Eukaryotic translation
Eukaryotic translation is the biological process by which
Initiation

Translation initiation is the process by which the ribosome and its associated factors bind to an mRNA and are assembled at the start codon. This process is defined as either cap-dependent, in which the ribosome binds initially at the 5' cap and then travels to the stop codon, or as cap-independent, where the ribosome does not initially bind the 5' cap.
Cap-dependent initiation
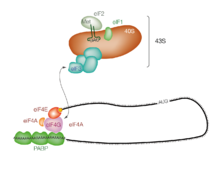
Initiation of translation usually involves the interaction of certain key proteins, the
This
Regulation of protein synthesis is partly influenced by phosphorylation of eIF2 (via the α subunit), which is a part of the eIF2-GTP-Met-tRNAiMet ternary complex (eIF2-TC). When large numbers of eIF2 are phosphorylated, protein synthesis is inhibited. This occurs under amino acid starvation or after viral infection. However, a small fraction of this initiation factor is naturally phosphorylated. Another regulator is 4EBP, which binds to the initiation factor eIF4E and inhibits its interactions with eIF4G, thus preventing cap-dependent initiation. To oppose the effects of 4EBP, growth factors phosphorylate 4EBP, reducing its affinity for eIF4E and permitting protein synthesis.[citation needed]
While protein synthesis is globally regulated by modulating the expression of key initiation factors as well as the number of ribosomes, individual mRNAs can have different translation rates due to the presence of regulatory sequence elements. This has been shown to be important in a variety of settings including yeast meiosis and ethylene response in plants. In addition, recent work in yeast and humans suggest that evolutionary divergence in cis-regulatory sequences can impact translation regulation.[4] Additionally, RNA helicases such as DHX29 and Ded1/DDX3 may participate in the process of translation initiation, especially for mRNAs with structured 5'UTRs.[5]
Cap-independent initiation
The best-studied example of cap-independent translation initiation in eukaryotes uses the
Elongation

Elongation depends on
Termination
Termination of elongation depends on
Regulation and modification of translation
Translation is one of the key energy consumers in cells, hence it is strictly regulated. Numerous mechanisms have evolved that control and regulate translation in eukaryotes as well as prokaryotes. Regulation of translation can impact the global rate of protein synthesis which is closely coupled to the metabolic and proliferative state of a cell. To delve deeper into this intricate process, scientists typically use a technique known as ribosome profiling.[9] This method enables researchers to take a snapshot of the translatome, showing which parts of the mRNA are being translated into proteins by ribosomes at a given time. Ribosome profiling provides valuable insights into translation dynamics, revealing the complex interplay between gene sequence, mRNA structure, and translation regulation. Expanding on this concept, a more recent development is single-cell ribosome profiling, a technique that allows us to study the translation process at the resolution of individual cells.[10] Single-cell ribosome profiling has the potential to shed light on the heterogeneous nature of cells, leading to a more nuanced understanding of how translation regulation can impact cell behavior, metabolic state, and responsiveness to various stimuli or conditions.
Amino acid substitution
In some cells certain amino acids can be depleted and thus affect translation efficiency. For instance, activated T cells secrete interferon-γ which triggers intracellular tryptophan shortage by upregulating the indoleamine 2,3-dioxygenase 1 (IDO1) enzyme. Surprisingly, despite tryptophan depletion, in-frame protein synthesis continues across tryptophan codons. This is achieved by incorporation of phenylalanine instead of tryptophan. The resulting peptides are called W>F "substitutants". Such W>F substitutants are abundant in certain cancer types and have been associated with increased IDO1 expression. Functionally, W>F substitutants can impair protein activity.[11]
See also
- 40S
- 60S
- 80S
- Eukaryotic initiation factor
- Eukaryotic elongation factors
- Eukaryotic release factors
