METAP2
| METAP2 | |||
|---|---|---|---|
Gene ontology | |||
| Molecular function | |||
| Cellular component | |||
| Biological process | |||
| Sources:Amigo / QuickGO | |||
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 12: 95.47 – 95.52 Mb | Chr 10: 93.69 – 93.73 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Methionine aminopeptidase 2 is an enzyme that in humans is encoded by the METAP2 gene.[5][6]
Methionine aminopeptidase 2, a member of the dimetallohydrolase family, is a
- peptide-methionine peptide + methionine
MetAP2 is found in all organisms and is especially important because of its critical role in tissue repair and protein degradation.
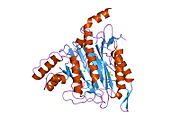
Structure
In living organisms, the start
Active site
The active site of MetAP2 has a structural motif characteristic of many metalloenzymes—including the dioxygen carrier protein,
Dimetal center
The identity of the
Mechanism

The bridging water or hydroxide ligand acts as a nucleophile during the hydrolysis reaction, but the exact mechanism of catalysis is not yet known.[10][19][28] The catalytic mechanisms of hydrolase enzymes depend greatly on the identity of the bridging ligand,[29] which can be challenging to determine due to the difficulty of studying hydrogen atoms via x-ray crystallography.
The histidine residues shown in the mechanism to the right, H178 and H79, are conserved in all MetAPs (MetAP1s and MetAP2s) sequenced to date, suggesting their presence is important to catalytic activity.[30] Based upon X-ray crystallographic data, histidine 79 (H79) has been proposed to help position the methionine residue in the active site and transfer a proton to the newly exposed N-terminal amine.[12] Lowther and Colleagues have proposed two possible mechanisms for MetAP2 in E. coli, shown at the right.[14]
Function
While previous studies have indicated MetAP2 catalyzes the removal of N-terminal methionine residues in vitro, the function of this enzyme in vivo may be more complex. For example, a significant correlation exists between the inhibition of the enzymatic activity of MetAP2 and inhibition of cell growth, thus implicating the enzyme in
Clinical significance

Numerous studies implicate MetAP2 in angiogenesis.
The fumagillin-derived METAP2 inhibitor beloranib (ZGN-433, CDK-732) has shown efficacy in reducing weight in severely obese subjects.[35] MetAP2 inhibitors work by re-establishing insulin sensitivity and balance to the ways the body metabolizes fat, leading to substantial loss of body weight. Development of beloranib was halted in 2016 after two deaths during clinical trials for patients with Praeder-Willi Syndrome.[36] A polymer-drug conjugate of a novel MetAP2 inhibitor called evexomostat being developed by SynDevRx, Inc. entered clinical development for late-stage cancer patients in 2016. Phase 1 dose escalation studies were completed in 2020. In 2022, SynDevRx initiated a Phase 2 clinical study of evexomostat in collaboration with Memorial Sloan Kettering Cancer Center of New York to assess the safety and efficacy in recurrent, metastatic triple-negative breast cancer in combination with the drug eribulin (Halaven(R)). In 2023, SynDevRx initiated another Phase 2 clinical study of evexomostat in combination with the alpelisib (Piqray (R)) and fulvestrant (Faslodex (R)) in metastatic HR+/Her2- breast cancer patients.
Interactions
METAP2 has been shown to
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000111142 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000036112 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- PMID 7644482.
- PMID 8858118.
- ^ .
- ^ PMID 18494467.
- PMID 15453765.
- ^ S2CID 5924813.
- PMID 9263636.
- ^ PMID 9177176.
- ^ PMID 9224570.
- ^ PMID 9770455.
- PMID 2544569.
- PMID 8618900.
- PMID 11511213.
- .
- ^ PMID 11681912.
- ^ PMID 18921993.
- PMID 11848832.
- PMID 10460163.
- PMID 1569059.
- PMID 9733678.
- PMID 12718546.
- PMID 17523636.
- PMID 10555963.
- .
- PMID 16296818.
- PMID 14976199.
- PMID 8679583.
- PMID 18587385.
- ^ PMID 15187415.
- ^ PMID 19246010.
- ^ "Zafgen Announces Positive Topline Phase 1b Data for ZGN-433 in Obesity". MedNews. Drugs.com. 2011-01-01. Retrieved 2011-04-13.
- ^ "Zafgen Halts Development of Beloranib, to Cut Jobs by ~34%". nasdaq.com. July 20, 2016.
- PMID 11123929.
Further reading
- Prigmore E, Ahmed S, Best A, Kozma R, Manser E, Segal AW, et al. (May 1995). "A 68-kDa kinase and NADPH oxidase component p67phox are targets for Cdc42Hs and Rac1 in neutrophils". J. Biol. Chem. 270 (18): 10717–22. PMID 7738010.
- Li X, Chang YH (February 1995). "Molecular cloning of a human complementary DNA encoding an initiation factor 2-associated protein (p67)". Biochim. Biophys. Acta. 1260 (3): 333–6. PMID 7873610.
- Ray MK, Chakraborty A, Datta B, Chattopadhyay A, Saha D, Bose A, et al. (May 1993). "Characteristics of the eukaryotic initiation factor 2 associated 67-kDa polypeptide". Biochemistry. 32 (19): 5151–9. PMID 8098621.
- Liu S, Widom J, Kemp CW, Crews CM, Clardy J (November 1998). "Structure of human methionine aminopeptidase-2 complexed with fumagillin". Science. 282 (5392): 1324–7. PMID 9812898.
- Griffith EC, Su Z, Niwayama S, Ramsay CA, Chang YH, Liu JO (December 1998). "Molecular recognition of angiogenesis inhibitors fumagillin and ovalicin by methionine aminopeptidase 2". Proc. Natl. Acad. Sci. U.S.A. 95 (26): 15183–8. PMID 9860943.
- Datta B, Datta R, Mukherjee S, Zhang Z (1999). "Increased phosphorylation of eukaryotic initiation factor 2alpha at the G2/M boundary in human osteosarcoma cells correlates with deglycosylation of p67 and a decreased rate of protein synthesis". Exp. Cell Res. 250 (1): 223–30. PMID 10388536.
- Gil J, Esteban M, Roth D (2001). "In vivo regulation of the dsRNA-dependent protein kinase PKR by the cellular glycoprotein p67". Biochemistry. 39 (51): 16016–25. PMID 11123929.
- Catalano A, Romano M, Robuffo I, Strizzi L, Procopio A (August 2001). "Methionine aminopeptidase-2 regulates human mesothelioma cell survival: role of Bcl-2 expression and telomerase activity". Am. J. Pathol. 159 (2): 721–31. PMID 11485930.
- Endo H, Takenaga K, Kanno T, Satoh H, Mori S (2002). "Methionine aminopeptidase 2 is a new target for the metastasis-associated protein, S100A4". J. Biol. Chem. 277 (29): 26396–402. PMID 11994292.
- Kanno T, Endo H, Takeuchi K, Morishita Y, Fukayama M, Mori S (2002). "High expression of methionine aminopeptidase type 2 in germinal center B cells and their neoplastic counterparts". Lab. Invest. 82 (7): 893–901. PMID 12118091.
- Datta R, Tammali R, Datta B (2003). "Negative regulation of the protection of eIF2alpha phosphorylation activity by a unique acidic domain present at the N-terminus of p67". Exp. Cell Res. 283 (2): 237–46. PMID 12581743.
- Serero A, Giglione C, Sardini A, Martinez-Sanz J, Meinnel T (December 2003). "An unusual peptide deformylase features in the human mitochondrial N-terminal methionine excision pathway". J. Biol. Chem. 278 (52): 52953–63. PMID 14532271.
- Selvakumar P, Lakshmikuttyamma A, Kanthan R, Kanthan SC, Dimmock JR, Sharma RK (April 2004). "High expression of methionine aminopeptidase 2 in human colorectal adenocarcinomas". Clin. Cancer Res. 10 (8): 2771–5. S2CID 577037.
- Kim S, LaMontagne K, Sabio M, Sharma S, Versace RW, Yusuff N, et al. (May 2004). "Depletion of methionine aminopeptidase 2 does not alter cell response to fumagillin or bengamides". Cancer Res. 64 (9): 2984–7. PMID 15126329.