Caenorhabditis elegans
| Caenorhabditis elegans | |
|---|---|
| |
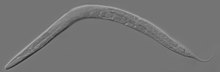
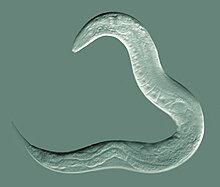
| An adult hermaphrodite C. elegans worm | |
| Scientific classification | |
| Domain: | Eukaryota |
| Kingdom: | Animalia |
| Phylum: | Nematoda |
| Class: | Chromadorea |
| Order: | Rhabditida |
| Family: | Rhabditidae |
| Genus: | Caenorhabditis |
| Species: | C. elegans
|
| Binomial name | |
| Caenorhabditis elegans | |
| Subspecies | |
Caenorhabditis elegans (/ˌsiːnoʊræbˈdaɪtəs ˈɛləɡæns/[6]) is a free-living transparent nematode about 1 mm in length[7] that lives in temperate soil environments. It is the type species of its genus.[8] The name is a blend of the Greek caeno- (recent), rhabditis (rod-like)[9] and Latin elegans (elegant). In 1900, Maupas initially named it Rhabditides elegans. Osche placed it in the subgenus Caenorhabditis in 1952, and in 1955, Dougherty raised Caenorhabditis to the status of genus.[10]
C. elegans is an unsegmented
In 1963, Sydney Brenner proposed research into C. elegans, primarily in the area of neuronal development. In 1974, he began research into the molecular and developmental biology of C. elegans, which has since been extensively used as a model organism.[13] It was the first multicellular organism to have its whole genome sequenced, and in 2019[14] it was the first organism to have its connectome (neuronal "wiring diagram") completed.[15][16][17]
Anatomy

C. elegans is



In relation to lipid metabolism, C. elegans does not have any specialized adipose tissues, a pancreas, a liver, or even blood to deliver nutrients compared to mammals. Neutral lipids are instead stored in the intestine, epidermis, and embryos. The epidermis corresponds to the mammalian adipocytes by being the main triglyceride depot.[19]
The pharynx is a muscular food pump in the head of C. elegans, which is triangular in cross-section. This grinds food and transports it directly to the intestine. A set of "valve cells" connects the pharynx to the intestine, but how this valve operates is not understood. After digestion, the contents of the intestine are released via the rectum, as is the case with all other nematodes.[20] No direct connection exists between the pharynx and the excretory canal, which functions in the release of liquid urine.
Males have a single-lobed gonad, a
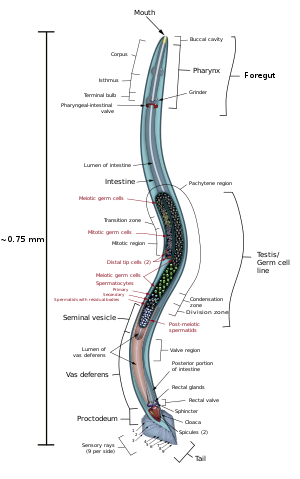
There are 302 neurons in C. elegans, approximately one-third of all the cells in the whole body.
Gut granules
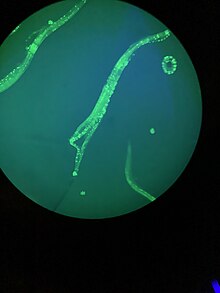
Numerous gut granules are present in the intestine of C. elegans, the functions of which are still not fully known, as are many other aspects of this nematode, despite the many years that it has been studied. These gut granules are found in all of the Rhabditida orders. They are very similar to
Many theories have been posited on the functions of the gut granules, with earlier ones being eliminated by later findings. They are thought to store zinc as one of their functions. Recent chemical analysis has identified the blue fluorescent material they contain as a
Reproduction
The hermaphroditic worm is considered to be a specialized form of self-fertile female, as its
The male can
C. elegans has five pairs of
Sex determination
C. elegans are mostly hermaphroditic organisms, producing both sperms and oocytes.[28] Males do occur in the population in a rate of approximately 1 in 200 hermaphrodites, but the two sexes are highly differentiated.[29][30] Males differ from their hermaphroditic counterparts in that they are smaller and can be identified through the shape of their tail.[30] C.elegans reproduce through a process called androdioecy. This means that they can reproduce in two ways: either through self-fertilization in hermaphrodites or through hermaphrodites breeding with males. Males are produced through non-disjunction of the X chromosomes during meiosis. The worms that reproduce through self-fertilization are at risk for high linkage disequilibrium, which leads to lower genetic diversity in populations and an increase in accumulation of deleterious alleles.[31] In C. elegans, somatic sex determination is attributed to the tra-1 gene.[32] The tra-1 is a gene within the TRA-1 transcription factor sex determination pathway that is regulated post-transcriptionally and works by promoting female development.[32] In hermaphrodites (XX), there are high levels of tra-1 activity, which produces the female reproductive system and inhibits male development.[28][32] At a certain time in their life cycle, one day before adulthood, hermaphrodites can be identified through the addition of a vulva near their tail. In males (XO), there are low levels of tra-1 activity, resulting in a male reproductive system.[32] Recent research has shown that there are three other genes, fem-1, fem-2, and fem-3, that negatively regulate the TRA-1 pathway and act as the final determiner of sex in C. elegans.[28]
Evolution
The sex determination system in C. elegans is a topic that has been of interest to scientists for years.[29] Since they are used as a model organism, any information discovered about the way their sex determination system might have evolved could further the same evolutionary biology research in other organisms. After almost 30 years of research, scientists have begun to put together the pieces in the evolution of such a system.[29] What they have discovered is that there is a complex pathway involved that has several layers of regulation.[29] The closely related organism Caenorhabditis briggsae has been studied extensively and its whole genome sequence has helped put together the missing pieces in the evolution of C. elegans sex determination.[29] It has been discovered that two genes have assimilated, leading to the proteins XOL-1 and MIX-1 having an effect on sex determination in C. elegans as well.[29] Mutations in the XOL-1 pathway leads to feminization in C. elegans .[33] The mix-1 gene is known to hypoactivate the X chromosome and regulates the morphology of the male tail in C. elegans.[34] Looking at the nematode as a whole, the male and hermaphrodite sex likely evolved from parallel evolution.[29] Parallel evolution is defined as similar traits evolving from an ancestor in similar conditions; simply put, the two species evolve in similar ways over time. An example of this would be marsupial and placental mammals. Scientists have also hypothesized that hermaphrodite asexual reproduction, or "selfing", could have evolved convergently by studying species similar to C. elegans[29] Other studies on the sex determination evolution suggest that genes involving sperm evolve at the faster rate than female genes.[35] However, sperm genes on the X chromosome have reduced evolution rates. Sperm genes have short coding sequences, high codon bias, and disproportionate representation among orphan genes.[35] These characteristics of sperm genes may be the reason for their high rates of evolution and may also suggest how sperm genes evolved out of hermaphrodite worms. Overall, scientists have a general idea of the sex determination pathway in C. elegans, however, the evolution of how this pathway came to be is not yet well defined.
Development

Embryonic development
The fertilized zygote undergoes rotational holoblastic cleavage.
Sperm entry into the oocyte commences formation of an anterior-posterior axis.
All cells of the
Axis formation
The resulting daughter cells of the first cell division are called the AB cell (containing PAR-6 and PAR-3) and the P1 cell (containing PAR-1 and PAR-2). A second cell division produces the ABp and ABa cells from the AB cell, and the EMS and P2 cells from the P1 cell. This division establishes the dorsal-ventral axis, with the ABp cell forming the dorsal side and the EMS cell marking the ventral side.[42] Through Wnt signaling, the P2 cell instructs the EMS cell to divide along the anterior-posterior axis.[43] Through Notch signaling, the P2 cell differentially specifies the ABp and ABa cells, which further defines the dorsal-ventral axis. The left-right axis also becomes apparent early in embryogenesis, although it is unclear exactly when specifically the axis is determined. However, most theories of the L-R axis development involve some kind of differences in cells derived from the AB cell.[44]
Gastrulation
Gastrulation occurs after the embryo reaches the 24-cell stage.[45] C. elegans are a species of protostomes, so the blastopore eventually forms the mouth. Involution into the blastopore begins with movement of the endoderm cells and subsequent formation of the gut, followed by the P4 germline precursor, and finally the mesoderm cells, including the cells that eventually form the pharynx. Gastrulation ends when epiboly of the hypoblasts closes the blastopore.[46]
Post-embryonic development


Under environmental conditions favourable for
Each stage transition is punctuated by a molt of the worm's transparent cuticle. Transitions through these stages are controlled by genes of the heterochronic pathway, an evolutionarily conserved set of regulatory factors.[51] Many heterochronic genes code for microRNAs, which repress the expression of heterochronic transcription factors and other heterochronic miRNAs.[52] miRNAs were originally discovered in C. elegans.[53] Important developmental events controlled by heterochronic genes include the division and eventual syncitial fusion of the hypodermic seam cells, and their subsequent secretion of the alae in young adults. It is believed that the heterochronic pathway represents an evolutionarily conserved predecessor to circadian clocks.[54]
Some nematodes have a fixed, genetically determined number of cells, a phenomenon known as eutely. The adult C. elegans hermaphrodite has 959 somatic cells and the male has 1033 cells,[55][56][57] although it has been suggested that the number of their intestinal cells can increase by one to three in response to gut microbes experienced by mothers.[58] Much of the literature describes the cell number in males as 1031, but the discovery of a pair of left and right MCM neurons increased the number by two in 2015.[57] The number of cells does not change after cell division ceases at the end of the larval period, and subsequent growth is due solely to an increase in the size of individual cells.[59]
Ecology
The different
Invertebrates such as
Nematodes can survive
C. elegans, as other nematodes, can be eaten by predator nematodes and other omnivores, including some insects.[71]
The Orsay virus is a virus that affects C. elegans, as well as the Caenorhabditis elegans Cer1 virus[72] and the Caenorhabditis elegans Cer13 virus.
- Interactions with fungi
Wild isolates of Caenorhabditis elegans are regularly found with infections by Microsporidia fungi. One such species, Nematocida parisii, replicates in the intestines of C. elegans.[73]
Arthrobotrys oligospora is the model organism for interactions between fungi and nematodes.[74] It is the most common and widespread nematode capturing fungus.
Use as a model organism
In 1963,
Research has explored the neural and molecular mechanisms that control several behaviors of C. elegans, including chemotaxis, thermotaxis, mechanotransduction, learning, memory, and mating behaviour.[79] In 2019 the connectome of the male was published using a technique distinct from that used for the hermaphrodite. The same paper used the new technique to redo the hermaphrodite connectome, finding 1,500 new synapses.[80]
It has been used as a model organism to study molecular mechanisms in metabolic diseases.[81]
Brenner also chose it as it is easy to grow in bulk populations, and convenient for genetic analysis.
Cell lineage mapping
The developmental fate of every single somatic cell (959 in the adult hermaphrodite; 1031 in the adult male) has been mapped.[85][86] These patterns of cell lineage are largely invariant between individuals, whereas in mammals, cell development is more dependent on cellular cues from the embryo.
As mentioned previously, the first cell divisions of early
Programmed cell death
Programmed cell death (apoptosis) eliminates many additional cells (131 in the hermaphrodite, most of which would otherwise become neurons); this "apoptotic predictability" has contributed to the elucidation of some apoptotic genes. Cell death-promoting genes and a single cell-death inhibitor have been identified.[87]
RNA interference and gene silencing
RNA interference (RNAi) is a relatively straightforward method of disrupting the function of specific genes. Silencing the function of a gene can sometimes allow a researcher to infer its possible function. The nematode can be soaked in, injected with,[88] or fed with genetically transformed bacteria that express the double-stranded RNA of interest, the sequence of which complements the sequence of the gene that the researcher wishes to disable.[89] RNAi has emerged as a powerful tool in the study of functional genomics. C. elegans has been used to analyse gene functions and claim the promise of future findings in the systematic genetic interactions.[90]
Environmental RNAi uptake is much worse in other species of worms in the genus Caenorhabditis. Although injecting RNA into the body cavity of the animal induces gene silencing in most species, only C. elegans and a few other distantly related nematodes can take up RNA from the bacteria they eat for RNAi.[91] This ability has been mapped down to a single gene, sid-2, which, when inserted as a transgene in other species, allows them to take up RNA for RNAi as C. elegans does.[92]
Cell division and cell cycle
Research into
Drug abuse and addiction
Nicotine dependence can also be studied using C. elegans because it exhibits behavioral responses to nicotine that parallel those of mammals. These responses include acute response, tolerance, withdrawal, and sensitization.[98]
Biological databases
As for most model organisms, scientists that work in the field curate a dedicated online database and the WormBase is that for C. elegans. The WormBase attempts to collate all published information on C. elegans and other related nematodes. Their website has advertised a reward of $4000 for the finder of a new species of closely related nematode.[99] Such a discovery would broaden research opportunities with the worm.[100]
Ageing
C. elegans has been a model organism for research into ageing; for example, the inhibition of an insulin-like growth factor signaling pathway has been shown to increase adult lifespan threefold;[101][102] while glucose feeding promotes oxidative stress and reduce adult lifespan by a half.[81] Similarly, induced degradation of an insulin/IGF-1 receptor late in life extended life expectancy of worms dramatically.[103] Long-lived mutants of C. elegans were demonstrated to be resistant to oxidative stress and UV light.[104] These long-lived mutants had a higher DNA repair capability than wild-type C. elegans.[104] Knockdown of the nucleotide excision repair gene Xpa-1 increased sensitivity to UV and reduced the life span of the long-lived mutants. These findings indicate that DNA repair capability underlies longevity.
The capacity to repair DNA damage by the process of nucleotide excision repair declines with age.[105]
C. elegans exposed to 5mM lithium chloride (LiCl) showed lengthened life spans.[106] When exposed to 10μM LiCl, reduced mortality was observed, but not with 1μM.[107]
C. elegans has been instrumental in the identification of the functions of genes implicated in
Sleep
C. elegans is notable in
Sensory biology
While the worm has no eyes, it has been found to be sensitive to light due to a third type of light-sensitive animal
C. elegans is remarkably adept at tolerating acceleration. It can withstand 400,000 g's, according to geneticists at the University of São Paulo in Brazil. In an experiment, 96% of them were still alive without adverse effects after an hour in an ultracentrifuge.[123]
Spaceflight research
C. elegans made news when specimens were discovered to have survived the
Genetics
Genome
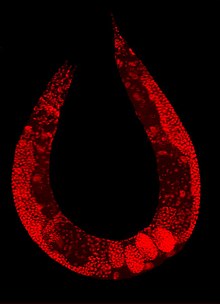
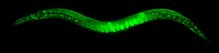
 Karyotype of C. elegans explanation of colors
Mitotic chromosomes of C. elegans. DNA (red)/ Kinetochores (green). Holocentric organisms, including C. elegans, assemble diffuse kinetochores along the entire poleward face of each sister chromatid. | |
| NCBI genome ID | 41 |
|---|---|
| Ploidy | diploid |
| Genome size | 101.169 Mb (haploid) |
| Number of chromosomes | 5 pairs of autosomes (I, II, III, IV and V) + 1 or 2 sex chromosomes (X[128]) |
| Year of completion | 1998 |
| Sequenced organelle | mitochondrion |
| Organelle size | 0,01 Mb |
C. elegans was the first multicellular organism to have its whole genome sequenced. The sequence was published in 1998,[129] although some small gaps were present; the last gap was finished by October 2002.[citation needed] In the run up to the whole genome the C. elegans Sequencing Consortium/C. elegans Genome Project released several partial scans including Wilson et al. 1994.[130][131][132]
Size and gene content
The C. elegans genome is about 100 million
The genome contains an estimated 20,470 protein-coding genes.[135] About 35% of C. elegans genes have human homologs. Remarkably, human genes have been shown repeatedly to replace their C. elegans homologs when introduced into C. elegans. Conversely, many C. elegans genes can function similarly to mammalian genes.[48]
The number of known
Scientific curators continue to appraise the set of known genes; new gene models continue to be added and incorrect ones modified or removed.
The reference C. elegans genome sequence continues to change as new evidence reveals errors in the original sequencing. Most changes are minor, adding or removing only a few base pairs of DNA. For example, the WS202 release of WormBase (April 2009) added two base pairs to the genome sequence.[138] Sometimes, more extensive changes are made as noted in the WS197 release of December 2008, which added a region of over 4,300 bp to the sequence.[139][140]
The C. elegans Genome Project's Wilson et al. 1994 found
Related genomes
In 2003, the genome sequence of the related nematode C. briggsae was also determined, allowing researchers to study the comparative genomics of these two organisms.[141] The genome sequences of more nematodes from the same genus e.g., C. remanei,[142] C. japonica[143] and C. brenneri (named after Brenner), have also been studied using the shotgun sequencing technique.[144] These sequences have now been completed.[145][146]
Other genetic studies
As of 2014, C. elegans is the most basal species in the 'Elegans' group (10 species) of the 'Elegans' supergroup (17 species) in phylogenetic studies. It forms a branch of its own distinct to any other species of the group.[147]
Scientific community
In 2002, the
Many scientists who research C. elegans closely connect to Sydney Brenner, with whom almost all research in this field began in the 1970s; they have worked as either a
See also
References
- ^ Maupas, É (1900). "Modes et formes de reproduction des nématodes". Archives de Zoologie Expérimentale et Générale. 8: 463–624.
- ^ Nigon V (1949). "Les modalités de la reproduction et le déterminisme du sexe chez quelques nematodes libres". Ann. Sci. Nat. Zool. Biol. Anim. 11: 1–132.
- PMID 6096205.
- S2CID 11275388.
- PMID 2550981.
- ^ "Caenorhabditis". Merriam-Webster.com Dictionary.
- ^
Wood, WB (1988). The Nematode Caenorhabditis elegans. ISBN 978-0-87969-433-3.
- .
- ^ καινός (caenos) = new, recent; ῥάβδος (rhabdos) = rod, wand.
- ^ Ferris, H (30 November 2013). "Caenorhabditis elegans". University of California, Davis. Archived from the original on 9 December 2013. Retrieved 2013-11-19.
- JSTOR 3227041.
- ^ "Introduction to sex determination". www.wormbook.org. Retrieved 2017-03-15.
- ^ PMID 4366476.
- ^ a b
Cook, SJ; Jarrell, TA; Brittin, CA; Wang, Y; Bloniarz, AE; Yakovlev, MA; Nguyen, KCQ; Tang, Lt-H; Bayer, EA; Duerr, JS; Bulow, HE; Hobert, O; Hall, DH; Emmons, SW (3 December 2019). "Whole-animal connectomes of both Caenorhabditis elegans sexes". Nature. 571 (7763). US National Library of Medicine, National Institutes of Health: 63–71. PMID 31270481.
- PMID 22462104.
- PMID 23801597.
- ^ Jabr F (2012-10-02). "The Connectome Debate: Is Mapping the Mind of a Worm Worth It?". Scientific American. Retrieved 2014-01-18.
- ISBN 978-0-8153-4105-5.
- PMID 27289335.
- ^ "The C. elegans pharynx: a model for organogenesis". www.wormbook.org. Retrieved 2017-03-15.
- PMID 28889277.
- ^ Nonet, M. (2004) About the nematode Caenorhabdtis elegans
- PMID 18050401.
- ^
Coburn C, Gems D (2013). "The mysterious case of the C. elegans gut granule: death fluorescence, anthranilic acid and the kynurenine pathway". Frontiers in Genetics. 4: 151. PMID 23967012.
- ^
Nayak S, Goree J, Schedl T (January 2005). "fog-2 and the evolution of self-fertile hermaphroditism in Caenorhabditis". PLOS Biology. 3 (1): e6. PMID 15630478.
- PMID 8254383.
- PMID 22903434.
- ^ PMID 17609115.
- ^ PMID 18050417.
- ^ a b "Handbook - Male Introduction". www.wormatlas.org. Retrieved 2021-03-30.
- PMID 25822066.
- ^ S2CID 14951719.
- S2CID 5005906.
- ^ "mix-1 (gene) - WormBase : Nematode Information Resource". wormbase.org. Retrieved 2021-04-23.
- ^ PMID 15371532.
- PMID 8625834.
- ISBN 9781605354705.
- PMID 7758115.
- ^ PMID 18050411.
- ^ Kimble J, Crittenden SL. Germline proliferation and its control. 2005 Aug 15. In: WormBook: The Online Review of C. elegans Biology [Internet]. Pasadena (CA): WormBook; 2005-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK19769/
- ^ a b "WBbt:0006773 (anatomy term)". WormBase (WS242 ed.). May 14, 2014. WBbt:0006773.
- ISBN 9781605354705.
- PMID 9288749.
- ISBN 9781605354705.
- PMID 1601187.
- ISBN 9781605354705.
- PMID 23355522.
- ^ a b "Introduction to C. Elegans". C. Elegans as a model organism. Rutgers University. Archived from the original on 2002-08-18. Retrieved August 15, 2014.
- ^ "Handbook - Introduction".
- ^ "Dauer". www.wormbook.org. Retrieved 2018-09-27.
- PMID 20232378.
- )
- PMID 8252621.
- PMID 15691769.
- PMID 838129.
- PMID 6684600.
- ^ PMID 26469050.
- S2CID 227060212.
- ISBN 978-81-315-0104-7.
- PMID 38190542.
- ^
Félix MA, Braendle C (November 2010). "The natural history of Caenorhabditis elegans". Current Biology. 20 (22): R965–9. PMID 21093785.
- S2CID 259047038.
- PMID 12438649.
- PMID 12654843.
- PMID 9892699.
- S2CID 37703980.
- PMID 32958848.
- PMID 35034106.
- ^
Kiontke K, Sudhaus W (January 2006). "Ecology of Caenorhabditis species". WormBook: 1–14. PMID 18050464.
- ^
Gal TZ, Glazer I, Koltai H (November 2004). "An LEA group 3 family member is involved in survival of C. elegans during exposure to stress". FEBS Letters. 577 (1–2): 21–6. S2CID 21960486.
- ^ Elaine R. Ingham Soil biology primer USDA
- PMID 10523521.
- PMID 22813931.
- .
- PMID 11084387.
- Bibcode:2007AcPPB..38.2201K.
- .
- S2CID 4429113.
- PMID 16139205.
- PMID 31270481.
- ^ PMID 29990370.
- Southwestern Medical Center. Archived from the original on August 15, 2011. Alt. URL Archived 2013-12-08 at the Wayback Machine
- PMID 4366476.
- ^ "Behavior". www.wormbook.org. Retrieved 2018-09-26.
- ^
Sulston JE, Horvitz HR (March 1977). "Post-embryonic cell lineages of the nematode, Caenorhabditis elegans". Developmental Biology. 56 (1): 110–56. PMID 838129.
- ^
Kimble J, Hirsh D (June 1979). "The postembryonic cell lineages of the hermaphrodite and male gonads in Caenorhabditis elegans". Developmental Biology. 70 (2): 396–417. PMID 478167.
- PMID 18719375.
- ^ NIDDK, National Institute of Diabetes and Digestive and Kidney Diseases (March 5, 2015). "Injection of C. elegans Gonads". YouTube. Archived from the original on 2021-11-17. Retrieved March 21, 2020.
- ^
Kamath RS, Fraser AG, Dong Y, Poulin G, Durbin R, Gotta M, Kanapin A, Le Bot N, Moreno S, Sohrmann M, Welchman DP, Zipperlen P, Ahringer J (January 2003). "Systematic functional analysis of the Caenorhabditis elegans genome using RNAi". Nature. 421 (6920): 231–7. S2CID 15745225.
- S2CID 6983519.
- ^
Félix MA (November 2008). "RNA interference in nematodes and the chance that favored Sydney Brenner". Journal of Biology. 7 (9): 34. PMID 19014674.
- ^
Winston WM, Sutherlin M, Wright AJ, Feinberg EH, Hunter CP (June 2007). "Caenorhabditis elegans SID-2 is required for environmental RNA interference". Proceedings of the National Academy of Sciences of the United States of America. 104 (25): 10565–70. PMID 17563372.
- PMID 11058122.
- PMID 14646232.
- PMID 11238374.
- S2CID 2229622.
- ^ Bernstein H, Bernstein C (July 2010) "Evolutionary Origin of Recombination during Meiosis," BioScience 60(7), 498-505. https://doi.org/10.1525/bio.2010.60.7.5
- PMID 17081982.
- ^ "Caenorhabditis isolation guide". WormBase. Archived from the original on November 7, 2007. Retrieved 2007-08-30. Alt. URL Archived 2014-09-05 at the Wayback Machine
- S2CID 85163742.
- PMID 11021802.
- PMID 25517099.
- PMID 34505574.
- ^ PMID 18203746.
- PMID 17472752.
- PMID 17959600.
- PMID 21301855.
- PMID 20012092.
- PMID 21402787.
- PMID 30559463.
- PMID 16151516.
- PMID 32876842.
- PMID 21821789.
- PMID 16477310.
- S2CID 19224032.
- PMID 27593554.
- PMID 23390606.
- PMID 27761361.
- S2CID 210829489.
- PMID 23449971.
- PMID 25264259.
- ^ Teensy, Eyeless Worms Have Completely New Light-Detecting Cells
- ^ Scientific American, August 2018, page 14
- ^ "Worms survived Columbia disaster". BBC News. 1 May 2003. Retrieved 2008-07-11.
- ^ "University sends worms into space". BBC News. 17 January 2009. Retrieved 2009-07-09.
- Discovery News. Archived from the originalon 2012-06-16. Retrieved 2011-05-17.
- PMID 36344513.
- PMID 24591522.
- ^
The C. elegans Sequencing Consortium (December 1998). "Genome sequence of the nematode C. elegans: a platform for investigating biology". Science. 282 (5396): 2012–8. PMID 9851916.
- ^ PMID 9759493.
- ^ PMID 12213389.
- ^ PMID 10669724.
- S2CID 4351788.
- PMID 15642184.
- ^ "WS227 Release Letter". WormBase. 10 August 2011. Archived from the original on 28 November 2013. Retrieved 2013-11-19.
- PMID 17174894.
- PMID 18023116.
- ^ "WS202 Release Letter". WormBase. 29 May 2009. Archived from the original on December 1, 2013. Retrieved 2013-11-19.
- ^ "WS197 Release Letter". WormBase. 27 November 2008. Archived from the original on 17 October 2019. Retrieved 2013-11-19.
- ^ "Genome sequence changes". WormBase. 15 June 2011. Archived from the original on 17 October 2019. Retrieved 2011-08-13.
- PMID 14624247.
- ^ Genome Sequencing Center. "Caenorhabditis remanei: Background". Washington University School of Medicine. Archived from the original on 2008-06-16. Retrieved 2008-07-11.
- ^ Genome Sequencing Center. "Caenorhabditis japonica: Background". Washington University School of Medicine. Archived from the original on 2008-06-26. Retrieved 2008-07-11.
- PMID 461197.
- ^ "UCSC genome browser". Retrieved 8 July 2014.
- PMID 18996895.
- PMID 24727800.
- ^
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (February 1998). "Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans". Nature. 391 (6669): 806–11. S2CID 4355692.
- PMID 19910365.
Further reading
- Bird J, Bird AC (1991). The structure of nematodes. ISBN 978-0-12-099651-3.
- Hope, IA (1999). C. elegans: a practical approach. ISBN 978-0-19-963738-6.
- Riddle DL, Blumenthal T, Meyer RJ, Priess JR (1997). C. elegans II. PMID 21413221.
External links
- Brenner S (2002) Nature's Gift to Science. In. http://nobelprize.org/nobel_prizes/medicine/laureates/2002/brenner-lecture.pdf (also Horvitz and Sulston lectures)
- WormBase – an extensive online database covering the biology and genomics of C. elegans and other nematodes
- WormAtlas – online database on all aspects of C. elegans anatomy with detailed explanations and high-quality images
- WormBook – online review of C. elegans biology
- AceView WormGenes – another genome database for C. elegans, maintained at the NCBI
- C. elegans II – a free online textbook.
- WormWeb Neural Network – an online tool for visualizing and navigating the connectome of C. elegans
- C. elegans movies – a visual introduction to C. elegans
- View the ce11 genome assembly in the UCSC Genome Browser.
- Caenorhabditis elegans at eppo.int (EPPO codeCAEOEL)
- Cylinder, Drew (2022). "Using C. elegans to study anesthesia". )
