Transfer-messenger RNA
| transfer-messenger RNA | |
|---|---|
 | |
| Identifiers | |
| Symbol | tmRNA |
| Rfam | RF00023 |
| Other data | |
| RNA type | gene |
| PDB structures | PDBe |
Transfer-messenger RNA (abbreviated tmRNA, also known as 10Sa RNA and by its genetic name SsrA) is a bacterial

Discovery and early work
tmRNA was first designated 10Sa RNA in 1979, after a mixed "10S" electrophoretic fraction of
Structure
Secondary structure of the standard one-piece tmRNAs

The complete E. coli tmRNA
A prominent feature of every tmRNA is the
tmRNAs typically contain four pseudoknots, one (pk1) upstream of the tag peptide CDS, and the other three pseudoknots (pk2 to pk4) downstream of the CDS. The pseudoknot regions, although generally conserved, are evolutionarily plastic. For example, in the (one-piece) tmRNAs of cyanobacteria, pk4 is substituted with two tandemly arranged smaller pseudoknots. This suggests that tmRNA folding outside the TLD can be important, yet the pseudoknot region lacks conserved residues and pseudoknots are among the first structures to be lost as ssrA sequences diverge in plastid and endosymbiont lineages. Base pairing in the three-pseudoknot region of E. coli tmRNA is disrupted during trans-translation.[7][10]
Two-piece tmRNAs
Circularly permuted ssrA has been reported in three major lineages: i) all alphaproteobacteria and the primitive mitochondria of jakobid protists, ii) two disjoint groups of cyanobacteria (Gloeobacter and a clade containing Prochlorococcus and many Synechococcus), and iii) some members of the betaproteobacteria (Cupriavidus and some Rhodocyclales).[11][12] All produce the same overall two-piece (acceptor and coding pieces) form, equivalent to the standard form nicked downstream of the reading frame. None retain more than two pseudoknots compared to the four (or more) of standard tmRNA.
Alphaproteobacteria have two signature sequences: replacement of the typical T-loop sequence TΨCRANY with GGCRGUA, and the sequence AACAGAA in the large loop of the 3´-terminal pseudoknot. In mitochondria, the MLR has been lost, and a remarkable re-permutation of mitochondrial ssrA results in a small one-piece product in Jakoba libera.[13]
The cyanobacteria provide the most plausible case for evolution of a permuted gene from a standard gene, due to remarkable sequence similarities between the two gene types as they occur in different Synechococcus strains.
tmRNA processing
Most tmRNAs are transcribed as larger precursors which are processed much like
Similar processing at internal sites of permuted precursor tmRNA explains its physical splitting into two pieces. The two-piece tmRNAs have two additional ends whose processing must be considered. For alphaproteobacteria, one 5´ end is the unprocessed start site of transcription.[16] The far 3´ end may in some cases be the result of rho-independent termination.
Three-dimensional structures


High-resolution structures of the complete tmRNA molecules are currently unavailable and may be difficult to obtain due to the inherent flexibility of the MLR. In 2007, the crystal structure of the Thermus thermophilus TLD bound to the SmpB protein was obtained at 3 Å resolution. This structure shows that SmpB mimics the D stem and the anticodon of a canonical tRNA whereas helical section 2a of tmRNA corresponds to the variable arm of tRNA.[18] A
Trans-translation

Coding by tmRNA was discovered in 1995
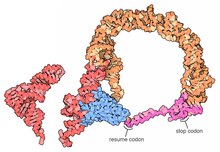
While details of the trans-translation mechanism are under investigation it is generally agreed that tmRNA first occupies the empty A site of the stalled
Mobile genetic elements and the tmRNA gene

ssrA is both a target for some mobile DNAs and a passenger on others. It has been found interrupted by three types of mobile elements. By different strategies none of these disrupt gene function: group I
Non-chromosomal ssrA was first detected in a genomic survey of mycobacteriophages (in 10% of the phages).
Mitochondrial tmRNAs (ssrA gene)
A mitochondrion-encoded, structurally reduced form of tmRNA (mt-tmRNA) was first postulated for the
Identification of ssrA in mitochondrial genomes

Mitochondrial tmRNA genes were initially recognized as short sequences that are conserved among
mt-tmRNA Structure
The standard bacterial tmRNA consists of a tRNA(Ala)-like domain (allowing addition of a non-encoded alanine to mRNAs that happen to lack a stop coding), and an mRNA-like domain coding for a protein tag that destines the polypeptide for proteolysis. The mRNA-like domain was lost in mt-tmRNAs. Comparative sequence analysis indicates features typical for mt-tmRNAs.[30] Most conserved is the primary sequence of the amino acyl acceptor stem. This portion of the molecule has an invariable A residue in the discriminator position and a G-U pair at position 3 (except in Seculamonas ecuadoriensis, which has a G-C pair); this position is the recognition site for alanyl tRNA synthase. P2 is a helix of variable length (3 to 10 base pairs) and corresponds to the anticodon stem of tRNAs, yet without an anticodon loop (as not required for tmRNA function). P2 stabilizes the tRNA-like structure, but four nucleotides invariant across oomycetes and jakobids suggest an additional, currently unidentified function. P3 has five base pairs and corresponds to the T-arm of tRNAs, yet with different consensus nucleotides both in the paired region and the loop. The T-loop sequence is conserved across
mt-tmRNA processing and expression

RNA-Seq data of
See also
References
- PMID 18557701.
- S2CID 22699560.
- PMID 1371186.
- ^ PMID 7524073.
- PMID 8972778.
- PMID 8990402.
- ^ PMID 10219077.
- PMID 17804647.
- ^ PMID 18852454.
- PMID 15811920.
- ^ PMID 10884408.
- PMID 15326226.
- ^ PMID 15037770.
- PMID 1375563.
- PMID 9501180.
- PMID 19617710.
- PMID 12560085.
- ^ PMID 17488812.
- S2CID 28845151.
- PMID 7536743.
- S2CID 29254050.
- PMID 19043582.
- PMID 33815344.
- PMID 7511583.
- PMID 11752287.
- S2CID 5369275.
- PMID 12533482.
- PMID 16789831.
- ^ PMID 23335123.
- ^ PMID 23823571. Archived from the originalon 2014-02-21. Retrieved 2014-02-13.
- ^ Eddy, S. "Infernal website". Retrieved 14 August 2016.
- PMID 8029015.
- PMID 19307242.
Further reading
- Hong SJ, Tran QA, Keiler KC (July 2005). "Cell cycle-regulated degradation of tmRNA is controlled by RNase R and SmpB". Molecular Microbiology. 57 (2): 565–75. PMID 15978085.
