Type I topoisomerase
This article is missing information about pfam box for the actual catalytic domain; member of breaking-rejoining enzyme (BRE) superfamily; link between BRE and AraC homeobox per ECOD. (February 2021) |
| DNA topoisomerase I, N-terminal (non-catalytic), viral | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
In molecular biology Type I topoisomerases are
- Type IA topoisomerases change the linking number of a circular DNA strand by units of strictly 1.
- Type IB topoisomerases change the linking number by multiples of 1 (n).
Historically, type IA topoisomerases are referred to as prokaryotic topo I, while type IB topoisomerases are referred to as eukaryotic topoisomerase. This distinction, however, no longer applies as type IA and type IB topoisomerases exist in all domains of life.
Functionally, these subclasses perform very specialized functions.
Function
These
Structure
This domain assumes a beta(2)-alpha-beta-alpha-beta(2) fold, with a left-handed crossover between strands beta2 and beta3. It has a four criss-crossed beta-strands surrounded by four alpha-helices that are arranged in a Rossmann fold[3]
Mechanisms
Type I topoisomerases are
DNA topoisomerases regulate the number of topological links between two DNA strands (i.e. change the number of superhelical turns) by catalysing transient single- or double-strand breaks, crossing the strands through one another, then resealing the breaks.[4]
Classes
DNA topoisomerases are divided into two classes: type I
Type IA topoisomerases
Introduction
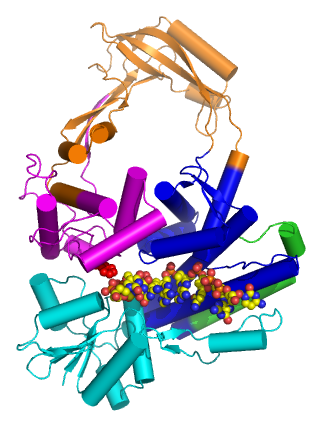
Type IA topoisomerases, historically said to be found in prokaryotes, create a single break in DNA and pass a second strand or duplex through the break. This strand passage mechanism shares several features with type IIA topoisomerases. They both form a 5' phosphotyrosine intermediate, and require a divalent metal ion to perform its work. Unlike type II topoisomerases, type IA topoisomerases do not use energy to do its work (with the notable exception of reverse gyrase, see below).
Structure
Type IA topoisomerases have several domains, often number Domain 1-4. Domain I contains a Toprim domain (a Rossman fold known to coordinate Magnesium ions), domain IV and domain III each consist of a helix-turn-helix (HTH) domain; the catalytic tyrosine resides on the HTH of domain III. Domain II is a flexible bridge between domains III and IV. The structure of type IA topoisomerase resembles a lock, with Domains I, III and IV lying on the bottom of the structure.[6] The structure of topo III (see below) bound to single-stranded DNA[7] (pdb id = 1I7D) shows how the HTH and Toprim domain are coordinated about the DNA.
Type IA topoisomerase variants
There are several variants of Type IA topoisomerases, differing by appendages attached to the main core (sometimes referred to as the "topo-fold"). Members of this subclass include topo I, topo III (which contain additional Zinc-binding motifs), and reverse gyrase. Reverse gyrase is particularly interesting because an ATPase domain, which resembles the helicase-like domain of the Rho transcription factor, is attached (the structure of reverse gyrase was solved by Rodriguez and Stock, EMBO J 2002). The enzyme uses the hydrolysis of ATP to introduce positive supercoils and overwinds DNA, a feature attractive in hyperthermophiles, in which reverse gyrase is known to exist. Rodriguez and Stock have done further work to identify a "latch" that is involved in communicating the hydrolysis of ATP to the introduction of positive supercoils.
The topo III variant is likewise very interesting because it has zinc-binding motifs that is thought to bind single-stranded DNA. Topo III has been identified to be associated with the BLM (for Bloom Syndrome) helicase during recombination.
Mechanism
Type IA topoisomerases operate through a strand-passage mechanism, using a single gate (in contrast with type II topoisomerases). First, the single-stranded DNA binds domain III and I. The catalytic tyrosine cleaves the DNA backbone, creating a transient 5' phosphotyrosine intermediate. The break is then separated, using domain II as a hinge, and a second duplex or strand of DNA is passed through. Domain III and I close and the DNA is re-annealed.
Type IB topoisomerases
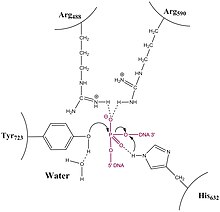

Introduction
In contrast to type IA topoisomerases, type 1B Topoisomerase solves the problem of overwound and underwound (also referred to as positively or negatively supercoiled) DNA through a hindered rotary mechanism. Crystal structures, biochemistry, and single molecule experiments have contributed to a general mechanism. The enzyme first wraps around DNA and creates a single, 3' phosphotyrosine intermediate. The 5' end is then free to rotate, twisting it about the other strand, to relax DNA until the topoisomerase re-ligates the broken strands.
Structure
The structure of topo IB bound to DNA has been solved (pdb id = 1A36). Topo IB is composed of an NTD, a capping lobe, a catalytic lobe, and a C-terminal domain. The capping lobe and catalytic lobe wrap around the DNA.
Mechanism
Relaxation is not an active process and energy (in the form of
Type IB topoisomerases were originally identified in eukaryotes and in viruses. Viral topo I is unique because it binds DNA in a sequence-specific manner.
See the article TOP1 for further details on this well-studied type 1B topoisomerase.
Type IC topoisomerases
A third type of topoisomerase I was identified, topo V, in the archaeon
Intermediates
All topoisomerases form a phosphotyrosine intermediate between the catalytic tyrosine of the enzyme and the scissile phosphoryl of the DNA backbone.
- Type IA topoisomerases form a covalent linkage between the catalytic tyrosine and the 5'-phosphoryl.
- Type IB enzymes form a covalent 3'-phosphotyrosine intermediate.
- Type IC topoisomerases form a covalent 3'-phosphotyrosine intermediate.
This intermediate is isoenergetic, meaning that the forward cleavage reaction and the backward religation reaction are both energetically equal. As such, no outside energy source is necessary to conduct this reaction.
Inhibition
As topoisomerases generate breaks in DNA, they are targets of small-molecule inhibitors that inhibit the enzyme. Type 1 topoisomerase is inhibited by
The human topoisomerase type IB enzyme forms a covalent 3'-phosphotyrosine intermediate, the topoisomerase 1-cleavage-complex (Top1cc). The active irinotecan metabolite, SN-38, acts by trapping (making a ternary complex with) a subset of Top1cc, those with a guanine +1 in the DNA sequence.[11] One irinotecan-derived SN-38 molecule stacks against the base pairs flanking the topoisomerase-induced cleavage site and poisons (inactivates) the topoisomerase 1 enzyme.[11]
Upon bacteriophage (phage) T4 infection of its bacterial host, Escherichia coli, the phage genome specifies a gene product (gp55.2) that inhibits the bacterial topoisomerase I.[12] Gp55.2 binds DNA and specifically blocks the relaxation of negatively supercoiled DNA by topoisomerase I. This inhibition appears to be an adaptation to subtly modulate host topoisomerase I activity during infection to ensure optimal phage yield.
Synthetic lethality
Synthetic lethality arises when a combination of deficiencies in the expression of two or more genes leads to cell death, whereas a deficiency in only one of these genes does not. The deficiencies can arise through mutation, epigenetic alteration or by inhibition of a gene's expression.
Topoisomerase 1 inhibition is synthetically lethal with deficiency of expression of certain DNA repair genes. In human patients the deficient DNA repair genes include
Autoantibodies
References
- S2CID 205496065.
- PMID 11395412.
- PMID 7994576.
- PMID 7770916.
- S2CID 4642743.
- S2CID 4314431.
- S2CID 4426078.
- PMID 16650908.
- PMID 16395333.
- PMID 17804808.
- ^ PMID 23259582.
- ^ Mattenberger Y, Silva F, Belin D. 55.2, a phage T4 ORFan gene, encodes an inhibitor of Escherichia coli topoisomerase I and increases phage fitness. PLoS One. 2015 Apr 14;10(4):e0124309. doi: 10.1371/journal.pone.0124309. PMID: 25875362; PMCID: PMC4396842
- PMID 16723399.
- PMID 25310185.
- PMID 27638859.
- PMID 23377825.
- PMID 27618784.
- ^ Product Name: SCL-70 Antigen Archived 2006-03-19 at the Wayback Machine at ImmunoVision.com, retrieved April 2011
External links
- DNA+Topoisomerases,+Type+I at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
