User:Linhui Li/sandbox
| Human gammaherpesvirus 8 | |
|---|---|
| |
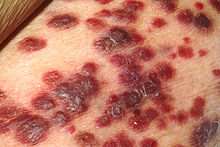
| Kaposi's sarcoma | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Duplodnaviria |
| Kingdom: | Heunggongvirae
|
| Phylum: | Peploviricota |
| Class: | Herviviricetes |
| Order: | Herpesvirales |
| Family: | Orthoherpesviridae
|
| Genus: | Rhadinovirus |
| Species: | Human gammaherpesvirus 8
|
Kaposi's sarcoma-associated herpesvirus (KSHV) is the ninth known human
History
In 1872,
Careful analysis of epidemiologic data by Valerie Beral, Thomas Peterman and Harold Jaffe,

As early as 1984, scientists reported seeing herpesvirus-like structures in KS tumors examined under
The discovery of this
Virology
KSHV is a
After infection, the virus enters into lymphocytes via macropinosomes where it remains in a latent ("quiet") state. Only a subset of genes that are encoded in KSHV latency associated region (KLAR) are expressed during latency including latency-associated nuclear antigen (LANA), vFLIP, vCyclin and 12 microRNA. Latency is the hallmark of all KSHV associated etiologies known till date including all the KSHV associated oncogenesis. It has been shown that both protein coding genes such and noncoding gene (microRNA) encoded in KLAR region are important for KSHV associated tumorigenesis. To study the functions of microRNA, a detailed protocol of bacmid mutagenesis and a complete set of cell-lines carrying microRNA deletion mutants have been established by leading researchers in the field, that are available as a resource to any international researcher working in field of virus-associated cancer.[11] Additionally, in another study, it has been shown that vFLIP and vCyclin interfere with the TGF-β signaling pathway indirectly by inducing oncogenic host mir17-92 cluster.[12] These observations represents a novel mechanism that may be important for KSHV tumorigenesis and angiogenesis, a hallmark of KS. The development of crucial tools such as complete set of 12 microRNA deletion mutants are important development in studying the functions of KLAR gene in context of KSHV associated tumorigenesis [13] Crucial for the Entry of the KSHV [14] is the EPH receptor A2,[15] Hrs,[16] TSG101[17] and a few Integrins, whose identity has yet to be confirmed.[18] The virus exists as a circular piece of DNA called an episome and uses the cellular replication machinery to replicate itself. LANA is the only viral protein that is required for virus DNA replication during latency. The viral episome is chromatinized upon entry into the host cell nucleus.[19] LANA tethers the viral DNA to cellular chromosomes, inhibits p53 and retinoblastoma protein and suppresses viral genes needed for full virus production and assembly ("lytic replication"). Why only a subset of virus genes expressed during latency is not fully understood. But it has been shown that the latency associated gene expression can be explained in part by a characteristic epigenetic state that KSHV episome acquires during latency. LANA play an important role during latency. It is shown to regulate both host and virus transcripts and binds to multiple active promoters including host promoters during latency.[20] In the same study, it has also been shown that LANA associates with host machinary hSET1 that creats H3K4me3 marks.[20] However, exact mechanism why KLAR region remains active during latency is not known. It is also shown that Various signals such as inflammation may provoke the virus to enter into lytic replication.When lytic replication occurs, the viral episome starts replicating itself in the form of linear DNA molecules that are packaged into virus particles which are expelled from the cell, to infect new cells or to be transmitted to a new host. When the virus enters into lytic replication, thousands of virus particles can be made from a single cell, which usually results in death of the infected cell.[citation needed]
The viral genome consists of a ~145 kbase long unique region, encoding all of expressed viral genes, which is flanked by ~20-30 kbases of terminal repeat sequences.[21] Each terminal repeat unit is 801 bp in length, has 85% G+C content and is oriented in a repetitive head-to-tail fashion. During latency, the virus genome depends on the host replication machinery and replicates as a closed circular episome ("plasmid") using sequences within the terminal repeats as a replication origin. When the virus reactivates into lytic replication, it is believed that the virus genome is replicated as a continuous linear molecule off from an episome (so-called rolling circle model). As each unit genome is replicated, it is cut within the terminal repeat region, and then packaged into a virus particle (virion). The virus then becomes enveloped with a lipid membrane as it transits the nucleus and the cytoplasm to exit the cell. Thus, whereas KSHV genome is circular in the nucleus of latently infected cells, it is packaged into infectious viruses as a linear molecule. Once the virus newly infects a cell, the lipid membrane is shed and the virion travels to the nucleus. The viral genome is released where it recircularizes through a poorly understood process that appears to involve homologous recombination.[citation needed]
The primary viral protein responsible for the switch between latent and lytic replication is known as the ORF50 Replication Transactivation Activator (RTA). When cell signaling conditions activate the generation of RTA, it in turn activates synthesis of a stereotypic cascade of secondary and tertiary viral proteins that ultimately make components of the virus capsid and also the DNA synthesis enzymes required to replicate the virus genome.[22]
Pathophysiology
The mechanisms by which the virus is contracted are not well understood. Healthy individuals can be infected with the virus and show no signs or symptoms, due to the immune system's ability to keep the infection in check. Infection is of particular concern to the immunocompromised. Cancer patients receiving chemotherapy, AIDS patients, and organ transplant patients are all at a high risk of showing signs of infection.[citation needed].
Recent advances in sequencing technologies have uncovered that virus is chromatinized during latency. It has also been shown that virus encoded microRNA manipulates and interacts not only with host mRNA but also deregulate host long non-coding RNA (lncRNA).[23] More recently, circularRNAs (circRNAs) are recently discovered in both EBV and KSHV [24][25][26]
Infection with this virus is thought to be lifelong, but a healthy immune system will keep the virus in check. Many people infected with KSHV will never show any symptoms. Kaposi's sarcoma occurs when someone who has been infected with KSHV becomes immunocompromised due to AIDS, medical treatment, or very rarely aging.
KSHV is a known causative agent of four diseases:[4][27]
- Kaposi's sarcoma – an angioproliferative tumor that can involve skin (most often), lymph nodes, or viscera,
- lymphoproliferative disorder,
- Primary effusion lymphoma – KSHV-associated aggressive lymphoma involving B-cells that are immature plasma cells termed plasmablasts and regarded as one type of lymphoid neoplasms with plasmablastic differentiation,
- KSHV inflammatory cytokine syndrome – a syndrome with MCD-like symptoms but without associated pathology.
Epidemiology
In the 1970s, the global prevalence rate for HHV-8 is 2 to 10%.
In countries with low seroprevalence, HHV-8 is primarily limited to AIDS and KS patients.[38] In countries with high seroprevalence, infection is frequent in childhood,[39] indicating a likely mother-to-child transmission by saliva[40].[41] In a Zambian survey, all children with KS had mothers who were positive for HHV-8, whereas not all children whose mothers had KS were HHV-8 positive.[42] In another Zambian survey, 13.8% of children were seropositive for HHV-8 by age 4.[43] Seroprevalence has not been shown to vary significantly because of gender or marital status.[35]
Evolution
The most recent common ancestor of this virus in the Mediterranean, Iran, and Xinjiang, China, has been estimated to have evolved 29,872 years (95% highest probability density 26,851-32,760 years) ago.[44] the most recent common ancestor for viruses isolated in Xinjiang was 2037 years (95% highest probability density 1843–2229 years) ago. Given the historical links between the Mediterranean and Xinjiang during the Roman period it seems likely that this virus was introduced to Xinjiang along the Silk Road. The mutation rate was estimated to be 3.44 × 10−6 substitutions per site per year (95% highest probability density 2.26 × -6 to 4.71 × 10−6). However, the global distribution of different genotypes of KSHV and the potential transmission path need further studies.[citation needed]
Typing of isolates is based on the variable K1 membrane protein. Six types are recognised (A-F).[45]
Prevention
Since persons infected with KSHV will asymptomatically give the virus, caution should be used by
Treatment
Kaposi's sarcoma is usually a localized tumor that can be treated either surgically or through local irradiation. Chemotherapy with drugs such as liposomal
Although KSHV affects the host immune system, there is ample chance for clinical intervention to recover this change. One challenge is overexpression inhibitory of target cell repress immune. Under longtime inflammation stimulation, the target cell unable to respond, which leads to an exhausted phenotype. The activation immunotherapies can revive and enhance immune cell function. Comparing to other immunotherapies, therapies targeting the anti-programmed cell death protein 1 (PD-1)/programmed death-ligand 1 (PD-L1) has been a great success. Because of KSHV infection, the monocytes increase the expression of PD-1, which is an inhibitory molecule, and cause immune escape in many tumor types. There is high PD-1 expression in NK cells from KS-HIV patients and cause exhausted phenotype. The anti-PD-1 antibody, (nivolumab or pembrolizumab), demonstrated a significant antitumor effect. Nivolumab is currently an ongoing phase I clinical trial, and Pembrolizumab has shown its function in treatment for HIV and KS patients in phase I and is in a phase II trial for treatment. A thalidomide analog medicine - Pomalidomide was also granted by the FDA in 2011. Pomalidomide was shown to recover the expression of MHC-1, which help cell display intracellular proteins to cytotoxic T cells, and it also can repress the expression of PD-L1 and increase the CD8+ T cell killing.[48]
KSHV genes
KSHV encodes for ~90 genes and multiple non-coding RNAs, such as microRNAs.[49] The "ORF" genes are named based on genome position of the homologous genes in the first rhadinovirus described, herpesvirus saimiri. The "K" genes are unique to KSHV, Some KSHV genes have well-characterized functions, while others remain uncharacterized.
ORF2 – dihydrofolate reductase
ORF8 – gB – envelope glycoprotein involved in viral entry
ORF9 – Pol8 – DNA polymerase required for viral DNA replication
ORF10 – regulates RNA export and responses to type I IFNs
ORF16 – v
ORF18, ORF24, ORF30, ORF31, ORF34, ORF66 – viral transcription factors required for the expression of late genes
ORF21 – vTK – thymidine kinase
ORF22 – gH – envelope glycoprotein involved in viral entry
ORF23 – uncharacterized
ORF25, ORF26 and ORF65 – capsid proteins
ORF33 – involved in viral particle formation
ORF34 – unclear function
ORF35 – unclear function, mutant does not express early viral genes
ORF36 – vPK – viral protein kinase with multiple roles in replication cycle
ORF37 – SOX – dual function protein – DNase activity required for genome packaging and RNase activity regulates host gene expression
ORF38 – involved in viral particle formation
ORF39 – gM – envelope glycoprotein
ORF40 and ORF41 – helicase and primase – DNA replication
ORF42 – uncharacterized
ORF45 - tegument protein, binds and prevents dephosphorylation of p90 ribosomal S6 kinases (RSKs) and ERK for modulate the ERK/RSK MAPK signaling pathway
ORF47 – gL – envelope glycoprotein involved in viral entry
ORF49 – may be required for viral gene expression
ORF50 – RTA, replication and transcription activator – the major transcription factor driving lytic KSHV reactivation
ORF52 – KicGAS – tegument protein required for formation of virions and inhibition of cGAS DNA sensing
ORF53 – gN – envelope glycoprotein
ORF55 – uncharacterized
ORF57 – MTA – regulates RNA stability, export and translation of viral genes
ORF59 – PF–8 – polymerase processivity factor, accessory subunit of viral DNA polymerase
ORF67 and ORF69 – nuclear egress
ORF70 – thymidylate synthase
ORF72 – vCyclin
ORF73 – LANA, latency-associated nuclear antigen– tethers genome to chromosome during latency, also regulates host gene expression. A cytoplasmic form of LANA may inhibit activation of immune responses.
ORF74 – vGPCR
ORF75 – FGARAT
PAN,
K1 – involved in oncogenesis
K2 - Interleukin 6 homolog, Q2HRC7
K3 and K5 –
K4 – vCCL2 – chemokine
K4.1 – vCCL3 – chemokine
K8 – transcriptional repressor – modulates chromatin
K8.1 – envelope glycoprotein
K9 – v
K10 – v
K10.5 – vIRF3/LANA2 – expressed during latency in B cells
K11 – v
K12 – kaposin
K13 – vFLIP
See also
- Oncovirus (cancer virus)
- 22nd International Workshop on Karposi Sarcoma Herpesvirus (KSHV) and Related Agents, 30 June – 3 July 2019, New York City, USA
References
- ^ "ICTV Master Species List 2018b.v2". International Committee on Taxonomy of Viruses. Retrieved 13 June 2019.
- ^ S2CID 13513517.
- PMID 7700311.
- ^ PMID 27662501.
- S2CID 72037330.
- S2CID 35639169.
- PMID 7997879.
- S2CID 13465834.
- PMID 10749966.
- PMID 10793096.
- PMID 26907327.
- PMID 26545119.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 26907327.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 27854239.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 22635007.
- PMID 26819309.
- PMID 27764233.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 27501260.
- PMID 26870016.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 8962146.
- PMID 18715905.
- PMID 28715488.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 30150410.
- PMID 30455306.
- PMID 30567979.
- PMID 31725418.
- PMID 9874635.
- PMID 27531516.
- ^ PMID 10343069.
- PMID 16095940.
- ^ PMID 19551832.
- ^ PMID 19777527.
- ^ PMID 22168313.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - ^ S2CID 1734745.
- S2CID 7592869.
- PMID 15543582.
- PMID 15231079.
- PMID 10855768.
- )
- PMID 12599072.
- PMID 9815235.
- PMID 18515794.
- ^ Liu Z, Fang Q, Zuo J, Minhas V, Wood C, He N, Zhang T (2017) Was Kaposi's sarcoma-associated herpesvirus introduced into China via the ancient Silk Road? An evolutionary perspective. Arch Virol
- ^ Jary A, Leducq V, Desire N, Petit H, Palich R, Joly V, Canestri A, Gothland A, Lambert-Niclot S, Surgers L, Amiel C, Descamps D, Spano JP, Katlama C, Calvez V, Marcelin AG (2020) New Kaposi's sarcoma-associated herpesvirus variant in men who have sex with men associated with severe pathologies. J Infect Dis
- PMID 10194235.
- S2CID 23476060.
- PMID 32117196.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 15994824.
Further reading
- Edelman DC (2005). "Human herpesvirus 8 – A novel human pathogen". Virology Journal. 2: 78. PMID 16138925.)
{{cite journal}}: CS1 maint: unflagged free DOI (link
External links
- Human Herpesvirus-8: Related Resources HIV InSite
- KSHV Annual Workshop 22nd International Workshop on Karposi Sarcoma Herpesvirus (KSHV) and Related Agents, 30 June – 3 July 2019, New York City, USA
Category:Gammaherpesvirinae
Category:Sexually transmitted diseases and infections
Category:Infectious causes of cancer
Category:IARC Group 1 carcinogens
