Myosin
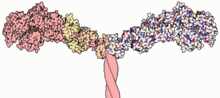
Myosins (/ˈmaɪəsɪn, -oʊ-/[1][2]) are a superfamily of motor proteins best known for their roles in muscle contraction and in a wide range of other motility processes in eukaryotes. They are ATP-dependent and responsible for actin-based motility.
The first myosin (M2) to be discovered was in 1864 by
Following the discovery in 1973 of enzymes with myosin-like function in Acanthamoeba castellanii, a global range of divergent myosin genes have been discovered throughout the realm of eukaryotes.[5]
Although myosin was originally thought to be restricted to muscle cells (hence myo-(s) + -in), there is no single "myosin"; rather it is a very large superfamily of genes whose protein products share the basic properties of actin binding, ATP hydrolysis (ATPase enzyme activity), and force transduction. Virtually all eukaryotic cells contain myosin isoforms. Some isoforms have specialized functions in certain cell types (such as muscle), while other isoforms are ubiquitous. The structure and function of myosin is globally conserved across species, to the extent that rabbit muscle myosin II will bind to actin from an amoeba.[6]
Structure and functions
Domains
Most myosin molecules are composed of a head, neck, and tail domain.
- The head domain binds the filamentous actin, and uses ATP hydrolysis to generate force and to "walk" along the filament towards the barbed (+) end (with the exception of myosin VI, which moves towards the pointed (-) end).
- the neck domain acts as a linker and as a lever arm for transducing force generated by the catalytic motor domain. The neck domain can also serve as a binding site for myosin light chains which are distinct proteins that form part of a macromolecular complex and generally have regulatory functions.
- The tail domain generally mediates interaction with cargo molecules and/or other myosin subunits. In some cases, the tail domain may play a role in regulating motor activity.
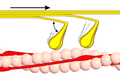
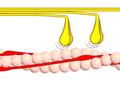
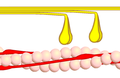
Power stroke
Multiple myosin II molecules generate force in skeletal muscle through a power stroke mechanism fuelled by the energy released from ATP hydrolysis.[7] The power stroke occurs at the release of phosphate from the myosin molecule after the ATP hydrolysis while myosin is tightly bound to actin. The effect of this release is a conformational change in the molecule that pulls against the actin. The release of the ADP molecule leads to the so-called rigor state of myosin.[8] The binding of a new ATP molecule will release myosin from actin. ATP hydrolysis within the myosin will cause it to bind to actin again to repeat the cycle. The combined effect of the myriad power strokes causes the muscle to contract.
Nomenclature, evolution, and the family tree

The wide variety of myosin genes found throughout the eukaryotic phyla were named according to different schemes as they were discovered. The nomenclature can therefore be somewhat confusing when attempting to compare the functions of myosin proteins within and between organisms.
Skeletal muscle myosin, the most conspicuous of the myosin superfamily due to its abundance in
Analysis of the amino acid sequences of different myosins shows great variability among the tail domains, but strong conservation of head domain sequences. Presumably this is so the myosins may interact, via their tails, with a large number of different cargoes, while the goal in each case – to move along actin filaments – remains the same and therefore requires the same machinery in the motor. For example, the
These differences in shape also determine the speed at which myosins can move along actin filaments. The hydrolysis of ATP and the subsequent release of the phosphate group causes the "power stroke", in which the "lever arm" or "neck" region of the heavy chain is dragged forward. Since the power stroke always moves the lever arm by the same angle, the length of the lever arm determines the displacement of the cargo relative to the actin filament. A longer lever arm will cause the cargo to traverse a greater distance even though the lever arm undergoes the same angular displacement – just as a person with longer legs can move farther with each individual step. The velocity of a myosin motor depends upon the rate at which it passes through a complete kinetic cycle of ATP binding to the release of ADP.
Myosin classes
Myosin I
Myosin I, a ubiquitous cellular protein, functions as monomer and functions in
Myosin II
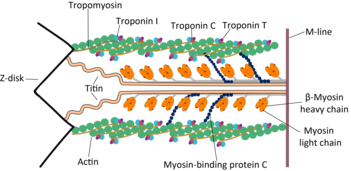
Myosin II (also known as conventional myosin) is the myosin type responsible for producing muscle contraction in muscle cells in most animal cell types. It is also found in non-muscle cells in contractile bundles called stress fibers.[17]
- Myosin II contains two heavy chains, each about 2000 MYH11)[19]) codes for the heavy chains myosin II, but splice variants of this gene result in four distinct isoforms.[18]
- It also contains 4 kDa.[18]These bind the heavy chains in the "neck" region between the head and tail.
- The MLC20 is also known as the regulatory light chain and actively participates in muscle contraction.[18]
- The MLC17 is also known as the essential light chain.[18] Its exact function is unclear, but is believed to contribute to the structural stability of the myosin head along with MLC20.[18] Two variants of MLC17 (MLC17a/b) exist as a result of alternative splicing at the MLC17 gene.[18]
In muscle cells, the long
Myosin III
Myosin III is a poorly understood member of the myosin family. It has been studied 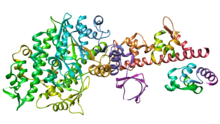
Myosin IV
Myosin IV has a single
Myosin V
Myosin V is an unconventional myosin motor, which is processive as a
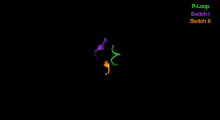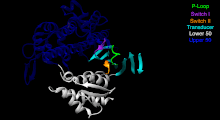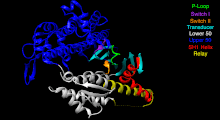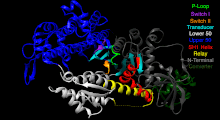
The Myosin V motor head can be subdivided into the following functional regions:[27]
- Nucleotide-binding site - These elements together coordinate di-valent metal cations (usually magnesium) and catalyze hydrolysis:
- Switch I - This contains a highly conserved SSR motif. Isomerizes in the presence of ATP.
- Switch II - This is the Kinase-GTPase version of the Walker B motifDxxG. Isomerizes in the presence of ATP.
- P-loop - This contains the Walker A motifGxxxxGK(S,T). This is the primary ATP binding site.
- Transducer - The seven β-strands that underpin the motor head's structure.[28]
- U50 and L50 - The Upper (U50) and Lower (L50) domains are each around 50 and some regulatory compounds.
- SH1 helix and Relay - These elements together provide an essential mechanism for coupling the enzymatic state of the motor domain to the powerstroke-producing region (converter domain, lever arm, and light chains).[30][31]
- Converter - This converts a change of conformation in the motor head to an angular displacement of the lever arm (in most cases reinforced with light chains).[31]
Myosin VI

Myosin VI is an unconventional myosin motor, which is primarily processive as a dimer, but also acts as a nonprocessive monomer. It walks along actin filaments, travelling towards the pointed end (- end) of the filaments.
Myosin VII
Myosin VII is an unconventional myosin with two FERM domains in the tail region. It has an extended lever arm consisting of five calmodulin binding IQ motifs followed by a single alpha helix (SAH)[35] Myosin VII is required for phagocytosis in Dictyostelium discoideum, spermatogenesis in C. elegans and stereocilia formation in mice and zebrafish.[36]
Myosin VIII
Myosin VIII is a plant-specific myosin linked to cell division;[37] specifically, it is involved in regulating the flow of cytoplasm between cells[38] and in the localization of vesicles to the phragmoplast.[39]
Myosin IX
Myosin IX is a group of single-headed motor proteins. It was first shown to be minus-end directed,[40] but a later study showed that it is plus-end directed.[41] The movement mechanism for this myosin is poorly understood.
Myosin X
Myosin X is an unconventional myosin motor, which is functional as a
Myosin XI
Myosin XI directs the movement of organelles such as
Myosin XII
Myosin XIII
Myosin XIV
This myosin group has been found in the Apicomplexa phylum.[47] The myosins localize to plasma membranes of the intracellular parasites and may then be involved in the cell invasion process.[48]
This myosin is also found in the ciliated protozoan Tetrahymena thermaphila. Known functions include: transporting phagosomes to the nucleus and perturbing the developmentally regulated elimination of the macronucleus during conjugation.
Myosin XV
Myosin XV is necessary for the development of the actin core structure of the non-motile stereocilia located in the inner ear. It is thought to be functional as a monomer.
Myosin XVI
Myosin XVII
Myosin XVIII
MYO18A A gene on chromosome 17q11.2 that encodes actin-based motor molecules with ATPase activity, which may be involved in maintaining stromal cell scaffolding required for maintaining intercellular contact.
Myosin XIX
Unconventional myosin XIX (Myo19) is a mitochondrial associated myosin motor.[49]
Genes in humans
Note that not all of these genes are active.
- Class I:
- Class II: MYH16
- Class III: MYO3A, MYO3B
- Class V:
- Class VI: MYO6
- Class VII: MYO7A, MYO7B
- Class IX: MYO9A, MYO9B
- Class X: MYO10
- Class XV: MYO15A
- Class XVI: MYO16
- Class XVIII: MYO18A, MYO18B
- Class XIX: MYO19
Myosin light chains are distinct and have their own properties. They are not considered "myosins" but are components of the macromolecular complexes that make up the functional myosin enzymes.
Paramyosin
Paramyosin is a large, 93-115kDa
Paramyosins can be found in seafood. A recent computational study showed that following human intestinal digestion, paramyosins of common octopus, Humboldt squid, Japanese abalone, Japanese scallop, Mediterranean mussel, Pacific oyster, sea cucumber, and Whiteleg shrimp could release short peptides that inhibit the enzymatic activities of angiotensin converting enzyme and dipeptidyl peptidase.[52]
References
- ^ "Myosin". Merriam-Webster.com Dictionary.
- OxfordDictionaries.com. Archived from the originalon 24 August 2012. Retrieved 20 January 2016.
- PMID 22566666.
- PMID 15173217.
- ^ from the original on 6 January 2016.
- ISBN 978-0-691-02376-2
- PMID 11810692.
- S2CID 4472407.
- PMID 1558751.
- PMID 8477454.
- PMID 7939893.
- PMID 10984423.
- PMID 11294886.
- S2CID 23407921.
- ^ Macive S (4 June 2003). "Myosin I". Archived from the original on 2 June 2007. Retrieved 23 May 2007.
- PMID 15014434.
- PMID 19851336.
- ^ PMID 20551073.
- PMID 7684189.
- ^ "New Page 2". Archived from the original on 7 July 2009. Retrieved 16 December 2015.
- ^ EntrezGene 53904
- PMID 10722873.
- PMID 22291143.
- PMID 19808891.
- S2CID 11853457.
- PMID 26190073.
- ^ PMID 20192767.
- S2CID 25508217.
- PMID 11005804.
- PMID 28630813.
- ^ PMID 25505494.
- S2CID 14102005.
- PMID 18068125.
- PMID 15473855.
- PMID 28262393.
- ISBN 978-3-527-30594-0.
- PMID 11516337.
- PMID 11351069.
- PMID 10504577.
- S2CID 12158370.
- S2CID 687308.
- PMID 23012428.
- PMID 27580874.
- PMID 19995734.
- PMID 18178669.
- PMID 12628919.
- PMID 11706051.
- PMID 10749937.
- ^ "MYO19 - Unconventional myosin-XIX - Homo sapiens (Human) - MYO19 gene & protein". www.uniprot.org. Retrieved 21 April 2022.
- ^ PMID 987889.
- PMID 185635.
- PMID 35744987.
Further reading
- Gavin RH (2001). "Myosins in protists". A Survey of Cell Biology. International Review of Cytology. Vol. 206. pp. 97–134. PMID 11407764.
- Mooseker MS, Cheney RE (1995). "Unconventional myosins". Annual Review of Cell and Developmental Biology. 11: 633–75. PMID 8689571.
- Sellers JR (March 2000). "Myosins: a diverse superfamily". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1496 (1): 3–22. PMID 10722873.
- Soldati T, Geissler H, Schwarz EC (1999). "How many is enough? Exploring the myosin repertoire in the model eukaryote Dictyostelium discoideum". Cell Biochemistry and Biophysics. 30 (3): 389–411. S2CID 13319819.
- Molecular Biology of the Cell. Alberts, Johnson, Lewis, Raff, Roberts, and Walter. 4th Edition. 949–952.
Additional images
-
Phase 1
-
Phase 2
-
Phase 3
-
Phase 4
External links
- MBInfo – Myosin Isoforms
- MBInfo – The Myosin Powerstroke
- Myosin Video A video of a moving myosin motor protein.
- Myosins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- The Myosin Homepage
- http://cellimages.ascb.org/cdm4/item_viewer.php?CISOROOT=/p4041coll12&CISOPTR=101&CISOBOX=1&REC=2[dead link] Animation of a moving myosin motor protein
- EC 3.6.4.1
- 3D macromolecular structures of myosin from the EM Data Bank(EMDB)