Z-DNA
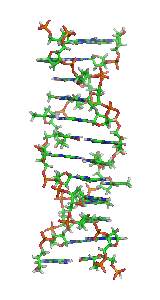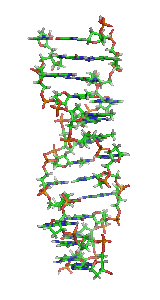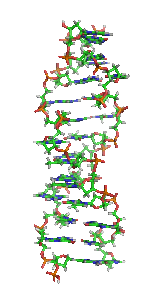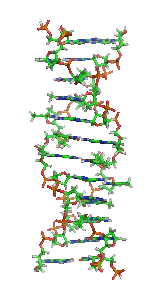
Z-DNA is one of the many possible
History
Left-handed DNA was first proposed by
In 2007, the RNA version of Z-DNA, Z-RNA, was described as a transformed version of an A-RNA double helix into a left-handed helix.[6] The transition from A-RNA to Z-RNA, however, was already described in 1984.[7]
Structure

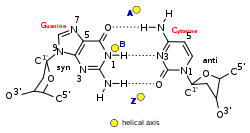
Z-DNA is quite different from the right-handed forms. In fact, Z-DNA is often compared against B-DNA in order to illustrate the major differences. The Z-DNA helix is left-handed and has a structure that repeats every other base pair. The major and minor grooves, unlike A- and B-DNA, show little difference in width. Formation of this structure is generally unfavourable, although certain conditions can promote it; such as alternating
Predicting Z-DNA structure
It is possible to predict the likelihood of a DNA sequence forming a Z-DNA structure. An algorithm for predicting the propensity of DNA to flip from the B-form to the Z-form, ZHunt, was written by P. Shing Ho in 1984 at MIT.[10] This algorithm was later developed by Tracy Camp, P. Christoph Champ, Sandor Maurice, and Jeffrey M. Vargason for genome-wide mapping of Z-DNA (with Ho as the principal investigator).[11]
Pathway of formation of Z-DNA from B-DNA
Since the discovery and crystallization of Z-DNA in 1979, the configuration has left scientists puzzled about the pathway and mechanism from the B-DNA configuration to the Z-DNA configuration.[12] The conformational change from B-DNA to the Z-DNA structure was unknown at the atomic level, but in 2010, computer simulations conducted by Lee et al. were able to computationally determine that the step-wise propagation of a B-to-Z transition would provide a lower energy barrier than the previously hypothesized concerted mechanism.[13] Since this was computationally proven, the pathway would still need to be tested experimentally in the lab for further confirmation and validity, in which Lee et al. specifically states in their journal article, "The current [computational] result could be tested by Single-molecule FRET (smFRET) experiments in the future."[13] In 2018, the pathway from B-DNA to Z-DNA was experimentally proven using smFRET assays.[14] This was performed by measuring the intensity values between the donor and acceptor fluorescent dyes, also known as Fluorophores, in relation to each other as they exchange electrons, while tagged onto a DNA molecule.[15][16] The distances between the fluorophores could be used to quantitatively calculate the changes in proximity of the dyes and conformational changes in the DNA. A Z-DNA high affinity binding protein, hZαADAR1,[17] was used at varying concentrations to induce the transformation from B-DNA to Z-DNA.[14] The smFRET assays revealed a B* transition state, which formed as the binding of hZαADAR1 accumulated on the B-DNA structure and stabilized it.[14] This step occurs to avoid high junction energy, in which the B-DNA structure is allowed to undergo a conformational change to the Z-DNA structure without a major, disruptive change in energy. This result coincides with the computational results of Lee et al. proving the mechanism to be step-wise and its purpose being that it provides a lower energy barrier for the conformational change from the B-DNA to Z-DNA configuration.[13] Contrary to the previous notion, the binding proteins do not actually stabilize the Z-DNA conformation after it is formed, but instead they actually promote the formation of the Z-DNA directly from the B* conformation, which is formed by the B-DNA structure being bound by high affinity proteins.[14]
Biological significance
A biological role for Z-DNA in the regulation of type I interferon responses has been confirmed in studies of three well-characterized rare Mendelian Diseases: Dyschromatosis Symmetrica Hereditaria (OMIM: 127400), Aicardi-Goutières syndrome (OMIM: 615010) and Bilateral Striatal Necrosis/Dystonia. Families with haploid ADAR transcriptome enabled mapping of Zα variants directly to disease, showing that genetic information is encoded in DNA by both shape and sequence.[18] A role in regulating type I interferon responses in cancer is also supported by findings that 40% of a panel of tumors were dependent on the ADAR enzyme for survival.[19]
In previous studies, Z-DNA was linked to both
Z-DNA in transcription
Z-DNA is commonly believed to provide
A study of
Z-DNA sequences upstream of promoter regions have been shown to stimulate transcription. The greatest increase in activity is observed when the Z-DNA sequence is placed three helical turns after the promoter sequence. Furthermore, using micrococcal nuclease-crosslinking technique,[24] Z-DNA is unlikely to form nucleosomes, which are often located before and/or after a Z-DNA forming sequence. Because of this property, Z-DNA is hypothesized to code for the boundary in nucleosome positioning. Since the placement of nucleosomes influences the binding of transcription factors, Z-DNA is thought to regulate the rate of transcription.[24]
Developed behind the pathway of
The toxic effect of ethidium bromide (EtBr) on trypanosomas is caused by shift of their kinetoplastid DNA to Z-form. The shift is caused by intercalation of EtBr and subsequent loosening of DNA structure that leads to unwinding of DNA, shift to Z-form and inhibition of DNA replication.[27]
Discovery of the Zα domain
The first domain to bind Z-DNA with high affinity was discovered in ADAR1 using an approach developed by Alan Herbert.[28][29] Crystallographic and NMR studies confirmed the biochemical findings that this domain bound Z-DNA in a non-sequence-specific manner.[30][31][32] Related domains were identified in a number of other proteins through sequence homology.[29] The identification of the Zα domain provided a tool for other crystallographic studies that lead to the characterization of Z-RNA and the B–Z junction. Biological studies suggested that the Z-DNA binding domain of ADAR1 may localize this enzyme that modifies the sequence of the newly formed RNA to sites of active transcription.[33][34] A role for Zα, Z-DNA and Z-RNA in defense of the genome against the invasion of Alu retro-elements in humans has evolved into a mechanism for the regulation of innate immune responses to dsRNA. Mutations in Zα are causal for human interferonopathies such as the Mendelian Aicardi-Goutières Syndrome.[35][18]Additionally, Zα domains are demonstrated to localize at the stress granules because of their innate ability in binding nucleic acid. Furthermore, different Zα domains bind to the Z conformation of nucleic acid differently providing important avenues for specific targeting in drug discovery.
Consequences of Z-DNA binding to vaccinia E3L protein
As Z-DNA has been researched more thoroughly, it has been discovered that the structure of Z-DNA can bind to Z-DNA binding proteins through
Furthermore, Alexander Rich and Jin-Ah Kwon found that E3L acts as a
Comparison geometries of some DNA forms

| A-form | B-form | Z-form | |
|---|---|---|---|
| Helix sense | right-handed | right-handed | left-handed |
| Repeating unit | 1 bp | 1 bp | 2 bp |
| Rotation/bp | 32.7° | 34.3° | 30° |
| bp/turn | 11 | 10 | 12 |
| Inclination of bp to axis | +19° | −1.2° | −9° |
| Rise/bp along axis | 2.3 Å (0.23 nm) | 3.32 Å (0.332 nm) | 3.8 Å (0.38 nm) |
| Pitch/turn of helix | 28.2 Å (2.82 nm) | 33.2 Å (3.32 nm) | 45.6 Å (4.56 nm) |
| Mean propeller twist | +18° | +16° | 0° |
| Glycosyl angle | anti | anti | C: anti, G: syn |
Sugar pucker |
C3′-endo | C2′-endo | C: C2′-endo, G: C3′-endo |
| Diameter | 23 Å (2.3 nm) | 20 Å (2.0 nm) | 18 Å (1.8 nm) |
See also
- ADAR1
- DNA supercoil
- E3L
- Mechanical properties of DNA
- Proteopedia Z-DNA
- Satellite DNA
- Z-DNA binding protein 1 (ZBP1)
- Zuotin
References
- S2CID 4248932.
- PMID 5045303.
- PMID 7301594.
- S2CID 4337955.
- ^ S2CID 2539819.
- PMID 17437712.
- S2CID 4316862.
- PMID 20439751.
- PMID 16473901.
- PMID 3780676.
- ^ PMID 15598822.
- S2CID 4337955.
- ^ PMID 20666528.
- ^ PMID 29584891.
- PMID 23733413.
- PMID 11730017.
- PMID 9237992.
- ^ PMID 31320745.
- S2CID 155209484.
- ^ S2CID 29059186.
- PMID 6822666.
- S2CID 835548.
- PMID 2006166.
- ^ PMID 17284586.
- ^ PMID 16473937.
- ^ PMID 2552445.
- PMID 21187912.
- PMID 8332463.
- ^ PMID 9237992.
- PMID 9671809.
- PMID 10364558.
- PMID 10535945.
- PMID 11593027.
- ^ Halber, D. (1999-09-11). "Scientists observe biological activities of 'left-handed' DNA". MIT News Office. Retrieved 2008-09-29.
- PMID 30729177.
- ^ PMID 16126896.
- ^ PMID 12777633.
- PMID 14757814.
- ISBN 978-0-126-45750-6.
- PMID 6383204.
- PMID 7937803.
