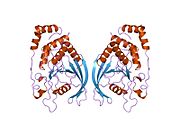
PTPRM
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) |
| ||||||||
| RefSeq (protein) |
| ||||||||
| Location (UCSC) | Chr 18: 7.57 – 8.41 Mb | Chr 17: 66.97 – 67.66 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Receptor-type tyrosine-protein phosphatase mu is an enzyme that in humans is encoded by the PTPRM gene.[5][6][7]
Function
The protein encoded by this gene is a member of the protein
Structure
Transmembrane PTPs are known as receptor
PTPmu possesses an extracellular region, a single transmembrane region, a 158 amino acid long juxtamembrane domain and two tandem tyrosine phosphatase domains (referred to as D1 and D2) in its intracellular domain, and thus represents an RPTP.
Homophilic binding
PTPmu protein expressed on the surface of cells is able to mediate binding between two cells, which results in the clustering of the cells, known as cell–cell aggregation.[33][34] PTPmu accomplishes this by interacting with another PTPmu molecule on an adjacent cell, known as homophilic binding. The Ig domain of PTPmu is responsible for promoting homophilic binding.[35] The Ig domain is also responsible for localizing PTPmu to the plasma membrane surface of the cell.[36] The ability of closely related molecules like PTPmu and PTPkappa to separate themselves to associate only with their identically matched (homologous) molecules, known as sorting, is attributed to the MAM domain.[37] The MAM, Ig, and the first two FNIII repeats are the minimum extracellular domains required for efficient cell–cell adhesion.[35][36][37][38][39][40][41] Crystallographic studies demonstrated that the MAM and Ig domains are tightly associated into one functional entity.[39] Additional crystal structure analysis by Aricescu and colleagues predicted that the adhesive interface between two PTPµ proteins is between the MAM and Ig domains of one PTPµ protein interacts with the first and second FN III domains of the second PTPµ protein.[40] The type IIb RPTPs mediate adhesion, with the exception of PCP-2.[42]
Tyrosine phosphatase activity
There are a number of ways that RPTP catalytic activity can be regulated (for reviews, see [11][14][17][43]). Dimerization of identical RPTP proteins at the cell surface leaves the PTP domains either in an open active conformation, as in the case of PTPmu[44] and LAR,[45] or in an inhibited conformation that leaves the catalytic domain inaccessible, in the case of CD45,[46] PTPalpha,[47] and PTPzeta/beta.[48] The binding of different parts of the protein with itself (ex. by folding to interact with itself), known as intramolecular interactions, can affect the activity of RPTPs. The cytoplasmic domains of different RPTPs can interact[49][50] to yield heterodimers of RPTP proteins, which then influence catalytic activity (for example, see [51]).
The regulation of PTPmu catalytic activity is complex. Like most RPTPs, the membrane proximal (or D1) phosphatase domain of PTPmu is catalytically active.
Crystal structure analysis of the D1 of PTPmu demonstrated that PTPmu dimers are in an open active conformation.[44] Even though PTPmu dimers may be active, an additional study suggests that the extracellular domain of PTPmu reduces phosphatase activity. In this study, it was shown that the cytoplasmic domain of PTPmu (a PTPmu molecule lacking the extracellular domain) has greater phosphatase activity than the full-length protein in an enzymatic phosphatase assay.[56]
PTPmu has a long juxtamembrane domain, which likely influences catalytic activity. The juxtamembrane domain of PTPmu can bind to either the D1 and/or D2 of PTPmu, but only within the same PTPmu monomer.[57] Removal of the juxtamembrane domain from PTPmu has been suggested to reduce PTPmu phosphatase activity.[52] The D2 domain of PTPmu also regulates its activity. Although originally demonstrated to positively regulate phosphatase activity,[52] the D2 domain has been shown to negatively affect PTPmu catalytic activity.[58] A wedge-shaped motif located by D1 also regulates catalytic activity.[59] Use of a peptide with the same sequence as the wedge motif inhibits PTPmu mediated functions.[59][60][61][62]
Certain stimuli may also influence PTP activity. For example, alteration of cell oxidation induces conformational changes in the cytoplasmic domain of PTPmu, which may affect its tyrosine phosphatase activity or binding of extracellular ligands.[55]
Cadherin-dependent adhesion
Classical
In addition to catenins and cadherins, PTPmu dephosphorylates PIPKIγ90 and nectin-3 (
Endothelial cell adhesion
PTPµ is expressed in human umbilical cord vein endothelial cells (HUVEC)[70] and in capillaries in the developing brain.[24] The expression of PTPµ in HUVEC cells increases at higher cell density.[70] Studies of PTPµ expression in animal tissues have demonstrated that PTPµ is preferentially expressed in endothelial cells of arteries and capillaries and in cardiac smooth muscle, in addition to brain cells.[25][26] Because of this specialized expression in arterial endothelial cells, and because PTPµ is found to associate with proteins involved in maintaining endothelial cell–cell junctions, such as VE-cadherin,[71] PTPµ is hypothesized to regulate endothelial cell junction formation or permeability. PTPµ has been shown to be involved in mechanotransduction that results from changes in blood flow to influence endothelial cell-mediated blood vessel dilation, a process induced by “shear stress.”[72] When PTPmu is missing in mice (PTPmu -/- knock-out mice), cannulated mesenteric arteries show reduced flow-induced (or “shear stress” induced) dilation.[72] PTPmu tyrosine phosphatase activity is activated by shear stress.[73] Caveolin 1 is a scaffolding protein enriched in endothelial cell junctions that is also linked to shear stress regulated responses.[73] Caveolin 1 is dephosphorylated on tyrosine 14 in response to shear stress and PTPmu is hypothesized to catalyze this reaction.[73]
Cell migration
Neurite outgrowth
PTPmu is expressed in the developing brain and retina.[27][28][29][30][31][74] A brain cell, or neuron, has a cell body that contains the nucleus and two types of extensions or processes that grow out from the cell body, the dendrites and axons. Dendrites generally receive input from other neurons, while axons send output to adjacent neurons. These processes are called neurites when grown ‘’in vitro’’ on tissue culture plates, because it is not clear whether they are dendrites or axons. ‘’In vitro’’ growth studies are useful for evaluating the mechanisms that neurons use to grow and function. A neurite outgrowth assay is a type of experiment where neurons are placed on different adhesive substrates on tissue culture plates. A neurite outgrowth assay is meant to mimic how neurons grow inside the body. During development of the nervous system, neuronal axons reach their often-distant targets by reacting to different substrates in their environment, so-called guidance cues, that are attractive, repulsive or simply permissive, meaning these substrates pull axons toward them, away from them, or act in a way that allows growth, respectively. When PTPmu is applied to a dish as an ‘’in vitro’’ substrate, it promotes neurite outgrowth.[27] PTPmu also acts as a guidance cue during development of the nervous system, by repelling neurites of the temporal neural retina, while permitting growth of neurites from the nasal neural retina.[28] Expression of PTPmu protein capable of dephosphorylating tyrosine residues is required for mediating both nasal neurite outgrowth and temporal neurite repulsion.[75] By blocking the expression of PTPmu protein with antisense technology, or by expressing catalytically inactive mutants of PTPmu (molecules of PTPmu that can not dephosphorylate their target proteins) in the developing retina, it was shown that PTPmu is required for the development of the neural retina.[29]
PTPmu also regulates neurite outgrowth on classical cadherins. PTPmu tyrosine phosphatase activity is necessary for neurite outgrowth on the classical cadherins E-, N- and R-cadherin,[27][60][61] suggesting that PTPmu dephosphorylates key components of the cadherin-catenin complex to regulate axonal migration. Again, this emphasizes that PTPmu likely regulates cadherin-dependent processes via its cytoplasmic domain.
Various signals required for PTPmu-mediated neurite outgrowth and repulsion have been identified. Some of these signals are proteins that interact with, or bind, to PTPmu, whereas, others may be dephosphorylated by PTPmu. PTPmu interacts with the scaffolding proteins RACK1/
The proteins PLCγ1 (PLCG1), PKCδ (PRKCD) and BCCIP are PTPmu substrates.[80] PKCδ activity is required for PTPmu mediated neurite outgrowth[81] and PTPmu-mediated neurite repulsion.[82] Expression of BCCIP is necessary for PTPmu-mediated neurite outgrowth.[83] PTPmu is cleaved in certain brain cancers, which results in nuclear translocation of the cytoplasmic domain of PTPmu (see below). A possible function for the BCCIP-PTPmu interaction may be to shuttle the intracellular PTPmu fragment into the cell nucleus. In summary, PTPmu dephosphorylates PKCδ, PLCγ1, and BCCIP, and binds to IQGAP1. The expression and/or activity of all these proteins and Cdc42 is necessary for PTPmu-mediated neurite outgrowth. Also, the activity of the GTPase Rac1 promotes PTPmu-mediated neurite repulsion.
Cancer
PTPmu is downregulated in
Interactions
PTPRM has been shown to
- BCCIP,[83]
- c-Met,[65]
- CDH4 R-cadherin (cadherin-4),[64]
- CDH5 VE-cadherin (cadherin 5, CDH5),[71]
- CTNND1 (p120catenin),[54]
- GNB2L1/RACK1,[76]
- GJA1 connexin43 (gap junction protein, alpha 1),[69]
- IQGAP1,[77]
- PVRL3 (nectin3),[68]
- PIPKIγ90,[68]
- PRKCD (PKCδ),[80] and
- PLCG1 (PLCγ1).[80]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000173482 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000033278 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ S2CID 7237197.
- PMID 8404049.
- ^ "Entrez Gene: PTPRM protein tyrosine phosphatase, receptor type, M".
- PMID 1339708.
- PMID 14731595.
- ^ a b c Brady-Kalnay S (1998). "Ig-superfamily phosphatases". In Peter Sonderegger (ed.). Ig Superfamily Molecules in the Nervous System (6 ed.). Zurich: Harwood Academic Publishers.
- ^ a b c Brady-Kalnay S (2001). "Protein tyrosine phosphatases". In Beckerle, M. (ed.). Cell Adhesion: Frontiers in Molecular Biology (39 ed.). Oxford, UK.: Oxford University Press. pp. 217–258.
- PMID 8573339.
- ISSN 0775-051X.
- ^ S2CID 44938812.
- PMID 12456340.
- PMID 12506125.
- ^ PMID 15464569.
- PMID 16497668.
- PMID 16497667.
- ^ PMID 21084269.
- ^ PMID 21235433.
- ^ PMID 7642713.
- S2CID 35492083.
- ^ PMID 8989520.
- ^ PMID 10094839.
- ^ PMID 12895029.
- ^ PMID 10087273.
- ^ PMID 11978837.
- ^ PMID 14623235.
- ^ S2CID 24084590.
- ^ PMID 10213455.
- PMID 10666036.
- PMID 8394372.
- PMID 8393854.
- ^ PMID 7961788.
- ^ PMID 15491993.
- ^ PMID 7782276.
- PMID 15084579.
- ^ PMID 16456543.
- ^ S2CID 15702183.
- PMID 18363557.
- PMID 20521994.
- PMID 10852814.
- ^ PMID 9346878.
- S2CID 14417598.
- PMID 9417031.
- S2CID 4233685.
- PMID 10706604.
- PMID 10777529.
- PMID 12376545.
- PMID 12364328.
- ^ PMID 7504951.
- ^ PMID 7559782.
- ^ PMID 10753936.
- ^ S2CID 199555986.
- S2CID 24662451.
- PMID 10809770.
- PMID 11162517.
- ^ PMID 16613844.
- ^ PMID 17276081.
- ^ PMID 20197094.
- ^ PMID 19690139.
- PMID 11801604.
- ^ PMID 9531566.
- ^ PMID 10425198.
- PMID 9772302.
- PMID 14691142.
- ^ PMID 17965016.
- ^ PMID 14681016.
- ^ PMID 8620001.
- ^ PMID 15793303.
- ^ S2CID 40391996.
- ^ PMID 16325778.
- S2CID 9560178.
- S2CID 3813261.
- ^ PMID 11278757.
- ^ PMID 16380380.
- PMID 12884260.
- ^ PMID 17234431.
- ^ PMID 20506511.
- S2CID 54361970.
- S2CID 54311542.
- ^ PMID 18773424.
- ^ PMID 19304959.
- ^ "NIH Researchers Identify Key Factor that Stimulates Brain Cancer Cells to Spread". News Release. National Institutes of Health (NIH). 2009-08-18. Retrieved 2011-07-21.
- S2CID 56680336.
- ^ Seper C (2009-08-18). "First, cure cancer. Second, build an iPhone app". MedCity News. Retrieved 2011-07-21.
- ^ PMID 20360941.
- PMID 1547787.
- PMID 1317540.
Further reading
- Serra-Pagès C, Medley QG, Tang M, Hart A, Streuli M (June 1998). "Liprins, a family of LAR transmembrane protein-tyrosine phosphatase-interacting proteins". J. Biol. Chem. 273 (25): 15611–20. PMID 9624153.
- Feiken E, van Etten I, Gebbink MF, Moolenaar WH, Zondag GC (May 2000). "Intramolecular interactions between the juxtamembrane domain and phosphatase domains of receptor protein-tyrosine phosphatase RPTPmu. Regulation of catalytic activity". J. Biol. Chem. 275 (20): 15350–6. PMID 10809770.