Enhancer RNA
Enhancer RNAs (eRNAs) represent a class of relatively long
Discovery
Biogenesis
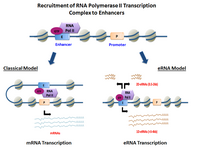
Summary
eRNAs are transcribed from
Depending on the directionality of
1D eRNAs
In most cases, unidirectional
2D eRNAs
Bidirectional
Frequency and timing of eRNA expression
Arner et al.[12] identified 65,423 transcribed enhancers (producing eRNA) among 33 different cell types under different conditions and different timings of stimulation. The transcription of enhancers generally preceded transcription of transcription factors which, in turn, generally preceded messenger RNA(mRNA) transcription of genes.
Carullo et al.
While some enhancers can activate their target promoters at their target genes without transcribing eRNA, most active enhancers do transcribe eRNA during activation of their target promoters.[14]
Functions of eRNA found in the period 2013 to 2021
The functions for eRNA described below have been reported in diverse biological systems, often demonstrated with a small number of specific enhancer-target gene pairs. It is not clear to what extent the functions of eRNA described here can be generalized to most eRNAs.
eRNAs in loop formation
The chromosome loops shown in the figure, bringing an enhancer to the promoter of its target gene, may be directed and formed by the eRNA transcribed from the enhancer after the enhancer is activated.
A transcribed enhancer RNA (eRNA) interacting with the complex of Mediator proteins (see Figure), especially Mediator subunit 12 (MED12), appears to be essential in forming the chromosome loop that brings the enhancer into close association with the promoter of the target gene of the enhancer in the case of five genes studied by Lai et al.[15][16][17] Hou and Kraus,[18] describe two other studies reporting similar results. Arnold et al.[19] review another 5 instances where eRNA is active in forming the enhancer-promoter loop.
eRNAs interact with proteins to affect transcription
One well-studied eRNA is the eRNA of the enhancer that interacts with the promoter of the prostate specific antigen (PSA) gene.[20] The PSA eRNA is strongly up-regulated by the androgen receptor. High PSA eRNA then has a domino effect. PSA eRNA binds to and activates the positive transcription elongation factor P-TEFb protein complex which can then phosphorylate RNA polymerase II (RNAP II), initiating its activity in producing mRNA. P-TEFb can also phosphorylate the negative elongation factor NELF (which pauses RNAP II within 60 nucleotides after mRNA initiation begins). Phosphorylated NELF is released from RNAP II, then allowing RNAP II to have productive mRNA progression (see Figure). Up-regulated PSA eRNA thereby increases expression of 586 androgen receptor-responsive genes. Knockdown of PSA eRNA or deleting a set of nucleotides from PSA eRNA causes decreased presence of phosphorylated (active) RNAP II at these genes causing their reduced transcription.
The negative elongation factor NELF protein can also be released from its interaction with RNAP II by direct interaction with some eRNAs. Schaukowitch et al.[21] showed that the eRNAs of two immediate early genes (IEGs) directly interacted with the NELF protein to release NELF from the RNAP II paused at the promoters of these two genes, allowing these two genes to then be expressed.
In addition, eRNAs appear to interact with as many as 30 other proteins.[19][17][18]
Proposed mechanisms of function up until 2013

The notions that not all
Transcriptional Noise
Since multiple studies have shown that
Transcription-dependent effects
Functional activity in cis
While the two previous models implied that eRNAs were not functionally relevant, this mechanism states that eRNAs are functional
Functional activity in trans
The last model involves
Experimental detection
The detection of eRNAs is fairly recent (2010) and has been made possible through the use of genome-wide investigation techniques such as
Implications in development and disease
Evidence that eRNAs cause downstream effects on the efficiency of enhancer activation and gene transcription suggests its functional capabilities and potential importance. The
Variations in
References
- ^ PMID 20393465.
- ^ PMID 20485488.
- ^ PMID 22905871.
- ^ PMID 28870212.
- ^ PMID 20485488.
- S2CID 1595885.
- ^ S2CID 12778976.
- S2CID 14304312.
- ^ PMID 18509338.
- PMID 22264824.
- PMID 21572438.
- PMID 25678556.
- ^ PMID 32810208.
- PMID 29378788.
- PMID 23417068.
- PMID 25693131.
- ^ PMID 32514177.
- ^ PMID 32888773.
- ^ PMID 31993419.
- PMID 27068475.
- PMID 25263592.
- ^ PMID 15935759.
- PMID 20463730.
- PMID 18710935.
- PMID 16705037.
- PMID 19015660.
- ^ PMID 17512414.
- PMID 22082242.
- PMID 21106759.
- PMID 35177836.
- PMID 27442863.
- PMID 23273978.
- S2CID 6412605.
- PMID 38203707.
External links
- Vista Enhancer Database
- Mouse ENCODE Project Archived 2012-12-13 at the Wayback Machine
- ENCODE Project at UCSC
- PEDB
