Galactose epimerase deficiency
This article needs additional citations for verification. (May 2020) |
| Galactose epimerase deficiency | |
|---|---|
| Other names | Uridine diphosphate galactose-4-epimerase deficiency |
 | |
| Uridine diphosphate glucose | |
Galactose epimerase deficiency, also known as GALE deficiency, Galactosemia III
Symptoms and signs
Symptoms of congenital Type III Galactosemia are apparent from birth, but vary in severity depending on whether the peripheral or generalized disease form is present. Symptoms may include:[3][4]
- Infantile jaundice
- Infantile hypotonia
- Dysmorphic features
- Sensorineural hearing loss
- Impaired growth
- Cognitive deficiencies
- Depletion of cerebellar Purkinje cells
- Ovarian failure (POI) and hypertrophic hypergonadism
- Liver failure
- Renal failure
- Splenomegaly
- Cataracts
Studies of Type III galactosemia symptoms are mostly descriptive, and precise pathogenic mechanisms remain unknown. This is largely due to a lack of functional animal models of classic galactosemia. The recent development of a Drosophila melanogaster GALE mutant exhibiting galactosemic symptoms may yield a promising future animal model.[3]
Genetics

Galactose
Genetic basis
Various human GALE mutations resulting in Type III galactosemia have been identified.[6] Functional analysis of these mutant GALE isoforms suggests that reduced catalytic efficiency and increased likelihood of proteolytic digestion act causatively in Type III galactosemia.[6]
| Mutated Residue | Biochemical Effect | Clinical Manifestation |
|---|---|---|
| V94M, K257R, L313M, R335H | Strongly impaired turnover number and specificity constant | Severe generalized galactosemia.[3] |
| S81R, T150M, P293L | Mild turnover number impairment | Intermediate galactosemia.[6] |
| L183P, D103G, G90E, N34S | Strongly impaired turnover number and specificity constant; increased proteolytic digestion. | Severe generalized galactosemia.[3] |
Biochemical basis
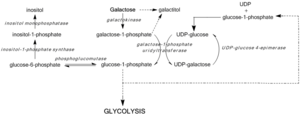
GALE deficiency inhibits UDP-glucose regeneration, preventing the formation of glucose-1-phosphate and leading to the accumulation of galactose and galactose-1-phosphate. High galactose-1-phosphate levels have been shown to interfere with phosphoglucomutase,[7] glycogen phosphorylase,[8] UDP-glycopyrophosphorylase,[9] activity in bacterial models and in vitro, yet in vivo mechanisms toxicity have yet to be confirmed.[3] Regardless, median galactose-1-phosphate levels act as the most accurate predictors of the severity of symptoms associated with Type III galactosemia.[10]
Blockage of the Leloir pathway by GALE deficiency or dysfunction activates alternate pathways of glucose metabolism and leads to galactitol and galactonate formation. Galactonate is metabolized by the pentose phosphate pathway, and is not considered toxic.[11] Galactitol, however, may accumulate in lens fibers, perturbing lens epithelial cell permeability and leading to cell death and cataract formation.[12] GALE deficiency also perturbs glycolipid and glycoprotein biosynthesis due to decreased production of UDP-GalNAc from UDP-GlcNAc.[3]
Diagnosis
Screening for elevated galactose levels may detect GALE deficiency or dysfunction in infants, and mutation studies for GALE are clinically available.[13]
Classification
There are 2 forms of epimerase deficiency: benign RBC deficiency and Severe liver deficiency. Severe form is similar to galactosemia.[citation needed]
Treatment
Individuals presenting with Type III galactosemia must consume a lactose- and galactose-restricted diet devoid of dairy products and mucilaginous plants.[4] Dietary restriction is the only current treatment available for GALE deficiency. As glycoprotein and glycolipid metabolism generate endogenous galactose, however, Type III galactosemia may not be resolved solely through dietary restriction.[3]
References
- ^ Online Mendelian Inheritance in Man (OMIM): Galactose epimerase deficiency - 230350
- ^ Online Mendelian Inheritance in Man (OMIM): UDP-Galactose-4-Epimerase - 606953
- ^ PMID 19859980.
- ^ PMID 10086948.
- PMID 16301867.
- ^ PMID 16302980.
- S2CID 205497901.
- PMID 5920799.
- PMID 10799308.
- PMID 11113841.
- PMID 9396569.
- PMID 5660111.
- S2CID 27586949.
