto-genes” | De novo gene birth more common than new genes from duplication; proto-genes are unique to Saccharomyces ( Sensu stricto ) yeasts
|
[59] |
| Saccharomyces cerevisiae | BLASTN, TBLASTX, against nt/nr, manual inspection of syntenic alignment | transcripts believed to be non-coding, manual inspection of ribosome profiling traces | None | None | 1 de novo candidate gene, 217 ribosome-associated transcripts | Candidate de novo gene is polymorphic. Ribosomal profiling data is the same as in [59] | [122] |
| Saccharomyces paradoxus | Intergenic ORFs (iORFs) were annotated in orthologous intergenic regions identified by microsynteny; BLASTP against NCBI refseq protein database against 417 species, including 237 fungi species. | Ribo-seq, RNA-seq
In vivo translation assay for subset (45) of translated iORFS |
Global dn/ds ratio for translated iORFs in 24 S. paradoxus strains showed no evidence for purifying selection. | None | 447 iORFs with significant translation that were specific to 3 S. paradoxus lineages and 1 S. cerevisiae lineage. | iORF translation efficiency was generally lower than that for annotated genes with significant overlap; only ~2% of the 19,689 iORFs found showed significant translation, but they add up to >8% of the canonical proteome in wild yeast populations. | [68] |
| Saccharomyces sensu stricto | BLASTP against NCBI nonredundant protein database, TBLASTN against ten outgroup species; BLASTP and phmmer against 20 yeast species reannotated using syntenic alignments | Transcript isoform sequencing (TIF-seq), Ribosome Profiling | Most genes weakly constrained but a subset under strong selection, according to Neutrality Index, Direction of Selection, Ka/Ks, and McDonald-Kreitman tests | Subcellular localization demonstrated for five genes | ~13,000 de novo genes | >65% of de novo genes are isoforms of ancient genes; >97% from TIF-seq dataset | [52] |
Note: For purposes of this table, genes are defined as orphan genes (when species-specific) or TRGs (when limited to a closely related group of species) when the mechanism of origination has not been investigated, and as de novo genes when de novo origination has been inferred, irrespective of method of inference. The designation of de novo genes as “candidates” or “proto-genes” reflects the language used by the authors of the respective studies.
See also
References
![]() This article was adapted from the following source under a CC BY 4.0 license (2019) (reviewer reports):
Stephen Branden Van Oss; Anne-Ruxandra Carvunis (23 May 2019). "De novo gene birth".
This article was adapted from the following source under a CC BY 4.0 license (2019) (reviewer reports):
Stephen Branden Van Oss; Anne-Ruxandra Carvunis (23 May 2019). "De novo gene birth". {{cite journal}}: CS1 maint: unflagged free DOI (link
- ^ S2CID 33999892.
- PMID 15064762.
- PMID 31619796.
- PMID 28163910.
- ^ PMID 25773713.
- ^ PMID 20651121.
- ^ S2CID 29756896.
- PMID 31120894.
- PMID 19716618.
- ^ S2CID 31738556.
- ^ Ohno S (1970) Evolution by Gene Duplication Allen & Unwin; Springer-Verlag
- S2CID 29552265.
- ^ Grassé P-P (1977) Evolution of living organisms: evidence for a new theory of transformation Academic Press
- S2CID 4264796.
- S2CID 4218777.
- S2CID 4206886.
- PMID 1329098.
- PMID 6585807.
- PMID 22821011.
- PMID 17939861.
- PMID 29083303.
- ^ PMID 32138667.
- PMID 8073532.
- PMID 17709368.
- ^ PMID 1822278.
- S2CID 4355476.
- S2CID 4271784.
- ^ PMID 8763498.
- ^ PMID 17435230.
- ^ PMID 16777968.
- ^ PMID 16361246.
- ^ PMID 26323763.
- ^ PMID 18493065.
- ^ PMID 29033289.
- ^ PMID 19154206.
- ^ S2CID 12446879.
- ^ PMID 19064677.
- ^ PMID 19726446.
- ^ PMID 18029048.
- ^ PMID 26116928.
- S2CID 14441902.
- ^ S2CID 6033249.[permanent dead link]
- PMID 16151190.
- PMID 17408474.
- PMID 26758516.
- PMID 25312911.
- ^ PMID 28087778.
- PMID 24932010.
- S2CID 2644451.
- PMID 29382321.
- PMID 11157786.
- ^ PMID 28981695.
- ^ PMID 27401176.
- ^ S2CID 7899890.
- ^ PMID 24457212.
- ^ PMID 18550802.
- ^ PMID 22102831.
- S2CID 52942639.
- ^ PMID 22722833.
- PMID 24814287.
- ^ PMID 24753594.
- ^ PMID 31674305.
- PMID 26032716.
- doi:10.1101/510214.
- PMID 26720152.
- PMID 6938980.
- ^ PMID 32253450.
- ^ PMID 31152050.
- ^ PMID 33210146.
- ^ PMID 23433480.
- ^ S2CID 52181376.
- ^ PMID 32066524.
- ^ PMID 24554240.
- ^ PMID 30232197.
- ^ PMID 23348040.
- ^ PMID 32499660.
- PMID 26836309.
- PMID 22844254.
- ^ PMID 24146629.
- ^ PMID 28104747.
- ^ PMID 33712569.
- PMID 32589737.
- ^ PMID 30765531.
- ^ PMID 29216381.
- PMID 31436535.
- PMID 20195295.
- PMID 25452167.
- ^ PMID 36099311.
- ^ PMID 19944701.
- PMID 14525923.
- PMID 18273382.
- PMID 19351897.
- ^ PMID 26296317.
- ^ PMID 21332978.
- ^ PMID 30232198.
- ^ PMID 29220506.
- PMID 30692195.
- ^ PMID 33416492.
- ^ S2CID 53216993.
- PMID 30065088.
- ^ PMID 30075701.
- doi:10.1101/332825.
- ^ PMID 37425675, retrieved 2023-12-25
- PMID 31227545.
- ^ PMID 32034123.
- PMID 35668555.
- ^ PMID 28355220.
- PMID 25736992.
- PMID 25843649.
- ^ PMID 28642936.
- PMID 25708804.
- PMID 33567396.
- PMID 35900020.
- S2CID 84338859.
- PMID 16569694.
- PMID 21771634.
- PMID 18451266.
- PMID 21765801.
- ^ PMID 25159147.
- S2CID 4959952.
- PMID 25233276.
- ^ PMID 21948395.
- PMID 32139545.
- ^ PMID 23593031.
- S2CID 4373304.
- ^ PMID 34810219.
- ^ S2CID 254966620.
- PMID 19043537.
- PMID 31545792.
- S2CID 25696990.
- S2CID 4423768.
- PMID 16388004.
- PMID 34478447.
- PMID 34791207.
- S2CID 198249413.
- ^ PMID 28854603.
- S2CID 14318566.
- PMID 21555408.
- S2CID 949694.
- ^ PMID 21199946.
- PMID 16387877.
- doi:10.1101/287193.
- PMID 30026186.
- PMID 24056411.
- PMID 17099057.
- ^ PMID 26517896.
- PMID 19679754.
- S2CID 812377.
- PMID 26002864.
- PMID 30247639.
- PMID 29802682.
- PMID 21115821.
- PMID 24050177.
- PMID 23949544.
- PMID 24391509.
- PMID 10706094.
- PMID 30049286.
- PMID 23418531.
- PMID 27056411.
- PMID 9307271.
- PMID 26568894.
- ^ PMID 27437030.
- PMID 20376170.
- ^ PMID 22028629.
- S2CID 205240839.
- PMID 9892612.
- PMID 18940856.
- PMID 23185269.
- S2CID 73728579.
- PMID 31088903.
