Pseudogene

Pseudogenes are nonfunctional segments of
Most non-bacterial genomes contain many pseudogenes, often as many as functional genes. This is not surprising, since various biological processes are expected to accidentally create pseudogenes, and there are no specialized mechanisms to remove them from genomes. Eventually pseudogenes may be deleted from their genomes by chance of DNA replication or DNA repair errors, or they may accumulate so many mutational changes that they are no longer recognizable as former genes. Analysis of these degeneration events helps clarify the effects of non-selective processes in genomes.
Pseudogene sequences may be transcribed into RNA at low levels, due to promoter elements inherited from the ancestral gene or arising by new mutations. Although most of these transcripts will have no more functional significance than chance transcripts from other parts of the genome, some have given rise to beneficial regulatory RNAs and new proteins.
Properties
This section is missing information about faster accumulation of mutation. (December 2023) |
Pseudogenes are usually characterized by a combination of similarity to a known gene and loss of some functionality. That is, although every pseudogene has a DNA sequence that is similar to some functional gene, they are usually unable to produce functional final protein products.[1] Pseudogenes are sometimes difficult to identify and characterize in genomes, because the two requirements of similarity and loss of functionality are usually implied through sequence alignments rather than biologically proven.
- Homology is implied by sequence similarity between the DNA sequences of the pseudogene and a known gene. After aligning the two sequences, the percentage of identical base pairs is computed. A high sequence identity means that it is highly likely that these two sequences diverged from a common ancestral sequence (are homologous), and highly unlikely that these two sequences have evolved independently (see Convergent evolution).
- Nonfunctionality can manifest itself in many ways. Normally, a gene must go through several steps to a fully functional protein: pre-mRNA processing, translation, and protein folding are all required parts of this process. If any of these steps fails, then the sequence may be considered nonfunctional. In high-throughput pseudogene identification, the most commonly identified disablements are premature stop codons and frameshifts, which almost universally prevent the translation of a functional protein product.
Pseudogenes for RNA genes are usually more difficult to discover as they do not need to be translated and thus do not have "reading frames". A number of rRNA pseudogenes have been identified on the basis of changes in rDNA array ends.[2]
Pseudogenes can complicate molecular genetic studies. For example, amplification of a gene by PCR may simultaneously amplify a pseudogene that shares similar sequences. This is known as PCR bias or amplification bias. Similarly, pseudogenes are sometimes annotated as genes in genome sequences.
Processed pseudogenes often pose a problem for gene prediction programs, often being misidentified as real genes or exons. It has been proposed that the identification of processed pseudogenes can help improve the accuracy of gene prediction methods.[3]
In 2014, 140 human pseudogenes have been shown to be translated.[4] However, the function, if any, of the protein products is unknown.
Types and origin
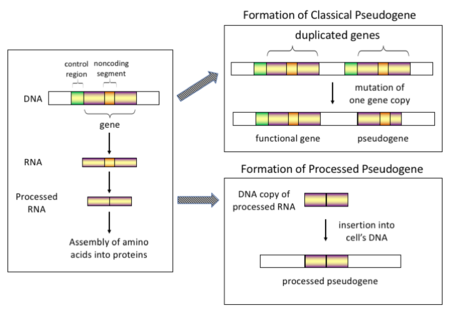
There are four main types of pseudogenes, all with distinct mechanisms of origin and characteristic features. The classifications of pseudogenes are as follows:
Processed

In higher
It has been shown that processed pseudogenes accumulate mutations faster than non-processed pseudogenes.[15]
Non-processed (duplicated)

Unitary pseudogenes

Various mutations (such as
Examples of pseudogene function
While the vast majority of pseudogenes have lost their function, some cases have emerged in which a pseudogene either re-gained its original or a similar function or evolved a new function. In the human genome, a number of examples have been identified that were originally classified as pseudogenes but later discovered to have a functional, although not necessarily protein-coding, role.[25][26]
Examples include the following:
Protein-coding: "pseudo-pseudogenes"

The rapid proliferation of DNA sequencing technologies has led to the identification of many apparent pseudogenes using gene prediction techniques. Pseudogenes are often identified by the appearance of a premature stop codon in a predicted mRNA sequence, which would, in theory, prevent synthesis (translation) of the normal protein product of the original gene. There have been some reports of translational readthrough of such premature stop codons in mammals. As alluded to in the figure above, a small amount of the protein product of such readthrough may still be recognizable and function at some level. If so, the pseudogene can be subject to natural selection. That appears to have happened during the evolution of Drosophila species.
In 2016 it was reported that four predicted pseudogenes in multiple Drosophila species actually encode proteins with biologically important functions,
As of 2012, it appeared that there are approximately 12,000–14,000 pseudogenes in the human genome.[31] A 2016 proteogenomics analysis using mass spectrometry of peptides identified at least 19,262 human proteins produced from 16,271 genes or clusters of genes, with 8 new protein-coding genes identified that were previously considered pseudogenes.[32] An earlier analysis found that human PGAM4 (phosphoglycerate mutase),[33] previously thought to be a pseudogene, is not only functional, but also causes infertility if mutated.[34][35]
A number of pseudo-pseudogenes were also found in prokaryotes, where some stop codon substitutions in essential genes appear to be retained, even positively selected for.[36][37]
Non-protein-coding
siRNAs. Some endogenous
piRNAs. Some
PTEN. The
Potogenes. Pseudogenes can, over evolutionary time scales, participate in gene conversion and other mutational events that may give rise to new or newly functional genes. This has led to the concept that pseudogenes could be viewed as potogenes: potential genes for evolutionary diversification.[46]
Bacterial pseudogenes
Pseudogenes are found in
Since most bacteria that carry pseudogenes are either symbionts or obligate intracellular parasites, genome size eventually reduces. An extreme example is the genome of Mycobacterium leprae, an obligate parasite and the causative agent of leprosy. It has been reported to have 1,133 pseudogenes which give rise to approximately 50% of its transcriptome.[48] The effect of pseudogenes and genome reduction can be further seen when compared to Mycobacterium marinum, a pathogen from the same family. Mycobacteirum marinum has a larger genome compared to Mycobacterium leprae because it can survive outside the host; therefore, the genome must contain the genes needed to do so.[49]
Although genome reduction focuses on what genes are not needed by getting rid of pseudogenes, selective pressures from the host can sway what is kept. In the case of a symbiont from the Verrucomicrobiota phylum, there are seven additional copies of the gene coding the mandelalide pathway.[50] The host, species from Lissoclinum, use mandelalides as part of its defense mechanism.[50]
The relationship between epistasis and the domino theory of gene loss was observed in Buchnera aphidicola. The domino theory suggests that if one gene of a cellular process becomes inactivated, then selection in other genes involved relaxes, leading to gene loss.[48] When comparing Buchnera aphidicola and Escherichia coli, it was found that positive epistasis furthers gene loss while negative epistasis hinders it.

This article is missing information about archaea. (December 2023) |
See also
- List of disabled human pseudogenes
- Molecular evolution
- Molecular paleontology
- Pseudogene (database)
- Retroposon
- Retrotransposon
References
- S2CID 42204036.
- PMID 28204512.
- PMID 16651666.
- PMID 24870542.
- ^ Max EE (1986). "Plagiarized Errors and Molecular Genetics". Creation Evolution Journal. 6 (3): 34–46.
- ^ Chandrasekaran C, Betrán E (2008). "Origins of new genes and pseudogenes". Nature Education. 1 (1): 181.
- PMID 15531153.
- S2CID 25083962.
- S2CID 32151696.
- S2CID 22437436.
- PMID 18842134.
- PMID 12468100.
- PMID 26224704.
- PMID 23359205.
- PMID 17568002.
- ^ Max EE (2003-05-05). "Plagiarized Errors and Molecular Genetics". TalkOrigins Archive. Retrieved 2008-07-22.
- ^ PMID 11073452.
- PMID 7705642.
- PMID 11779815.
- PMID 11827946.
- .
- PMID 1400507.
- PMID 8175804.
- PMID 16532395.
- S2CID 209393216.
- PMID 32421357.
- ^ PMID 27776356.
- S2CID 1665885.
- PMID 10958846.
- PMID 7682012.
- PMID 22951037.
- PMID 27250503.
- PMID 9370262.
- PMID 11961099.
- PMID 22590500.
- PMID 33672790.
- PMID 35489061.
- PMID 24823781.
- PMID 23376929.
- PMID 24279857.
- PMID 24178556.
- S2CID 5710813.
- PMID 25843629.
- PMID 9620558.
- PMID 20577206.
- PMID 14616058.
- PMID 25461580.
- ^ PMID 16237210.
- PMID 28854187.
- ^ PMID 29181447.
- S2CID 4307207.
Further reading
- Gerstein M, Zheng D (August 2006). "The real life of pseudogenes". Scientific American. 295 (2): 48–55. PMID 16866288.
- Torrents D, Suyama M, Zdobnov E, Bork P (December 2003). "A genome-wide survey of human pseudogenes". Genome Research. 13 (12): 2559–2567. PMID 14656963.
- Bischof JM, Chiang AP, Scheetz TE, Stone EM, Casavant TL, Sheffield VC, Braun TA (June 2006). "Genome-wide identification of pseudogenes capable of disease-causing gene conversion". Human Mutation. 27 (6): 545–552. S2CID 20219423.
- Syberg-Olsen MJ, Garber AI, Keeling PJ, McCutcheon JP, Husnik F (July 2022). "Pseudofinder: Detection of Pseudogenes in Prokaryotic Genomes". Molecular Biology and Evolution. 39 (7). PMID 35801562.
