Y-chromosomal Adam
In
Estimates of the time when Y-MRCA lived have also shifted as modern knowledge of human ancestry changes. For example, in 2013, the discovery of a previously unknown
By definition, it is not necessary that the Y-MRCA and the mt-MRCA should have lived at the same time.[3]
While estimates as of 2014 suggested the possibility that the two individuals may well have been roughly contemporaneous,
Y-chromosomal data taken from a
Definition
million years ago ) |
The Y-chromosomal most recent common ancestor is the
Due to the definition via the "currently living" population, the identity of a MRCA, and by extension of the human Y-MRCA, is time-dependent (it depends on the moment in time intended by the term "currently"). The MRCA of a population may move forward in time as archaic lineages within the population go extinct: once a lineage has died out, it is irretrievably lost. This mechanism can thus only shift the title of Y-MRCA forward in time. Such an event could be due to the total extinction of several basal haplogroups.[3] The same holds for the concepts of matrilineal and patrilineal MRCAs: it follows from the definition of Y-MRCA that he had at least two sons who both have unbroken lineages that have survived to the present day. If the lineages of all but one of those sons die out, then the title of Y-MRCA shifts forward from the remaining son through his patrilineal descendants, until the first descendant is reached who had at least two sons who both have living, patrilineal descendants. The title of Y-MRCA is not permanently fixed to a single individual, and the Y-MRCA for any given population would himself have been part of a population which had its own, more remote, Y-MRCA.
Although the informal name "Y-chromosomal Adam" is a reference to the biblical Adam, this should not be misconstrued as implying that the bearer of the chromosome was the only human male alive during his time.[7] His other male contemporaries may also have descendants alive today, but not, by definition, through solely patrilineal descent; in other words, none of them have an unbroken male line of descendants (son's son's son's … son) connecting them to currently living people.
By the nature of the concept of most recent common ancestors, these estimates can only represent a
Age estimate
Estimates on the age of the Y-MRCA crucially depend on the most archaic known haplogroup extant in contemporary populations. As of 2018[update], this is
Method
In addition to the tendency of the title of Y-MRCA to shift forward in time, the estimate of the Y-MRCA's DNA sequence, his position in the family tree, the time when he lived, and his place of origin, are all subject to future revisions.
The following events would change the estimate of who the individual designated as Y-MRCA was:
- Further sampling of Y chromosomes could uncover previously unknown divergent lineages. If this happens, Y-chromosome lineages would converge on an individual who lived further back in time.
- The discovery of additional deep rooting mutations in known lineages could lead to a rearrangement of the family tree.
- Revision of the Y-chromosome mutation rate (see below) can change the estimate of the time when he lived.
The time when Y-MRCA lived is determined by applying a molecular clock to human Y-chromosomes. In contrast to mitochondrial DNA (mtDNA), which has a short sequence of 16,000 base pairs, and mutates frequently, the Y chromosome is significantly longer at 60 million base pairs, and has a lower mutation rate. These features of the Y chromosome have slowed down the identification of its polymorphisms; as a consequence, they have reduced the accuracy of Y-chromosome mutation rate estimates.[9]
Methods of estimating the age of the Y-MRCA for a population of human males whose Y-chromosomes have been sequenced are based on applying the theories of
These mutations can be used as markers to identify shared patrilineal relationships. Y chromosomes that share a specific mutation are referred to as haplogroups. Y chromosomes within a specific haplogroup are assumed to share a common patrilineal ancestor who was the first to carry the defining mutation. (This assumption could be mistaken, as it is possible for the same mutation to occur more than once.)[citation needed] A family tree of Y chromosomes can be constructed, with the mutations serving as branching points along lineages. The Y-MRCA is positioned at the root of the family tree, as the Y chromosomes of all living males are descended from his Y chromosome.
Researchers can reconstruct ancestral Y chromosome DNA sequences by reversing mutated DNA segments to their original condition. The most likely original or ancestral state of a DNA sequence is determined by comparing human DNA sequences with those of a closely related species, usually non-human primates such as chimpanzees and gorillas. By reversing known mutations in a Y-chromosome lineage, a hypothetical ancestral sequence for the MRCA, Y-chromosomal Adam, can be inferred.
Determining the Y-MRCA's DNA sequence, and the time when he lived, involves identifying the human Y-chromosome lineages that are most divergent from each other—the lineages that share the fewest mutations with each other when compared to a non-human primate sequence in a phylogenetic tree. The common ancestor of the most divergent lineages is therefore the common ancestor of all lineages.
History of estimates
Early estimates of the age for the Y-MRCA published during the 1990s ranged between roughly 200 and 300 thousand years ago (kya).[10] Such estimates were later substantially revised downward, as in Thomson et al. 2000,[9] which proposed an age of about 59,000. This date suggested that the Y-MRCA lived about 84,000 years after his female counterpart
The "hyper-recent" estimate of significantly below 100 kya was again corrected upward in studies of the early 2010s, which ranged at about 120 kya to 160 kya. This revision was due to the rearrangement of the backbone of the Y-chromosome phylogeny following the resequencing of Haplogroup A lineages.[14] In 2013, Francalacci et al. reported the sequencing of male-specific single-nucleotide Y-chromosome polymorphisms (MSY-
The announcement by Mendez et al.
Family tree
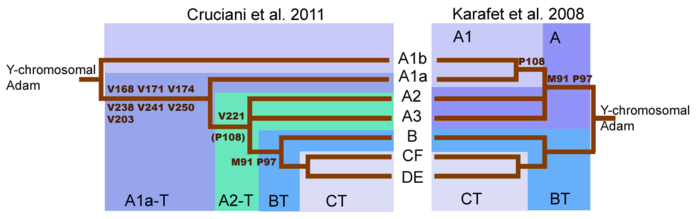
Initial sequencing (Karafet et al., 2008) of the human Y chromosome suggested that two most basal Y-chromosome lineages were
Cruciani et al. 2011, determined that the deepest split in the Y-chromosome tree was found between two previously reported subclades of Haplogroup A, rather than between Haplogroup A and Haplogroup BT. Later, group A00 was found, outside of the previously known tree. The rearrangement of the Y-chromosome family tree implies that lineages classified as Haplogroup A do not necessarily form a
The M91 and P97 mutations distinguish Haplogroup A from Haplogroup BT. Within Haplogroup A chromosomes, the M91 marker consists of a stretch of 8
But according to Cruciani et al. 2011, the region surrounding the M91 marker is a mutational hotspot prone to recurrent mutations. It is therefore possible that the 8T stretch of Haplogroup A may be the ancestral state of M91 and the 9T of Haplogroup BT may be the derived state that arose by an insertion of 1T. This would explain why subclades A1b and A1a-T, the deepest branches of Haplogroup A, both possess the same version of M91 with 8Ts. Furthermore, Cruciani et al. 2011 determined that the P97 marker, which is also used to identify Haplogroup A, possessed the ancestral state in Haplogroup A but the derived state in Haplogroup BT.[20]
Likely geographic origin
As current estimates on TMRCA converge with estimates for the age of
According to Cruciani et al. 2011, the most basal lineages have been detected in
Scozzari et al. (2012) agreed with a plausible placement in "the north-western quadrant of the African continent" for the emergence of the A1b haplogroup.
The revision of Y-chromosomal phylogeny since 2011 has affected estimates for the likely geographical origin of Y-MRCA as well as estimates on time depth. By the same reasoning, future discovery of presently-unknown archaic haplogroups in living people would again lead to such revisions. In particular, the possible presence of between 1% and 4%
See also
- Adam's Curse (book by Bryan Sykes)
- Biblical Adam (Y-chromosomal Adam's namesake)
- Archaeogenetics
- San people
- Eurasian Adam
- Mitochondrial Eve
- Genealogical DNA test
- Genetic genealogy
- Genographic project
- Human Y-chromosome DNA haplogroup
- Identical ancestors point
- Paternal mtDNA transmission
- Y-chromosomal Aaron
References
- ^ PMID 23453668. Archived from the original(PDF) on 24 September 2019. Retrieved 8 March 2013. (primary source)
- PMID 24448544. 'Y-Chromosomal Adam Lived 208,300 Years Ago, Says New Study' , Sci-News.com, 23 January 2014.
- ^ ISBN 9780618619160. Blaine Bettinger (20 July 2007). "Mitochondrial Eve and Y-chromosomal Adam". The Genetic Genealogist.
- ^ S2CID 206550892.
- ^ PMID 25770088. "we date the Y-chromosomal most recent common ancestor (MRCA) in Africa at 254 (95% CI 192–307) kya and detect a cluster of major non-African founder haplogroups in a narrow time interval at 47–52 kya, consistent with a rapid initial colonization model of Eurasia and Oceania after the out-of-Africa bottleneck. In contrast to demographic reconstructions based on mtDNA, we infer a second strong bottleneck in Y-chromosome lineages dating to the last 10 ky. We hypothesize that this bottleneck is caused by cultural changes affecting variance of reproductive success among males."
- ^ PMID 27058445.
- PMID 8450756.
- ^ PMID 24448544.
- ^ PMID 10860948.
- S2CID 1119082. Dorit RL, Akashi H, Gilbert W (1995). "Absence of polymorphism at the ZFY locus on the human Y chromosome". Science. 268 (5214): 1183–85.PMID 7761836. Huang W, Fu YX, Chang BH, Gu X, Jorde LB, Li WH (1998). "Sequence variation in ZFX introns in human populations". Mol Biol Evol. 15 (2): 138–42.PMID 9491612.
- ^ "Genetic 'Adam never met Eve'". BBC News. 2000-10-30. Retrieved 2013-03-08.
- ISBN 978-1-4051-3166-7.
- PMID 17408354.
- PMID 21601174.
- PMID 23908240.
- PMID 23908239.
- ^ University of Michigan Health System (1 August 2013). "The when and where of the Y: Research on Y chromosomes uncovers new clues about human ancestry". ScienceDaily. Retrieved 10 August 2013.
- ^ Rathi A (2 August 2013). "Genetic Adam and Eve may have walked on Earth at the same time". ars technica. Condé Nast. Retrieved 10 August 2013.
- ^ PMID 18385274.
- ^ a b c Fulvio Cruciani, Beniamino Trombetta, Andrea Massaia, Giovanni Destro-Biso, Daniele Sellitto y Rosaria Scozzari 2011, A Revised Root for the human Y-chromosomal Phylogenetic Tree: The Origin of Patrilineal Diversity in Africa
- ^ In a sample of 2204 African Y-chromosomes, 8 chromosomes belonged to either haplogroup A1b or A1a. Haplogroup A1a was identified in two Moroccan Berbers, one Fulbe, and one Tuareg person from Niger. Haplogroup A1b was identified in three Bakola pygmies from Southern Cameroon and one Algerian Berber. Cruciani et al. 2011
- PMID 23145109.
Further reading
- Gibbons, A. (2001). "Modern Men Trace Ancestry to African Migrants". S2CID 83688736.
- Ke, Yuehai (2001). "African Origin of Modern Humans in East Asia: A Tale of 12,000 Y Chromosomes". Science. 292 (5519): 1151–53. S2CID 32685801.
- Bateman, A. J. (1948). "Intra-sexual selection in Drosophila". Heredity. 2 (3): 349–68. PMID 18103134.
- Fu, YX; Li, WH; Donnelly, P.; Tavaré, S.; Balding, D. J.; Griffiths, R. C.; Weiss, G.; Von Haeseler, A.; Rogers, J. (1996). "Estimating the age of the common ancestor of men from the ZFY intron". Science. 272 (5266): 1356–57, author reply 1361–1362. PMID 8650550.
- Donnelly, P; Tavaré, S; Balding, DJ; Griffiths, RC (May 1996). "Estimating the age of the common ancestor of men from the ZFY intron". Science. 272 (5266): 1357–59, author reply 1361–62. PMID 8650551.
- Dorit, RL; Akashi, H; Gilbert, W (May 1995). "Absence of polymorphism at the ZFY locus on the human Y chromosome". Science. 268 (5214): 1183–85. PMID 7761836.
External links
- Documentary Redraws Humans' Family Tree (from National Geographic)
- DNA Mysteries – The Search for Adam (from National Geographic Channel)
- Mitochondrial Eve and Y-chromosomal Adam Diagrams Archived 2014-09-21 at the Wayback Machine
- Y-Chromosome Biallelic Haplogroups
- Most European males 'descended from farmers'
- Why study the Y: Chromosome reveals path of ancestral humans Archived 2009-10-06 at the Wayback Machine
