ATM serine/threonine kinase
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) |
| ||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 11: 108.22 – 108.37 Mb | Chr 9: 53.35 – 53.45 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
ATM serine/threonine kinase or Ataxia-telangiectasia mutated, symbol ATM, is a
In 1995, the gene was discovered by Yosef Shiloh[6] who named its product ATM since he found that its mutations are responsible for the disorder ataxia–telangiectasia.[7] In 1998, the Shiloh and Kastan laboratories independently showed that ATM is a protein kinase whose activity is enhanced by DNA damage.[8][9]
Introduction
Throughout the
Structure
The ATM gene codes for a 350 kDa protein consisting of 3056 amino acids.

Function
A complex of the three proteins

The protein kinase ATM may also be involved in mitochondrial homeostasis, as a regulator of mitochondrial autophagy (mitophagy) whereby old, dysfunctional mitochondria are removed.[16] Increased ATM activity also occurs in viral infection where ATM is activated early during dengue virus infection as part of autophagy induction and ER stress response.[17]
Regulation
A functional MRN complex is required for ATM activation after DSBs. The complex functions upstream of ATM in mammalian cells and induces conformational changes that facilitate an increase in the affinity of ATM towards its substrates, such as CHK2 and p53.[10]
Inactive ATM is present in the cells without DSBs as dimers or multimers. Upon DNA damage, ATM autophosphorylates on residue Ser1981. This phosphorylation provokes dissociation of ATM dimers, which is followed by the release of active ATM monomers.
Germline mutations and cancer risk
People who carry a
Somatic ATM mutations in sporadic cancers
Mutations in the ATM gene are found at relatively low frequencies in sporadic cancers. According to COSMIC, the Catalogue Of Somatic Mutations In Cancer, the frequencies with which heterozygous mutations in ATM are found in common cancers include 0.7% in 713 ovarian cancers, 0.9% in central nervous system cancers, 1.9% in 1,120 breast cancers, 2.1% in 847 kidney cancers, 4.6% in colon cancers, 7.2% among 1,040 lung cancers and 11.1% in 1790 hematopoietic and lymphoid tissue cancers.
Frequent epigenetic deficiencies of ATM in cancers
ATM is one of the DNA repair genes frequently hypermethylated in its promoter region in various cancers (see table of such genes in Cancer epigenetics). The promoter methylation of ATM causes reduced protein or mRNA expression of ATM.
More than 73% of brain tumors were found to be methylated in the ATM gene promoter and there was strong inverse correlation between ATM promoter methylation and its protein expression (p < 0.001).[30]
The ATM gene promoter was observed to be hypermethylated in 53% of small (impalpable) breast cancers[31] and was hypermethylated in 78% of stage II or greater breast cancers with a highly significant correlation (P = 0.0006) between reduced ATM mRNA abundance and aberrant methylation of the ATM gene promoter.[32]
In non-small cell lung cancer (NSCLC), the ATM promoter methylation status of paired tumors and surrounding histologically uninvolved lung tissue was found to be 69% and 59%, respectively. However, in more advanced NSCLC the frequency of ATM promoter methylation was lower at 22%.[33] The finding of ATM promoter methylation in surrounding histologically uninvolved lung tissue suggests that ATM deficiency may be present early in a field defect leading to progression to NSCLC.
In squamous cell carcinoma of the head and neck, 42% of tumors displayed ATM promoter methylation.[34]
DNA damage appears to be the primary underlying cause of cancer,
Meiosis
ATM functions during meiotic prophase.[39] The wild-type ATM gene is expressed at a four-fold increased level in human testes compared to somatic cells (such as skin fibroblasts).[40] In both mice and humans, ATM deficiency results in female and male infertility. Deficient ATM expression causes severe meiotic disruption during prophase I.[41] In addition, impaired ATM-mediated DNA DSB repair has been identified as a likely cause of aging of mouse and human oocytes.[42] Expression of the ATM gene, as well as other key DSB repair genes, declines with age in mouse and human oocytes and this decline is paralleled by an increase of DSBs in primordial follicles.[42] These findings indicate that ATM-mediated homologous recombinational repair is a crucial function of meiosis.
Inhibitors
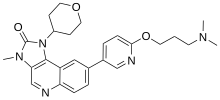
Several ATM kinase inhibitors are currently known, some of which are already in clinical trials.[43][44][45] One of the first discovered ATM inhibitors is caffeine with an IC50 of 0.2 mM and only a low selectivity within the PIKK family.[46][47] Wortmannin is an irreversible inhibitor of ATM with no selectivity over other related PIKK and PI3K kinases.[48] The most important group of inhibitors are compounds based on the 3-methyl-1,3-dihydro-2H-imidazo[4,5-c]quinolin-2-one scaffold. The first important representative is the inhibitor is Dactolisib (NVP-BEZ235), which was first published by Novartis as a selective mTOR/PI3K inhibitor.[49] It was later shown to also inhibit other PIKK kinases such as ATM, DNA-PK and ATR.[50] Various optimisation efforts by AstraZeneca (AZD0156, AZD1390), Merck (M4076) and Dimitrov et al. have led to highly active ATM inhibitors with greater potency.[51][52][53]

Interactions
Ataxia telangiectasia mutated has been shown to interact with:
Tefu
The Tefu protein of Drosophila melanogaster is a structural and functional homolog of the human ATM protein.[78] Tefu, like ATM, is required for DNA repair and normal levels of meiotic recombination in oocytes.
See also
- Ataxia telangiectasia
- Ataxia telangiectasia and Rad3 related
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000149311 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000034218 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- S2CID 237294441.
- PMID 7792600.
- ^ "Entrez Gene: ATM ataxia telangiectasia mutated (includes complementation groups A, C and D)".
- ^ PMID 9733514.
- ^ PMID 9733515.
- ^ PMID 18066086.
- ^ "Serine-protein kinase ATM - Homo sapiens (Human)".
- S2CID 237615473.
- ^ PMID 19779456.
- S2CID 32281084.
- ^ ISBN 978-0-19-920610-0.
- PMID 22144182.
- PMID 26938301.
- S2CID 4403303.
- PMID 33509806.
- PMID 9916992.
- PMID 15279774.
- ^ PMID 11016625.
- PMID 23851492.
- PMID 17683622.
- PMID 31676541.
- S2CID 220931196.
- PMID 26563132.
- S2CID 34149115.
- PMID 26517239.
- S2CID 35412479.
- PMID 26255234.
- PMID 15516988.
- PMID 15958624.
- PMID 16139561.
- PMID 18403632.
- PMID 18082599.
- PMID 18704159.
- PMID 17616978.
- PMID 14681204.
- S2CID 23743474.
- PMID 9735362.
- ^ PMID 23408054.
- ^ "CTG Labs - NCBI". clinicaltrials.gov. 16 September 2022. Retrieved 2023-08-29.
- ^ "CTG Labs - NCBI". clinicaltrials.gov. Retrieved 2023-08-29.
- ^ "CTG Labs - NCBI". clinicaltrials.gov. 18 July 2023. Retrieved 2023-08-29.
- PMID 10531013.
- PMID 10485486.
- PMID 9766667.
- PMID 18606717.
- PMID 21552262.
- PMID 29683659.
- PMID 35405736.
- S2CID 247356817.
- ^ PMID 10212258.
- ^ PMID 11375976.
- S2CID 4334242.
- ^ PMID 10608806.
- ^ PMID 10783165.
- PMID 10866324.
- PMID 10550055.
- PMID 11114888.
- S2CID 43554268.
- PMID 12034743.
- PMID 10464290.
- S2CID 16580666.
- PMID 14499622.
- PMID 15632067.
- PMID 15159397.
- S2CID 23994762.
- PMID 9135004.
- S2CID 4429058.
- S2CID 3266654.
- S2CID 3078706.
- PMID 19015526.
- PMID 11877376.
- S2CID 12380387.
- PMID 12697768.
- PMID 15256487.
Further reading
- Giaccia AJ, Kastan MB (October 1998). "The complexity of p53 modulation: emerging patterns from divergent signals". Genes & Development. 12 (19): 2973–83. PMID 9765199.
- Akst J (2015). "Another Telomere-Regulating Enzyme Found". The Scientist (November 12).
- Kastan MB, Lim DS (December 2000). "The many substrates and functions of ATM". Nature Reviews. Molecular Cell Biology. 1 (3): 179–86. S2CID 10691352.
- Shiloh Y (2002). "ATM: From Phenotype to Functional Genomics — and Back". The Human Genome. pp. 51–70. )
- Redon C, Pilch D, Rogakou E, Sedelnikova O, Newrock K, Bonner W (April 2002). "Histone H2A variants H2AX and H2AZ". Current Opinion in Genetics & Development. 12 (2): 162–9. PMID 11893489.
- Tang Y (February 2002). "[ATM and Cancer]". Zhongguo Shi Yan Xue Ye Xue Za Zhi. 10 (1): 77–80. PMID 12513844.
- Shiloh Y (March 2003). "ATM and related protein kinases: safeguarding genome integrity". Nature Reviews. Cancer. 3 (3): 155–68. S2CID 22770833.
- Gumy-Pause F, Wacker P, Sappino AP (February 2004). "ATM gene and lymphoid malignancies". Leukemia. 18 (2): 238–42. PMID 14628072.
- Kurz EU, Lees-Miller SP (2005). "DNA damage-induced activation of ATM and ATM-dependent signaling pathways". DNA Repair. 3 (8–9): 889–900. PMID 15279774.
- Abraham RT (2005). "The ATM-related kinase, hSMG-1, bridges genome and RNA surveillance pathways". DNA Repair. 3 (8–9): 919–25. PMID 15279777.
- Lavin MF, Scott S, Gueven N, Kozlov S, Peng C, Chen P (2005). "Functional consequences of sequence alterations in the ATM gene". DNA Repair. 3 (8–9): 1197–205. PMID 15279808.
- Meulmeester E, Pereg Y, Shiloh Y, Jochemsen AG (September 2005). "ATM-mediated phosphorylations inhibit Mdmx/Mdm2 stabilization by HAUSP in favor of p53 activation". Cell Cycle. 4 (9): 1166–70. PMID 16082221.
- Ahmed M, Rahman N (September 2006). "ATM and breast cancer susceptibility". Oncogene. 25 (43): 5906–11. PMID 16998505.
External links
- https://web.archive.org/web/20060107000211/http://www.hprd.org/protein/06347
- Drosophila telomere fusion - The Interactive Fly
- GeneReviews/NCBI/NIH/UW entry on Ataxia telangiectasia
- OMIM entries on Ataxia telangiectasia
- Human ATM genome location and ATM gene details page in the UCSC Genome Browser.
- Overview of all the structural information available in the PDB for UniProt: Q13315 (Serine-protein kinase ATM) at the PDBe-KB.
