Euchromatin

Euchromatin (also called "open chromatin") is a lightly packed form of
In
Structure
Euchromatin is composed of repeating subunits known as nucleosomes, reminiscent of an unfolded set of beads on a string, that are approximately 11 nm in diameter.[2] At the core of these nucleosomes are a set of four histone protein pairs: H3, H4, H2A, and H2B.[2] Each core histone protein possesses a 'tail' structure, which can vary in several ways; it is thought that these variations act as "master control switches" through different methylation and acetylation states, which determine the overall arrangement of the chromatin.[2] Approximately 147 base pairs of DNA are wound around the histone octamers, or a little less than 2 turns of the helix.[3] Nucleosomes along the strand are linked together via the histone, H1,[4] and a short space of open linker DNA, ranging from around 0-80 base pairs. The key distinction between the structure of euchromatin and heterochromatin is that the nucleosomes in euchromatin are much more widely spaced, which allows for easier access of different protein complexes to the DNA strand and thus increased gene transcription.[2]
Appearance
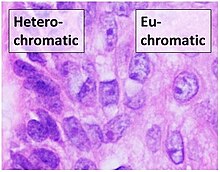
Euchromatin resembles a set of beads on a string at large magnifications.[2] From farther away, it can resemble a ball of tangled thread, such as in some electron microscope visualizations.[5] In both optical and electron microscopic visualizations, euchromatin appears lighter in color than heterochromatin - which is also present in the nucleus and appears darkly[6] - due to its less compact structure.[5] When visualizing chromosomes, such as in a karyogram, cytogenetic banding is used to stain the chromosomes. Cytogenetic banding allows us to see which parts of the chromosome are made up of euchromatin or heterochromatin in order to differentiate chromosomal subsections, irregularities or rearrangements.[7] One such example is G banding, otherwise known as Giemsa staining where euchromatin appears lighter than heterochromatin.[8]
| Giemsa (G-) Banding | Reverse (R-) Banding | Constitutive Heterochromatin (C-) banding | Quinacrine (Q-) banding | Telomeric R (T-) banding | |
|---|---|---|---|---|---|
| Euchromatin | Lighter | Darker | Lighter | Dull | Light |
| Heterochromatin | Darker | Lighter | Darker | Bright (Fluorescent) | Darker (Faint) |
Function

Transcription
Euchromatin participates in the active
It is thought that the cell uses transformation from euchromatin into heterochromatin as a method of controlling
Epigenetics
Epigenetics involves changes in the phenotype that can be inherited without changing the DNA sequence. This can occur through many types of environmental interactions.[16] Regarding euchromatin, post-translational modifications of the histones can alter the structure of chromatin, resulting in altered gene expression without changing the DNA.[17] Additionally, a loss of heterochromatin and increase in euchromatin has been shown to correlate with an accelerated aging process, especially in diseases known to resemble premature aging.[18] Research has shown epigenetic markers on histones for a number of additional diseases.[19][20]
Regulation
Euchromatin is primarily regulated by post-translational modifications to its nucleosomes' histones, conducted by many histone-modifying enzymes. These modifications occur on the histones' N-terminal tails that protrude from the nucleosome structure, and are thought of to recruit enzymes to either keep the chromatin in its open form, as euchromatin, or in its closed form, as heterochromatin.[21] Histone acetylation, for instance, is typically associated with euchromatin structure, whereas histone methylation promotes heterochromatin remodeling.[22] Acetylation makes the histone group more negatively charged, which in turn disrupts its interactions with the DNA strand, essentially "opening" the strand for easier access.[21] Acetylation can occur on multiple lysine residues of a histone's N-terminal tail and in different histones of the same nucleosome, which is thought to further increase DNA accessibility for transcription factors.[21]
Another method of regulation that incorporates a negative charge, thereby favoring the "open" form, is
See also
References
- S2CID 186242248.
- ^ PMID 2822591.
- ^ "Definition: nucleosome/nucleosomes". Scitable Nature Education. Retrieved 2021-10-06.
- ISBN 978-0-12-811014-0.
- ^ a b "The cell. 4. Nucleus. Chromatin. Atlas of plant and animal histology". mmegias.webs.uvigo.es. Retrieved 2021-12-02.
- PMID 23582201.
- S2CID 131915096.
- ^ a b "Giemsa banding". Biology Articles, Tutorials & Dictionary Online. 2019-10-07. Retrieved 2021-12-02.
- ^ "Reverse banding - Definition and Examples - Biology Online Dictionary". Biology Articles, Tutorials & Dictionary Online. 2020-09-18. Retrieved 2021-12-02.
- ^ "Constitutive heterochromatin banding". Biology Articles, Tutorials & Dictionary Online. 2019-10-07. Retrieved 2021-12-02.
- ^ "Quinacrine banding". Biology Articles, Tutorials & Dictionary Online. 2019-10-07. Retrieved 2021-12-02.
- ^ "T-banding". Biology Articles, Tutorials & Dictionary Online. 2019-10-07. Retrieved 2021-12-02.
- PMID 10508851.
- PMID 12702201.
- PMID 23810203.
- S2CID 24376600.
- PMID 21252296.
- PMID 27005444.
- ^ Simmons D (2008). "Epigenetic Influences and Disease". Nature Education. 1 (1): 6. Retrieved 2021-12-02.
- PMID 29796022.
- ^ PMID 21321607.
- ^ S2CID 235031860.
Further reading
- Heterochromatin formation involves changes in histone modifications over multiple cell generations – Katan-Khaykovich Y, Struhl K (June 2005). "Heterochromatin formation involves changes in histone modifications over multiple cell generations". The EMBO Journal. 24 (12): 2138–2149. PMID 15920479.
- Chromatin Velocity reveals epigenetic dynamics by single-cell profiling of heterochromatin and euchromatin – Tedesco M, Giannese F, Lazarević D, Giansanti V, Rosano D, Monzani S, et al. (October 2021). "Chromatin Velocity reveals epigenetic dynamics by single-cell profiling of heterochromatin and euchromatin". Nature Biotechnology. 40 (2): 235–244. S2CID 238637962.
- Epigenetic inheritance and the missing heritability – Trerotola M, Relli V, Simeone P, Alberti S (July 2015). "Epigenetic inheritance and the missing heritability". Human Genomics. 9 (1): 17. PMID 26216216.
- Histone epigenetic marks in heterochromatin and euchromatin of the Chagas' disease vector, Triatoma infestans – Alvarenga EM, Rodrigues VL, Moraes AS, Naves LS, Mondin M, Felisbino MB, Mello ML (May 2016). "Histone epigenetic marks in heterochromatin and euchromatin of the Chagas' disease vector, Triatoma infestans". Acta Histochemica. 118 (4): 401–412. PMID 27079857.
