PTEN (gene)
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 10: 87.86 – 87.97 Mb | Chr 19: 32.73 – 32.8 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Phosphatase and tensin homolog (PTEN) is a phosphatase in humans and is encoded by the PTEN
PTEN acts as a tumor suppressor gene through the action of its phosphatase protein product. This phosphatase is involved in the regulation of the cell cycle, preventing cells from growing and dividing too rapidly.[8] It is a target of many anticancer drugs.
The protein encoded by this gene is a
Function
PTEN protein acts as a
PTEN also has weak protein
PTEN is one of the targets for drug candidates such as the oncomiR, MIRN21.
Structure
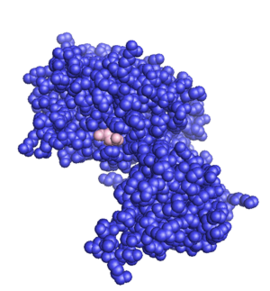
The structure of the core of PTEN (solved by X-ray crystallography, see figure to the upper right[5]) reveals that it consists primarily of a phosphatase domain, and a C2 domain: the phosphatase domain contains the active site, which carries out the enzymatic function of the protein, while the C2 domain binds the phospholipid membrane. Thus PTEN binds the membrane through both its phosphatase and C2 domains, bringing the active site to the membrane-bound PIP3 to dephosphorylate it.
The two domains of PTEN, a protein tyrosine phosphatase domain and a C2 domain, are inherited together as a single unit and thus constitute a superdomain, not only in PTEN but also in various other proteins in fungi, plants and animals, for example, tensin proteins and auxilin.[12]
The active site of PTEN consists of three loops, the
Not present in the crystal structure of PTEN is a short 10-amino-acid unstructured region N-terminal of the phosphatase domain (from residues 6 to 15), known variously as the PIP2 Binding Domain (PBD) or PIP2 Binding Motif (PBM)[13][14][15] This region increases PTEN's affinity for the plasma membrane by binding to Phosphatidylinositol 4,5-bisphosphate, or possibly any anionic lipid.
Also not present in the crystal structure is the intrinsically disordered C-terminal region (CTR) (spanning residues 353–403). The CTR is constitutively phosphorylated at various positions that effect various aspects of PTEN, including its ability to bind to lipid membranes, and also act as either a protein or lipid phosphatase.[16][17]
Additionally, PTEN can also be expressed as PTEN-L[18] (known as PTEN-Long, or PTEN-α[19]), a leucine initiator alternative start site variant, which adds an additional 173 amino acids to the N-terminus of PTEN. The exact role of this 173-amino acid extension is not yet known, either causing PTEN to be secreted from the cell, or to interact with the mitochondria. The N-terminal extension has been predicted to be largely disordered,[20] although there is evidence that there is some structure in the last twenty amino acids of the extension (most proximal to the start methionine of PTEN).[17]
Clinical significance
Cancer
PTEN is one of the most commonly lost tumor suppressors in human cancer; in fact, up to 70% of men with prostate cancer are estimated to have lost a copy of the PTEN gene at the time of diagnosis.[21] A number of studies have found increased frequency of PTEN loss in tumours which are more highly visible on diagnostic scans such as mpMRI, potentially reflecting increased proliferation and cell density in these tumours.[22]
During tumor development, mutations and deletions of PTEN occur that inactivate its enzymatic activity leading to increased cell proliferation and reduced cell death. Frequent genetic inactivation of PTEN occurs in glioblastoma, endometrial cancer, and prostate cancer; and reduced expression is found in many other tumor types such as lung and breast cancer. Furthermore, PTEN mutation also causes a variety of inherited predispositions to cancer.
Non-cancerous neoplasia
Researchers have identified more than 70
Mutations in the PTEN gene cause several other disorders that, like Cowden syndrome, are characterized by the development of non-cancerous tumors called
Brain function and autism
Defects of the PTEN gene have been cited to be a potential cause of
When defective, PTEN protein interacts with the protein of a second gene known as Tp53 to dampen energy production in neurons. This severe stress leads to a spike in harmful mitochondrial DNA changes and abnormal levels of energy production in the cerebellum and hippocampus, brain regions critical for social behavior and cognition. When PTEN protein is insufficient, its interaction with
Patients with defective PTEN can develop cerebellar mass lesions called dysplastic gangliocytomas or Lhermitte–Duclos disease.[23]
Cell regeneration
PTEN's strong link to cell growth inhibition is being studied as a possible therapeutic target in tissues that do not traditionally regenerate in mature animals, such as central neurons. PTEN deletion mutants have recently[26] been shown to allow nerve regeneration in mice.[27][28]
As a drug target
PTEN inhibitors
Cell lines
Cell lines with known PTEN mutations include:
Interactions
PTEN (gene) has been shown to
See also
References
- ^ a b c ENSG00000284792 GRCh38: Ensembl release 89: ENSG00000171862, ENSG00000284792 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000013663 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ S2CID 5624414.
- S2CID 41286105.
- ^ "OrthoMaM phylogenetic marker: PTEN coding sequence". Archived from the original on 2016-12-27. Retrieved 2009-12-02.
- ^ PMID 15448614.
- ^ "Entrez Gene: PTEN phosphatase and tensin homolog (mutated in multiple advanced cancers 1)".
- PMID 24814346.
- PMID 26399523.
- PMID 25694109.
- PMID 12857747.
- PMID 14764604.
- PMID 12534371.
- PMID 19114656.
- ^ PMID 26527737.
- PMID 23744781.
- PMID 24768297.
- PMID 24056727.
- PMID 16079851.
- PMID 33000006.
- ^ PMID 15121767.
- ^ PMID 22900024.
- ISBN 9780190681425.
- ^ "Rodent of the Week: Nerves regenerated after spinal cord injury". The Los Angeles Times. August 13, 2010.
- PMID 20694004.
- PMID 31453382.
- PMID 22253859.
- PMID 24882728.
- ^ S2CID 23093929.
- ^ S2CID 1093672.
- PMID 10748157.
- PMID 12177006.
- PMID 18498243.
- PMID 15205473.
- PMID 12620407.
- PMID 10400703.
- PMID 11857088.
Further reading
- Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, et al. (March 1997). "PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer". Science. 275 (5308): 1943–1947. S2CID 23093929.
- Simpson L, Parsons R (March 2001). "PTEN: life as a tumor suppressor". Experimental Cell Research. 264 (1): 29–41. PMID 11237521.
- Eng C (September 2003). "PTEN: one gene, many syndromes". Human Mutation. 22 (3): 183–198. S2CID 13417857.
- Hamada K, Sasaki T, Koni PA, Natsui M, Kishimoto H, Sasaki J, et al. (September 2005). "The PTEN/PI3K pathway governs normal vascular development and tumor angiogenesis". Genes & Development. 19 (17): 2054–2065. PMID 16107612.
- Leslie NR, Downes CP (August 2004). "PTEN function: how normal cells control it and tumour cells lose it". The Biochemical Journal. 382 (Pt 1): 1–11. PMID 15193142.
- Sansal I, Sellers WR (July 2004). "The biology and clinical relevance of the PTEN tumor suppressor pathway". Journal of Clinical Oncology. 22 (14): 2954–2963. PMID 15254063.
- Waite KA, Eng C (April 2002). "Protean PTEN: form and function". American Journal of Human Genetics. 70 (4): 829–844. PMID 11875759.
- Zhou XP, Waite KA, Pilarski R, Hampel H, Fernandez MJ, Bos C, et al. (August 2003). "Germline PTEN promoter mutations and deletions in Cowden/Bannayan-Riley-Ruvalcaba syndrome result in aberrant PTEN protein and dysregulation of the phosphoinositol-3-kinase/Akt pathway". American Journal of Human Genetics. 73 (2): 404–411. PMID 12844284.
- Ji SP, Zhang Y, Van Cleemput J, Jiang W, Liao M, Li L, et al. (March 2006). "Disruption of PTEN coupling with 5-HT2C receptors suppresses behavioral responses induced by drugs of abuse". Nature Medicine. 12 (3): 324–329. S2CID 22093776.
- Pulido R (May 2015). "PTEN: a yin-yang master regulator protein in health and disease". Methods. 77–78: 3–10. PMID 25843297.
- Pulido R (January 2018). "PTEN Inhibition in Human Disease Therapy". Molecules. 23 (2): 285. PMID 29385737.
External links
- GeneReviews/NCBI/NIH/UW entry on PTEN Hamartoma Tumor Syndrome (PHTS)
- PTEN+Protein at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- "PTEN Gene - phosphatase and tensin homolog". GeneCards. The Weizmann Institute of Science. Archived from the original on 2007-10-08. Retrieved 2009-03-12.
- "Gene overview of all published AD-association studies for PTEN". Alzforum: AlzGene. Alzheimer Research Forum. Archived from the original on 2009-02-10. Retrieved 2009-03-12.
- Research shows gene defect's role in autism-like behavior
- Dance Your PhD 2017 : A Story of Tumor Suppression Deepti Mathur. PTEN and cancer explained in dance. A metabolic pathway uses glutamine to create a component of DNA. This pathway is regulated in part by PTEN. Loss of PTEN allows the pathway to go into overdrive, leading to cancer. A drug that interrupts the PTEN pathway preferentially destroys cancer cells.
- PDBe-KB provides an overview of all the structure information available in the PDB for Human Phosphatidylinositol 3,4,5-trisphosphate 3-phosphatase and dual-specificity protein phosphatase PTEN
This article incorporates text from the United States National Library of Medicine, which is in the public domain.

