Histone acetylation and deacetylation
Histone acetylation and deacetylation are essential parts of
Acetylated
Mechanism of action
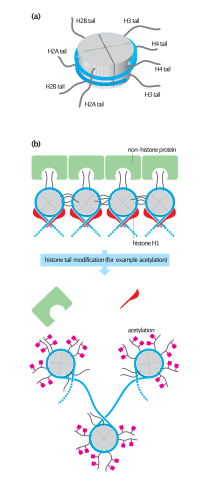
Nucleosomes are portions of double-stranded DNA (dsDNA) that are wrapped around protein complexes called histone cores. These histone cores are composed of 8 subunits, two each of H2A, H2B, H3 and H4 histones. This protein complex forms a cylindrical shape that dsDNA wraps around with approximately 147 base pairs. Nucleosomes are formed as a beginning step for DNA compaction that also contributes to structural support as well as serves functional roles.[2] These functional roles are contributed by the tails of the histone subunits. The histone tails insert themselves in the minor grooves of the DNA and extend through the double helix,[1] which leaves them open for modifications involved in transcriptional activation.[3] Acetylation has been closely associated with increases in transcriptional activation while deacetylation has been linked with transcriptional deactivation. These reactions occur post-translation and are reversible.[3]
The mechanism for acetylation and deacetylation takes place on the NH3+ groups of lysine amino acid residues. These residues are located on the tails of histones that make up the nucleosome of packaged dsDNA. The process is aided by factors known as
Acetylation has the effect of changing the overall charge of the histone tail from positive to neutral. Nucleosome formation is dependent on the positive charges of the H4 histones and the negative charge on the surface of H2A histone fold domains. Acetylation of the histone tails disrupts this association, leading to weaker binding of the nucleosomal components.[1] By doing this, the DNA is more accessible and leads to more transcription factors being able to reach the DNA. Thus, acetylation of histones is known to increase the expression of genes through transcription activation. Deacetylation performed by HDAC molecules has the opposite effect. By deacetylating the histone tails, the DNA becomes more tightly wrapped around the histone cores, making it harder for transcription factors to bind to the DNA. This leads to decreased levels of gene expression and is known as gene silencing.[5][6][7]
Acetylated histones, the octomeric protein cores of nucleosomes, represent a type of epigenetic marker within chromatin. Studies have shown that one modification has the tendency to influence whether another modification will take place. Modifications of histones can not only cause secondary structural changes at their specific points, but can cause many structural changes in distant locations which inevitably affects function.[8] As the chromosome is replicated, the modifications that exist on the parental chromosomes are handed down to daughter chromosomes. The modifications, as part of their function, can recruit enzymes for their particular function and can contribute to the continuation of modifications and their effects after replication has taken place.[1] It has been shown that, even past one replication, expression of genes may still be affected many cell generations later. A study showed that, upon inhibition of HDAC enzymes by Trichostatin A, genes inserted next to centric heterochromatin showed increased expression. Many cell generations later, in the absence of the inhibitor, the increased gene expression was still expressed, showing modifications can be carried through many replication processes such as mitosis and meiosis.[8]
Histone acetylation/deacetylation enzymes

Histone acetyltransferase (HATs)
Histone Acetyltransferases, also known as HATs, are a family of enzymes that acetylate the histone tails of the nucleosome. This, and other modifications, are expressed based on the varying states of the cellular environment.[2] Many proteins with acetylating abilities have been documented and, after a time, were categorized based on sequence similarities between them. These similarities are high among members of a family, but members from different families show very little resemblance.[9] Some of the major families identified so far are as follows.
GNAT family
General Control Non-Derepressible 5 (Gcn5) –related N-Acetyltransferases (GNATs) is one of the many studied families with acetylation abilities.[10] This superfamily includes the factors Gcn5 which is included in the SAGA, SLIK, STAGA, ADA, and A2 complexes, Gcn5L, p300/CREB-binding protein associated factor (PCAF), Elp3, HPA2 and HAT1.[10][11] Major features of the GNAT family include HAT domains approximately 160 residues in length and a conserved bromodomain that has been found to be an acetyl-lysine targeting motif.[9] Gcn5 has been shown to acetylate substrates when it is part of a complex.[11] Recombinant Gcn5 has been found to be involved in the acetylation of the H3 histones of the nucleosome.[2][11] To a lesser extent, it has been found to also acetylate H2B and H4 histones when involved with other complexes.[2][3][11] PCAF has the ability to act as a HAT protein and acetylate histones, it can acetylate non-histone proteins related to transcription, as well as act as a coactivator in many processes including myogenesis, nuclear-receptor-mediated activation and growth-factor-signaled activation. Elp3 has the ability to acetylate all histone subunits and also shows involvement in the RNA polymerase II holoenzyme.[2]
MYST family
MOZ (Monocytic Leukemia Zinc Finger Protein), Ybf2/Sas3, Sas2 and Tip60 (Tat Interacting Protein) all make up MYST, another well known family that exhibits acetylating capabilities. This family includes Sas3, essential SAS-related acetyltransferase (Esa1), Sas2, Tip60, MOF, MOZ, MORF, and HBO1. The members of this family have multiple functions, not only with activating and silencing genes, but also affect development and have implications in human diseases.
p300/CBP family
Adenoviral E1A-associated protein of 300kDa (p300) and the CREB-binding protein (CBP) make up the next family of HATs.[10] This family of HATs contain HAT domains that are approximately 500 residues long and contain bromodomains as well as three cysteine-histidine rich domains that help with protein interactions.[9] These HATs are known to acetylate all of the histone subunits in the nucleosome. They also have the ability to acetylate and mediate non-histone proteins involved in transcription and are also involved in the cell-cycle, differentiation and apoptosis.[2]
Other HATs
There are other proteins that have acetylating abilities but differ in structure to the previously mentioned families. One HAT is called
Histone deacetylase (HDACs)
There are a total of four classes that categorize Histone Deacetylases (HDACs). Class I includes HDACs
Class I HDACs
HDAC1 & HDAC2
Activity of these HDACs can be affected by
HDAC3
HDAC3 has been found to be most closely related to HDAC8. HDAC3 contains a non-conserved region in the C-terminal region that was found to be required for transcriptional repression as well as its deacetylase activity. It also contains two regions, one called a Nuclear Localization Signal (NLS) as well as a Nuclear Export Signal (NES). The NLS functions as a signal for nuclear action while an NES functions with HDACs that perform work outside of the nucleus. A presence of both signals for HDAC3 suggests it travels between the nucleus and the cytoplasm.[5] HDAC3 has even been found to interact with the plasma membrane.[6] Silencing Mediator for Retinoic Acid and Thyroid Hormone (SMRT) receptors and Nuclear Receptor Co-Repressor (N-CoR) factors must be utilized by HDAC3 in order to activate it.[5][6] Upon doing so, it gains the ability to co-precipitate with HDACs 4, 5, and 7. HDAC3 can also be found complexed together with HDAC-related protein (HDRP).[5] HDACs 1 and 3 have been found to mediate Rb-RbAp48 interactions which suggests that it functions in cell cycle progression.[5][6] HDAC3 also shows involvement in stem cell self-renewal and a transcription independent role in mitosis.[6]
HDAC8
HDAC8 has been found to be most similar to HDAC3. Its major feature is its catalytic domain which contains an NLS region in the center. Two transcripts of this HDAC have been found which include a 2.0kb transcript and a 2.4kb transcript.[5] Unlike the other HDAC molecules, when purified, this HDAC showed to be enzymatically active.[6] At this point, due to its recent discovery, it is not yet known if it is regulated by co-repressor protein complexes. Northern blots have revealed that different tissue types show varying degrees of HDAC8 expression[5] but has been observed in smooth muscles and is thought to contribute to contractility.[6]
Class II HDACs
Class IIA
The Class IIA HDACs includes
For HDACs 4, 5 and 7, conserved binding domains have been discovered that bind for
Class IIB
The Class IIB HDACs include HDAC6 and HDAC10. These two HDACs are most closely related to each other in overall sequence. However, HDAC6's catalytic domain is most similar to HDAC9.[5] A unique feature of HDAC6 is that it contains two catalytic domains in tandem of one another.[5][6] Another unique feature of HDAC6 is the HDAC6-, SP3, and Brap2-related zinc finger motif (HUB) domain in the C-terminus which shows some functions related to ubiquitination, meaning this HDAC is prone to degradation.[5] HDAC10 has two catalytic domains as well. One active domain is located in the N-terminus and a putative catalytic domain is located in the C-terminus[5][6] along with an NES domain.[5] Two putative Rb-binding domains have also been found on HDAC10 which shows it may have roles in the regulation of the cell cycle. Two variants of HDAC10 have been found, both having slight differences in length. HDAC6 is the only HDAC to be shown to act on tubulin, acting as a tubulin deacetylase which helps in the regulation of microtubule-dependent cell motility. It is mostly found in the cytoplasm but has been known to be found in the nucleus, complexed together with HDAC11. HDAC10 has been seen to act on HDACs 1, 2, 3 (or SMRT), 4, 5 and 7. Some evidence has been shown that it may have small interactions with HDAC6 as well. This leads researchers to believe that HDAC10 may function more as a recruiter rather than a factor for deacetylation. However, experiments conducted with HDAC10 did indeed show deacetylation activity.[5]
Class IV HDACs
HDAC11
HDAC11 has been shown to be related to HDACs 3 and 8, but its overall sequence is quite different from the other HDACs, leading it to be in its own category.[5][6] HDAC11 has a catalytic domain located in its N-terminus. It has not been found incorporated in any HDAC complexes such as Nurd or SMRT which means it may have a special function unique to itself. It has been found that HDAC11 remains mainly in the nucleus.[5]
Biological functions
Transcription regulation
The discovery of histone acetylation causing changes in
The regulatory mechanism is thought to be twofold. Lysine is an amino acid with a positive charge when unmodified. Lysines on the
Another implication of histone acetylation is to provide a platform for protein binding. As a
Histone code hypothesis
The Histone code hypothesis suggests the idea that patterns of post-translational modifications on histones, collectively, can direct specific cellular functions.[21] Chemical modifications of histone proteins often occur on particular amino acids. This specific addition of single or multiple modifications on histone cores can be interpreted by transcription factors and complexes which leads to functional implications. This process is facilitated by enzymes such as HATs and HDACs that add or remove modifications on histones, and transcription factors that process and "read" the modification codes. The outcome can be activation of transcription or repression of a gene. For example, the combination of acetylation and phosphorylation have synergistic effects on the chromosomes overall structural condensation level and, hence, induces transcription activation of immediate early gene.[22]
Experiments investigating acetylation patterns of H4 histones suggested that these modification patterns are collectively maintained in mitosis and meiosis in order to modify long-term gene expression.[8] The acetylation pattern is regulated by HAT and HADC enzymes and, in turn, sets the local chromatin structure. In this way, acetylation patterns are transmitted and interconnected with protein binding ability and functions in subsequent cell generation.
Bromodomain
The
Human diseases
Inflammatory diseases
Gene expression is regulated by histone acetylation and deacetylation, and this regulation is also applicable to inflammatory genes. Inflammatory lung diseases are characterized by expression of specific inflammatory genes such as
Specifically, gene expression data demonstrated increased activity of HAT and decreased level of HDAC activity in patients with Asthma.[26] Patients with chronic obstructive pulmonary disease showed there is an overall decrease in HDAC activity with unchanged levels of HAT activity.[27] Results have shown that there is an important role for HAT/HDAC activity balance in inflammatory lung diseases and provided insights on possible therapeutic targets.[28]
Cancer
Due to the regulatory role during transcription of epigenetic modifications in genes, it is not surprising that changes in
Approved in 2006 by the U.S.
Carbon source availability is reflected in histone acetylation in cancer. Glucose and glutamine are the major carbon sources of most mammalian cells, and glucose metabolism is closely related to histone acetylation and deacetylation. Glucose availability affects the intracellular pool of acetyl-CoA, a central metabolic intermediate that is also the acetyl donor in histone acetylation. Glucose is converted to acetyl-CoA by the pyruvate dehydrogenase complex (PDC), which produces acetyl-CoA from glucose-derived pyruvate; and by adenosine triphosphate-citrate lyase (ACLY), which generates acetyl-CoA from glucose-derived citrate. PDC and ACLY activity depend on glucose availability, which thereby influences histone acetylation and consequently modulates gene expression and cell cycle progression. Dysregulation of ACLY and PDC contributes to metabolic reprogramming and promotes the development of multiple cancers. At the same time, glucose metabolism maintains the NAD+/NADH ratio, and NAD+ participates in SIRT-mediated histone deacetylation. SIRT enzyme activity is altered in various malignancies, and inhibiting SIRT6, a histone deacetylase that acts on acetylated H3K9 and H3K56, promotes tumorigenesis. SIRT7, which deacetylates H3K18 and thereby represses transcription of target genes, is activated in cancer to stabilize cells in the transformed state. Nutrients appear to modulate SIRT activity. For example, long-chain fatty acids activate the deacetylase function of SIRT6, and this may affect histone acetylation.[33]
Addiction
Epigenetic modifications of histone tails in specific regions of the brain are of central importance in addictions, and much of the work on addiction has focused on histone acetylation.[34][35][36] Once particular epigenetic alterations occur, they appear to be long lasting "molecular scars" that may account for the persistence of addictions.[34][37]
About 7% of the US population is addicted to alcohol. In rats exposed to alcohol for up to 5 days, there was an increase in histone 3 lysine 9 acetylation in the pronociceptin promoter in the brain amygdala complex. This acetylation is an activating mark for pronociceptin. The nociceptin/nociceptin opioid receptor system is involved in the reinforcing or conditioning effects of alcohol.[43]
Cocaine addiction occurs in about 0.5% of the US population. Repeated cocaine administration in mice induces hyperacetylation of histone 3 (H3) or histone 4 (H4) at 1,696 genes in one brain "reward" region [the nucleus accumbens (NAc)] and deacetylation at 206 genes.[44][45] At least 45 genes, shown in previous studies to be upregulated in the NAc of mice after chronic cocaine exposure, were found to be associated with hyperacetylation of H3 or H4. Many of these individual genes are directly related to aspects of addiction associated with cocaine exposure.[45][46]
In rodent models, many agents causing addiction, including tobacco smoke products,[47] alcohol,[48] cocaine,[49] heroin[50] and methamphetamine,[51][52] cause DNA damage in the brain. During repair of DNA damages some individual repair events may alter the acetylations of histones at the sites of damage, or cause other epigenetic alterations, and thus leave an epigenetic scar on chromatin.[37] Such epigenetic scars likely contribute to the persistent epigenetic changes found in addictions.
In 2013, 22.7 million persons aged 12 or older needed treatment for an illicit drug or alcohol use problem (8.6 percent of persons aged 12 or older).[38]
Other disorders
Suggested by the idea that the structure of chromatin can be modified to allow or deny access of
Based on different
Epigenetic modifications also play a role in neurological disorders. Deregulation of histones modification are found to be responsible for deregulated gene expression and hence associated with neurological and psychological disorders, such as
See also
- Histone acetyltransferase
- Histone deacetylase
- Histone methylation
- Acetylation
- Phosphorylation
- Nucleosome
References
- ^ ISBN 978-0-321-76243-6.
- ^ PMID 16877467.
- ^ S2CID 35433573.
- S2CID 4419816.
- ^ PMID 12429021.
- ^ S2CID 30268983.
- PMID 9499396.
- ^ PMID 10944586.
- ^ PMID 11492997.
- ^ S2CID 10662910.
- ^ PMID 14969728.
- PMID 29795555.
- PMID 11250138.
- PMID 14172992.
- PMC 3265906.
- S2CID 23873925.
- PMID 26140054.
- PMID 18804549.
- S2CID 16237908.
- S2CID 22196542.
- PMID 20574448.
- PMID 8197135.
- PMID 19736624.
- S2CID 12172346.
- PMID 15738302.
- PMID 24527405.
- PMID 18204158.
- PMID 15738302.
- S2CID 2976852.
- PMID 21941619.
- PMID 16960145.
- S2CID 262487540.
- PMID 29784032.
- ^ PMID 21989194.
- )
- PMID 20425300.
- ^ PMID 27259203.
- ^ a b Substance Abuse and Mental Health Services Administration, Results from the 2013 National Survey on Drug Use and Health: Summary of National Findings, NSDUH Series H-48, HHS Publication No. (SMA) 14-4863. Rockville, MD: Substance Abuse and Mental Health Services Administration, 2014
- ^ "Is nicotine addictive?".
- PMID 22049069.
- S2CID 19157711.
- PMID 11572966.
- S2CID 14013417.
- )
- ^ PMID 19447090.
- ^ https://www.drugsandalcohol.ie/12728/1/NIDA_Cocaine.pdf [bare URL PDF]
- PMID 28912356.
- PMID 18482162.
- S2CID 20849951.
- PMID 16076083.
- S2CID 24182756.
- PMID 18797138.
- PMID 19568876.
- PMID 21367693.
- PMID 12202037.
- PMID 24310814.
- PMID 24693336.
- PMID 14517255.
- PMID 22832356.
- PMID 24006238.
- S2CID 3112549.
External links
- Animation of histone tail acetylation and deacetylation: [1]
