Arabidopsis thaliana
| Arabidopsis thaliana | |
|---|---|

| |
| Scientific classification | |
| Kingdom: | Plantae |
| Clade: | Tracheophytes |
| Clade: | Angiosperms |
| Clade: | Eudicots |
| Clade: | Rosids |
| Order: | Brassicales |
| Family: | Brassicaceae |
| Genus: | Arabidopsis |
| Species: | A. thaliana
|
| Binomial name | |
| Arabidopsis thaliana (
Heynh. | |
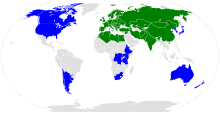
| |
The range of Arabidopsis thaliana.
| |
| Synonyms[1] | |
|
Arabis thaliana | |
Arabidopsis thaliana, the thale cress, mouse-ear cress or arabidopsis, is a small plant from the mustard family (Brassicaceae), native to Eurasia and Africa.[2][3][4][5][6][7] Commonly found along the shoulders of roads and in disturbed land, it is generally considered a weed.
A
Description

Arabidopsis thaliana is an

A. thaliana can complete its entire lifecycle in six weeks. The central stem that produces flowers grows after about 3 weeks, and the flowers naturally self-pollinate. In the lab, A. thaliana may be grown in Petri plates, pots, or hydroponics, under fluorescent lights or in a greenhouse.[15]
Taxonomy
The plant was first described in 1577 in the
Thousands of natural inbred accessions of A. thaliana have been collected from throughout its natural and introduced range.[16] These accessions exhibit considerable genetic and phenotypic variation, which can be used to study the adaptation of this species to different environments.[16]
Distribution and habitat
A. thaliana is native to Europe, Asia, and Africa, and its geographic distribution is rather continuous from the Mediterranean to Scandinavia and Spain to Greece.[17] It also appears to be native in tropical alpine ecosystems in Africa and perhaps South Africa.[18][19] It has been introduced and naturalized worldwide,[20] including in North America around the 17th century.[21]
A. thaliana readily grows and often pioneers rocky, sandy, and calcareous soils. It is generally considered a weed, due to its widespread distribution in agricultural fields, roadsides, railway lines, waste ground, and other disturbed habitats,[20][22] but due to its limited competitive ability and small size, it is not categorized as a noxious weed.[23] Like most Brassicaceae species, A. thaliana is edible by humans in a salad or cooked, but it does not enjoy widespread use as a spring vegetable.[24]
Use as a model organism
Botanists and biologists began to research A. thaliana in the early 1900s, and the first systematic description of mutants was done around 1945. Although A. thaliana the plant has little direct significance for agriculture, A. thaliana the model organism has revolutionized our understanding of the genetic, cellular, and molecular biology of flowering plants.

The first mutant in A. thaliana was documented in 1873 by
In the 1950s and 1960s, John Langridge and George Rédei played an important role in establishing A. thaliana as a useful organism for biological laboratory experiments. Rédei wrote several scholarly reviews instrumental in introducing the model to the scientific community. The start of the A. thaliana research community dates to a newsletter called Arabidopsis Information Service,[31] established in 1964. The first International Arabidopsis Conference was held in 1965, in Göttingen, Germany.
In the 1980s, A. thaliana started to become widely used in plant research laboratories around the world. It was one of several candidates that included maize,
Genomics

Nuclear genome
Due to the small size of its
The genome encodes ~27,600 protein-coding genes and about 6,500 non-coding genes.[41] However, the Uniprot database lists 39,342 proteins in their Arabidopsis reference proteome.[42] Among the 27,600 protein-coding genes 25,402 (91.8%) are now annotated with "meaningful" product names,[43] although a large fraction of these proteins is likely only poorly understood and only known in general terms (e.g. as "DNA-binding protein without known specificity"). Uniprot lists more than 3,000 proteins as "uncharacterized" as part of the reference proteome.
Chloroplast genome
The plastome of A. thaliana is a 154,478 base-pair-long DNA molecule,[34] a size typically encountered in most flowering plants (see the list of sequenced plastomes). It comprises 136 genes coding for small subunit ribosomal proteins (rps, in yellow: see figure), large subunit ribosomal proteins (rpl, orange), hypothetical chloroplast open reading frame proteins (ycf, lemon), proteins involved in photosynthetic reactions (green) or in other functions (red), ribosomal RNAs (rrn, blue), and transfer RNAs (trn, black).[35]
Mitochondrial genome
The mitochondrial genome of A. thaliana is 367,808 base pairs long and contains 57 genes.[44] There are many repeated regions in the Arabidopsis mitochondrial genome. The largest repeats recombine regularly and isomerize the genome.[45] Like most plant mitochondrial genomes, the Arabidopsis mitochondrial genome exists as a complex arrangement of overlapping branched and linear molecules in vivo.[46]
Genetics
The A. thaliana gene knockout collections are a unique resource for plant biology made possible by the availability of high-throughput transformation and funding for genomics resources. The site of T-DNA insertions has been determined for over 300,000 independent transgenic lines, with the information and seeds accessible through online T-DNA databases.[49] Through these collections, insertional mutants are available for most genes in A. thaliana.
Characterized accessions and mutant lines of A. thaliana serve as experimental material in laboratory studies. The most commonly used background lines are Ler (Landsberg erecta), and Col, or Columbia.[50] Other background lines less-often cited in the scientific literature are Ws, or Wassilewskija, C24, Cvi, or Cape Verde Islands, Nossen, etc. (see for ex.[51]) Sets of closely related accessions named Col-0, Col-1, etc., have been obtained and characterized; in general, mutant lines are available through stock centers, of which best-known are the Nottingham Arabidopsis Stock Center-NASC[50] and the Arabidopsis Biological Resource Center-ABRC in Ohio, USA.[52] The Col-0 accession was selected by Rédei from within a (nonirradiated) population of seeds designated 'Landsberg' which he received from Laibach.[53] Columbia (named for the location of Rédei's former institution, University of Missouri-Columbia) was the reference accession sequenced in the Arabidopsis Genome Initiative. The Later (Landsberg erecta) line was selected by Rédei (because of its short stature) from a Landsberg population he had mutagenized with X-rays. As the Ler collection of mutants is derived from this initial line, Ler-0 does not correspond to the Landsberg accessions, which designated La-0, La-1, etc.
Trichome formation is initiated by the GLABROUS1 protein. Knockouts of the corresponding gene lead to glabrous plants. This phenotype has already been used in gene editing experiments and might be of interest as visual marker for plant research to improve gene editing methods such as CRISPR/Cas9.[54][55]
Non-Mendelian inheritance controversy
In 2005, scientists at Purdue University proposed that A. thaliana possessed an alternative to previously known mechanisms of DNA repair, producing an unusual pattern of inheritance, but the phenomenon observed (reversion of mutant copies of the HOTHEAD gene to a wild-type state) was later suggested to be an artifact because the mutants show increased outcrossing due to organ fusion.[56][57][58]
Lifecycle
The plant's small size and rapid lifecycle are also advantageous for research. Having specialized as a
Cellular biology
Arabidopsis is often the model for study of
DNA repair
The DNA of plants is vulnerable to ultraviolet light, and DNA repair mechanisms have evolved to avoid or repair genome damage caused by UV. Kaiser et al.[60] showed that in A. thaliana cyclobutane pyrimidine dimers (CPDs) induced by UV light can be repaired by expression of CPD photolyase.
Germination in lunar regolith
On May 12, 2022,
Development
Flower development
A. thaliana has been extensively studied as a model for flower development. The developing flower has four basic organs -
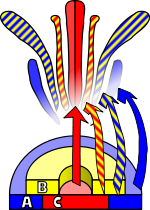

Observations of homeotic mutations led to the formulation of the
Leaf development
Studies of A. thaliana have provided considerable insights with regards to the genetics of leaf morphogenesis, particularly in dicotyledon-type plants.[63][64] Much of the understanding has come from analyzing mutants in leaf development, some of which were identified in the 1960s, but were not analysed with genetic and molecular techniques until the mid-1990s. A. thaliana leaves are well suited to studies of leaf development because they are relatively simple and stable.
Using A. thaliana, the genetics behind leaf shape development have become more clear and have been broken down into three stages: The initiation of the leaf primordium, the establishment of dorsiventrality, and the development of a marginal meristem. Leaf primordia are initiated by the suppression of the genes and proteins of class I KNOX family (such as SHOOT APICAL MERISTEMLESS). These class I KNOX proteins directly suppress gibberellin biosynthesis in the leaf primordium. Many genetic factors were found to be involved in the suppression of these class I KNOX genes in leaf primordia (such as ASYMMETRIC LEAVES1, BLADE-ON-PETIOLE1, SAWTOOTH1, etc.). Thus, with this suppression, the levels of gibberellin increase and leaf primordium initiate growth.
The establishment of leaf dorsiventrality is important since the dorsal (adaxial) surface of the leaf is different from the ventral (abaxial) surface.[65]
Microscopy
A. thaliana is well suited for
Physiology
Light sensing, light emission, and circadian biology
The photoreceptors
The
A. thaliana was used extensively in the study of the genetic basis of phototropism, chloroplast alignment, and stomal aperture and other blue light-influenced processes.[71] These traits respond to blue light, which is perceived by the phototropin light receptors. Arabidopsis has also been important in understanding the functions of another blue light receptor, cryptochrome, which is especially important for light entrainment to control the plants' circadian rhythms.[72] When the onset of darkness is unusually early, A. thaliana reduces its metabolism of starch by an amount that effectively requires division.[73]
Light responses were even found in roots, previously thought to be largely insensitive to light. While the gravitropic response of A. thaliana root organs is their predominant tropic response, specimens treated with mutagens and selected for the absence of gravitropic action showed negative phototropic response to blue or white light, and positive response to red light, indicating that the roots also show positive phototropism.[74]
In 2000, Dr.
Multiple efforts, including the Glowing Plant project, have sought to use A. thaliana to increase plant luminescence intensity towards commercially viable levels.
Thigmomorphogenesis (Touch response)
In 1990, Janet Braam and Ronald W. Davis determined that A. thaliana exhibits thigmomorphogenesis in response to wind, rain and touch.[76] Four or more touch induced genes in A. thaliana were found to be regulated by such stimuli.[76] In 2002, Massimo Pigliucci found that A. thaliana developed different patterns of branching in response to sustained exposure to wind, a display of phenotypic plasticity.[77]
On the Moon
On January 2, 2019, China's
Secondary metabolites
Thalianin is an Arabidopsis root triterpene.[80] Potter et al., 2018 finds synthesis is induced by a combination of at least 2 facts, cell-specific transcription factors (TFs) and the accessibility of the chromatin.[80]
Plant–pathogen interactions
Understanding how plants achieve resistance is important to protect the world's food production, and the agriculture industry. Many model systems have been developed to better understand interactions between plants and
| Pathogen type | Example in A. thaliana |
|---|---|
| Bacteria | Pseudomonas syringae, Xanthomonas campestris |
| Fungi | Colletotrichum destructivum, Botrytis cinerea, Golovinomyces orontii |
| Oomycete | Hyaloperonospora arabidopsidis |
| Viral | Cauliflower mosaic virus (CaMV), tobacco mosaic virus (TMV) |
| Nematode | Meloidogyne incognita, Heterodera schachtii
|

A schematic of PAMP-triggered immunity, to be specific recognition of flagellin by FLS2 (top left), effector-triggered immunity depicted through the recognition of avrRpt2 by RPS2 through RIN4 (top-right), microscopic view of callose deposition in an A. thaliana leaf (bottom left), an example of no hypersensitive response (HR), top, and HR in A. thaliana leaves (bottom right)

on the roots of Arabidopsis thaliana
a) Overview of an A. thaliana root (primary root) with numerous root hairs, b) Biofilm-forming bacteria, c) Fungal or oomycete hyphae surrounding the root surface, d) Primary root densely covered by spores and protists, e, f) Protists, most likely belonging to the Bacillariophyceae class, g) Bacteria and bacterial filaments, h, i) Different bacterial individuals showing great varieties of shapes and morphological features[81]
The use of A. thaliana has led to many breakthroughs in the advancement of knowledge of how plants manifest
In general, when a plant is exposed to a pathogen, or
The best-characterized PRR in A. thaliana is FLS2 (Flagellin-Sensing2), which recognizes bacterial
A second PRR, EF-Tu receptor (EFR), identified in A. thaliana, recognizes the bacterial
Both FLS2 and EFR use similar
PTI is able to combat pathogens in a nonspecific manner. A stronger and more specific response in plants is that of effector-triggered immunity (ETI), which is dependent upon the recognition of pathogen effectors, proteins secreted by the pathogen that alter functions in the host, by plant resistance genes (R-genes), often described as a gene-for-gene relationship. This recognition may occur directly or indirectly via a guardee protein in a hypothesis known as the guard hypothesis. The first R-gene cloned in A. thaliana was RPS2 (resistance to Pseudomonas syringae 2), which is responsible for recognition of the effector avrRpt2.[92] The bacterial effector avrRpt2 is delivered into A. thaliana via the Type III secretion system of P. syringae pv. tomato strain DC3000. Recognition of avrRpt2 by RPS2 occurs via the guardee protein RIN4, which is cleaved.[clarification needed] Recognition of a pathogen effector leads to a dramatic immune response known as the hypersensitive response, in which the infected plant cells undergo cell death to prevent the spread of the pathogen.[93]
Systemic acquired resistance (SAR) is another example of resistance that is better understood in plants because of research done in A. thaliana. Benzothiadiazol (BTH), a salicylic acid (SA) analog, has been used historically as an antifungal compound in crop plants. BTH, as well as SA, has been shown to induce SAR in plants. The initiation of the SAR pathway was first demonstrated in A. thaliana in which increased SA levels are recognized by nonexpresser of PR genes 1 (NPR1)[94] due to redox change in the cytosol, resulting in the reduction of NPR1. NPR1, which usually exists in a multiplex (oligomeric) state, becomes monomeric (a single unit) upon reduction.[95] When NPR1 becomes monomeric, it translocates to the nucleus, where it interacts with many TGA transcription factors, and is able to induce pathogen-related genes such as PR1.[96] Another example of SAR would be the research done with transgenic tobacco plants, which express bacterial salicylate hydroxylase, nahG gene, requires the accumulation of SA for its expression[97]
Although not directly immunological,
Evolutionary aspect of plant-pathogen resistance
Plants are affected by multiple
A. thaliana has also been used to study SAR.[101] This pathway uses benzothiadiazol, a chemical inducer, to induce transcription factors, mRNA, of SAR genes. This accumulation of transcription factors leads to inhibition of pathogen-related genes.[101]
Plant-pathogen interactions are important for an understanding of how plants have evolved to combat different types of pathogens that may affect them.
Research in A. thaliana suggests that the
Other research
Ongoing research on A. thaliana is being performed on the
Plant-on-a-chip devices in which A. thaliana tissues can be cultured in semi-in vitro conditions have been described.[107] Use of these devices may aid understanding of pollen-tube guidance and the mechanism of sexual reproduction in A. thaliana.
Researchers at the University of Florida were able to grow the plant in lunar soil originating from the Sea of Tranquillity.[108]
Self-pollination
A. thaliana is a predominantly self-pollinating plant with an outcrossing rate estimated at less than 0.3%.[109] An analysis of the genome-wide pattern of linkage disequilibrium suggested that self-pollination evolved roughly a million years ago or more.[110] Meioses that lead to self-pollination are unlikely to produce significant beneficial genetic variability. However, these meioses can provide the adaptive benefit of recombinational repair of DNA damages during formation of germ cells at each generation.[111] Such a benefit may have been sufficient to allow the long-term persistence of meioses even when followed by self-fertilization. A physical mechanism for self-pollination in A. thaliana is through pre-anthesis autogamy, such that fertilisation takes place largely before flower opening.
Databases and other resources
- TAIR and NASC:[50] curated sources for diverse genetic and molecular biology information, links to gene expression databases[112] etc.
- Arabidopsis Biological Resource Center (seed and DNA stocks)
- Nottingham Arabidopsis Stock Centre (seed and DNA stocks)
- Artade database
See also
- A. thaliana responses to salinity
- BZIP intron plant
- The Thaliana Bridge, installed in 2021 at Harlow Carr was inspired by the work of the botanical scientist Rachel Leech and represents the sequence of an Arabidopsis thaliana chromosome.[113]
- Novosphingobium arabidopsis, isolated from the rhizosphere of the plant
References
- from the original on 9 December 2018. Retrieved 1 June 2016.
- ^ "Arabidopsis thaliana". Germplasm Resources Information Network. Agricultural Research Service, United States Department of Agriculture. Retrieved 11 December 2017.
- S2CID 84959150.
- .
- S2CID 1788832.
- ^ PMID 25807084.
- PMID 28473417.
- ^ a b "Genome Assembly". The Arabidopsis Information Resource. Archived from the original on 7 March 2021. Retrieved 29 March 2016.
- ^ "Nifty 50: ARABIDOPSIS -- A PLANT GENOME PROJECT". www.nsf.gov. Retrieved 10 February 2023.
- ^ Flora of NW Europe: Arabidopsis thaliana Archived 8 December 2007 at the Wayback Machine
- ISBN 0-340-40170-2
- ^ Flora of Pakistan: Arabidopsis thaliana Archived 18 June 2008 at the Wayback Machine
- ^ Flora of China: Arabidopsis thaliana Archived 5 October 2018 at the Wayback Machine
- PMID 17313171.
- PMID 9784120.
- ^ PMID 27293186.)
{{cite journal}}: CS1 maint: numeric names: authors list (link - ^ "Arabidopsis thaliana (L.) Heynh". www.gbif.org. Archived from the original on 1 June 2019. Retrieved 8 December 2018.
- ^ Hedberg, Olov (1957). "Afroalpine Vascular Plants: A Taxonomic Revision". Acta Universitatis Upsaliensis: Symbolae Botanicae Upsalienses. 15 (1): 1–144.
- PMID 29862511.
- ^ a b "Arabidopsis thaliana – Overview". Encyclopedia of Life. Archived from the original on 10 June 2016. Retrieved 31 May 2016.
- PMID 29432421.
- ^ "Arabidopsis thaliana (thale cress)". Kew Gardens. Archived from the original on 28 February 2018. Retrieved 27 February 2018.
- ^ "State and Federal Noxious Weeds List | USDA PLANTS". plants.sc.egov.usda.gov. Archived from the original on 9 December 2018. Retrieved 8 December 2018.
- ^ "IRMNG". Encyclopedia of Life. Archived from the original on 1 April 2018.
- ^ [1] Archived 22 October 2016 at the Wayback Machine TAIR: About Arabidopsis
- PMID 15208410.
- (PDF) from the original on 9 July 2021. Retrieved 29 June 2021.
- PMID 20169178.
- S2CID 4323431.
- ^ PMID 11154286.
- ^ "About AIS". The Arabidopsis Information Resource. 8 November 2018. Archived from the original on 27 April 2021. Retrieved 25 April 2021.
- S2CID 22125701.
- PMID 2937058.
- ^ a b "Arabidopsis thaliana chloroplast, complete genome — NCBI accession number NC_000932.1". National Center for Biotechnology Information. Archived from the original on 4 November 2018. Retrieved 4 November 2018.
- ^ PMID 10574454.
- PMID 12646499.
- ^ (Leutwileret al., 1984). In our survey Arabidopsis ...
- PMID 25274549.
- PMID 11130711.
- ^ "TAIR - Genome Annotation". Archived from the original on 14 October 2008. Retrieved 29 December 2008.
- ^ "Details - Arabidopsis_thaliana - Ensembl Genomes 63". ensembl.gramene.org. Archived from the original on 24 June 2021. Retrieved 15 June 2021.
- ^ "Arabidopsis thaliana (Mouse-ear cress)". www.uniprot.org. Archived from the original on 21 May 2021. Retrieved 15 June 2021.
- S2CID 12155857.
- ^ "Arabidopsis thaliana ecotype Col-0 mitochondrion, complete genome — NCBI accession number BK010421". National Center for Biotechnology Information. 10 October 2018. Archived from the original on 12 April 2019. Retrieved 10 April 2019.
- PMID 7920724.
- PMID 24075874.
- S2CID 410286.
- S2CID 6906570.
- ^ "T-DNA Express: Arabidopsis Gene Mapping Tool". signal.salk.edu. Archived from the original on 25 November 2009. Retrieved 19 October 2009.
- ^ a b c "Eurasian Arabidopsis Stock Centre (uNASC)". arabidopsis.info. Archived from the original on 12 December 2001. Retrieved 19 October 2009.
- PMID 15908601.
- ^ "ABRC". abrc.osu.edu. Archived from the original on 25 February 2021. Retrieved 12 December 2020.
- ^ "NASC Collection Info". arabidopsis.info. Archived from the original on 19 July 2011. Retrieved 15 February 2011.
- PMID 28174584.
- PMID 29675030.
- S2CID 4420979.
- S2CID 82215542.
- PMID 27860488.
- ^ Kaiser G, Kleiner O, Beisswenger C, Batschauer A. Increased DNA repair in Arabidopsis plants overexpressing CPD photolyase. Planta. 2009 Aug;230(3):505-15. doi: 10.1007/s00425-009-0962-y. Epub 2009 Jun 12. PMID 19521716
- ^ Keeter, Bill (12 May 2022). "Scientists Grow Plants in Lunar Soil". NASA. Archived from the original on 14 May 2022. Retrieved 14 May 2022.
- S2CID 4276098.
- PMID 23864837.
- PMID 22303224.
- PMID 20435903.
- ^ Moreno N, Bougourd S, Haseloff J and Fiejo JA. 2006. Chapter 44: Imaging Plant Cells. In: Pawley JB (Editor). Handbook of Biological Confocal Microscopy - 3rd edition. SpringerScience+Business Media, New York. p769-787
- PMID 16441350.
- ^ PMID 11951029.
- S2CID 225237602.
- PMID 28693387.
- PMID 12921732.
- PMID 16096959.
- PMID 23805380.
- S2CID 28410755.
- ^ "Plants that Glow in the Dark" Archived 3 February 2014 at the Wayback Machine, Bioresearch Online, 18 May 2000
- ^ S2CID 38574940.
- S2CID 84173889.
- ^ a b Letzter, Rafi (4 January 2019). "There Are Plants and Animals on the Moon Now (Because of China)". Space.com. Archived from the original on 15 January 2019. Retrieved 15 January 2019.
- from the original on 12 January 2022. Retrieved 15 January 2019.
- ^ S2CID 219947907.
- doi:10.1186/s40168-018-0445-0.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License Archived 16 October 2017 at the Wayback Machine
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License Archived 16 October 2017 at the Wayback Machine - S2CID 4408024.
- from the original on 11 March 2022. Retrieved 5 September 2019.
- PMID 16473969.
- PMID 22303251.
- PMID 10377993.
- PMID 10911994.
- S2CID 90436989.
- S2CID 6856390.
- S2CID 7260214.,
- PMID 20713980.
- PMID 8400869.
- S2CID 1497625.
- PMID 12244227.
- S2CID 1562690.
- PMID 12897257.
- ^ S2CID 15507678.
- PMID 26243727.
- PMID 8091210.
- ^ S2CID 4332562.
- ^ PMID 8758979.
- PMID 25617321.
- S2CID 218617308.
- S2CID 215757180.
- PMID 25317938.
- PMID 20446867.
- S2CID 12989263.
- ^ "NASA-funded study breaks new ground in plant research". NASA. 12 May 2022. Archived from the original on 12 May 2022. Retrieved 13 May 2022.
- .
- S2CID 45853624.
- ^ Bernstein H; Byerly HC; Hopf FA; Michod RE (1985). "Genetic damage, mutation, and the evolution of sex". Science. 229 (4719): 1277–81. Bibcode:1985Sci...229.1277B. doi:10.1126/science.3898363. PMID 3898363
- ^ "TAIR - Gene Expression - Microarray - Public Datasets". Archived from the original on 4 December 2021. Retrieved 4 December 2021.
- ^ "Genome research inspires new bridge at Harlow Carr". The Garden (September 2021): 97. 2021.
External links
- Arabidopsis transcriptional regulatory map
- The Arabidopsis Information Resource (TAIR)
- Salk Institute Genomic Analysis Laboratory Archived 8 March 2021 at the Wayback Machine
- What Makes Plants Grow? The Arabidopsis genome knows Featured article in Genome News Network
- The Arabidopsis book - A comprehensive review published yearly related to research in Arabidopsis
- A. thaliana protein abundance
- The Arabidopsis Information Portal (Araport)
