Discovery and development of NS5A inhibitors

Hepatitis C virus
HCV is a
HCV is among the leading causes of liver disease around the world. It is transmitted by blood and is most commonly contracted through the use of infected needles.[10] Patients with chronic HCV infection are at significant risk of cirrhosis and hepatocellular carcinoma, which are the leading causes of death for those infected.[11][12]
The virus has been around for over a millennia and has been classified into six known genotypes, each of which contains numerous subtypes. The seventh remains uncharacterized. The genotype contracted dictates which specific treatments are viable.[13]
NS5A receptor
Basic structure and chemical properties
NS5A is a large hydrophilic phosphoprotein that is essential for the HCV life cycle and is found in association with virus-induced membrane vesicles, termed the membranous web.[14][15] NS5A is a proline-rich protein composed of approximately 447 amino acids, which is divided into three domains.[16][17] These domains are linked by two low-complexity sequences that are either serine- or proline-rich.[18] Domain I is a zinc binding domain and X-ray crystallography studies indicated alternative dimer conformations of domain I of NS5A.[19][20][21] Domain II and III are unstructured, shown by NMR studies.[16][22] Domain I is preceded by an N-terminal amphipathic helix which allows the protein to associate with endoplasmic reticulum-derived membranes.[22][23][24] Although X-ray crystallographic studies revealed dimer conformations of NS5A domain1, recent in solution structural characterization studies showed that NS5A proteins form higher-order structures by dimeric subunits of NS5A domain 1.[25] Moreover, the overall structural model of NS5A highlights the variability of intrinsic conformations of the D2 and D3 domains between HCV genotypes.[26] Therefore, it is still under debate which conformation\s of NS5A is functional and also targeted by NS5A inhibitors.[citation needed]
NS5A mainly exists in two distinct phosphorylated forms, a hypophosphorylated and a hyperphosphorylated form, but the exact function of the phosphorylation has not been determined.[17][18][27]
Function
The NS5A protein plays an important role in viral RNA replication, viral assembly, and complex interactions with cellular functions.[2][17] The protein has been implicated in the modulation of host defenses, apoptosis, the cell cycle, and stress-responsive pathways.[27] However, its function and complete structure have yet to be elucidated.[16]
NS5A seems to be key in triggering the formation of the membranous web in the absence of other similar nonstructural proteins.[15] Many proteins within the host cell can be affected by NS5A, e.g. phosphatidylinositol 4-kinase IIIα (PI4KIIIα), a kinase required for the replication of HCV. This kinase takes part in the biosynthesis of phosphatidylinositol 4-phosphate (PI4P) by interacting with NS5A, which stimulates its activity and appears to improve the integrity of the membranous web.[15][28][29]
Recently, the central role of NS5A in viral proliferation has made it the target for drug development. As a result, new antiviral agents have been introduced for the treatment of HCV.[2]
Mechanism of action
NS5A inhibitors have been developed to target the NS5A protein. These inhibitors have achieved a significant reduction in HCV RNA blood levels and can therefore be considered as potent antivirals. Their mechanism of action is thought to be diverse but the exact mechanism is not fully understood.[2][30] Most studies assume that NS5A inhibitors act on two essential stages of the HCV life cycle; the replication of the genomic RNA, and virion assembly. Other studies propose an alteration of host cell factors as a possible third mechanism.[2][15][31]
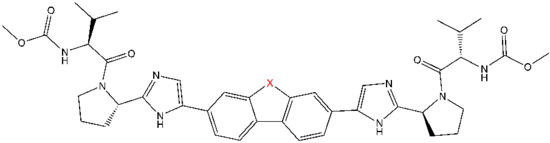
The structure of NS5A inhibitors is characterized by dimeric symmetry. This suggests that NS5A inhibitors act on dimers of NS5A.[32] A number of modeling studies have shown that daclatasvir, which is an NS5A inhibitor, only binds to the "back-to-back" NS5A dimer and that the binding has to be symmetrical. Other modeling studies have shown that binding to other conformations of NS5A might be possible, as well as asymmetrical binding.[30] Research has shown that daclatasvir's target is most likely domain I of NS5A.[31] Even though the mechanism is not completely understood, it has been demonstrated that the inhibitors downregulate NS5A hyperphosphorylation, leading to the suppression of HCV replication and its processing of polyproteins, as well as resulting in an unusual protein location.[31][33] Hitherto, this inhibition was thought to require only NS5A domain I, but not domains II and III.[33] However, recent studies have shown that both domains I and II are relevant to this disruption of RNA replication.[34]
NS5A inhibitors appear to furthermore disrupt the formation of new replicase complexes resulting in a gradual slowing of viral RNA synthesis. Effect on previously formed complexes has yet to be demonstrated.[34][35]
Available evidence suggests that NS5A inhibitors modify the location of NS5A inside the cell. This may cause abnormal assembly leading to malformed viruses.[2] Some studies have revealed that inhibition of the viral assembly has a more important role in RNA reduction than viral replication reduction.[35][36]
Studies have shown that NS5A inhibitors block the formation of the membranous web, which protects the viral genome and features the main sites for viral replication and assembly.[15][31][37] This mechanism is thought to be independent of RNA replication, but seems to be affected by NS5A inhibitors blocking the formation of the PI4KIIIα-NS5A complex, essential to the synthesis of the PI4P, resulting in decreased integrity of the membranous web and therefore reduced HCV RNA replication.[15][28][38]
History
HCV research has taken great strides in recent years with the discovery and clinical development of multiple new HCV drugs. Among those drugs are the DAAs which include NS5A inhibitors.[39] NS5A inhibitors have been found particularly effective in the treatment of HCV where they have been used in combination with other protease inhibitors such as NS5B inhibitors (e.g. sofosbuvir), pegylated interferons (e.g. peginterferon alfa-2a), and ribonucleic analogs (e.g. ribavirin).[40][41][42] The ever present risk of viral strains developing resistance has been a main factor in why they are used in combination with one or more complementary drug.[43]
Adverse effects, and extensive and complicated drug regimens with accompanying low compliance rates, have been a hindrance in the development of antiviral treatments. The combination of NS5A and NS5B inhibitors has produced positive results in this regard.[44]
Drug discovery and development
Discovery
The discovery of NS5A inhibitors took place within the context of a pursuit for a treatment for HCV. NS5A is among the seven nonstructural proteins that form a complex with viral RNA within infected cells to initiate HCV replication.[45] HCV research has produced several DAAs including NS3A, NS4A and NS5B inhibitors, as well as NS5A inhibitors.[46]
Development
The development of antiviral drugs capable of interfering with the proteins responsible for viral replication has been intimately linked with advancements in techniques for establishing the efficient cell culture systems needed to screen for them.[46]
In 1999 a breakthrough came when a full-length consensus genome cloned from HCV RNA was found to replicate at high levels when transfected into a human hepatoma cell line.[47] This method has since been improved upon with the use of cell culture-adaptive mutations that enhance RNA replication.[48]
Screening has now produced a number of NS5A inhibitors, which have been incorporated into treatments for HCV. The first in this new class of drugs was daclatasvir (Daklinza), gaining first global approval from the Japanese Ministry of Health, Labour and Welfare (MHLW) in July 2014 in combination with asunaprevir.[49] Daclatasvir received FDA approval in July 2015.[50] Other drugs have since been approved, among them notably the first FDA-approved NS5A inhibitor ledipasvir, approved October 2014 in combination with sofosbuvir to comprise the HCV drug Harvoni.[51][52]
Although NS5A inhibitors have proven effective antivirals, they must be used alongside complementary antiviral drugs due to how quickly they lead to the development of resistant mutations when given as a single agent.[53] This has shaped the focus of NS5A inhibitor development, from which asymmetrical variants that metabolize into analogues with complementary resistance profiles have emerged, amongst other discoveries.[54]
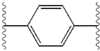
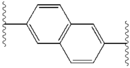
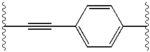
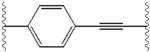
Structure-activity relationship
The structural similarities between the inhibitors are readily apparent.
Favorable characteristics in an NS5A inhibitor include high potency and long plasma
| Structure | Activity | |
|---|---|---|
| X | IC50 (nM) | Inhibitory activity |
| >44 | None | |
| >44 | None | |
| 11 | Very weak | |
| 1.7 | Weak | |
| 0.50 | Moderate | |
| 3.7 | Weak | |
| 0.11 | Moderate | |
| 0.20 | Moderate | |
| Structure | Activity | |
|---|---|---|
| X | IC50 (nM) | Inhibitory activity |
| >44 | None | |
| 0.071 | Moderate | |
| 2.5 | Weak | |
| 0.38 | Moderate | |
| 0.20 | Moderate | |
| 0.17 | Moderate | |
| 0.040 | Strong | |
| Structure | Activity | |
|---|---|---|
| X | IC50 (nM) | Inhibitory activity |
| CH2 | 0.094 | Strong |
| CO | 0.30 | Moderate |
| C(CH3)2 | 1.2 | Weak |
Resistance
The potential HCV resistance against DAA drugs is a concern.[6] Among the HCV quasispecies there are pre-existing variants with the potential to confer resistance to NS5A inhibitors without having any previous exposure to those drugs. Generally, the replication of these variants happens only in minute quantities, making them undetectable by current techniques. On the other hand, it is possible to selectively grow immune variants in the presence of NS5A inhibitors.[2] HCV resistance is characterized by a certain escape pattern. This pattern is often associated with amino acid substitutions that confer upon the virus a robust drug resistance without impairing the viral fitness.[2][57] It has been established that NS5A inhibitors possess a relatively low threshold for resistance, and variants that are associated with NS5A resistance have been shown to endure for up to six months in patients following treatment cessation.[58] Therefore, combination therapies produce higher efficacy and shorter treatment periods.[7]
Future research and new generations of NS5A inhibitors
DAA developers face foreseeable challenges in the years to come. Therapeutic gaps for individuals with complicating conditioned such as chronic kidney disease and cirrhosis will need to be bridged. Shorter therapies with milder side effects would yield greater adherence, and the ever present spectre of drug resistance is looming. The highly adaptive HCV has evolved into a number of different genomes that all need to be adequately treated, preferably with pan-genotypic regimens.[59]
Some of these challenges already have possible solutions in sight. The protease inhibitor ABT-493 and the next-generation NS5A inhibitor ABT-530 are considered active against all HCV genotypes, including the hard to treat genotype 3.[42][59] In vitro, ABT-530 showed potency against the resistance associated variants which are immune to the first generations of NS5A inhibitors, including ledipasvir, daclatasvir and ombitasvir.[42] Because this drug combination has the additional quality of being hepatically cleared, it holds the promise that patients with chronic kidney disease and HCV could receive a safe, non-sofosbuvir-based treatment in the near future.[59]
At least three drug combinations for the treatment of HCV are in the pipeline to be approved in 2016-2017: Sofosbuvir in combination with velpatasvir, ABT-493 in combination with ABT-530, and grazoprevir in combination with elbasvir, of which velpatasvir, ABT-530 and elbasvir are NS5A inhibitors.[7]
See also
References
- PMID 25761432.
- ^ PMID 23567084.
- PMID 26312999.
- ^ PMID 27634194.
- ^ PMID 21576451.
- ^ PMID 20585111.
- ^ PMID 26725897.
- PMID 21604986.
- PMID 7679746.
- PMID 24863270.
- PMID 12407575.
- S2CID 23226526.
- PMID 18971279.
- PMID 18032500.
- ^ PMID 25224012.
- ^ PMID 22239820.
- ^ PMID 24760886.
- ^ PMID 24257600.
- PMID 15902263.
- PMID 19244328.
- PMID 24639329.
- ^ PMID 20592076.
- PMID 21593143.
- ^ PMID 24755925.
- PMID 29300159.
- PMID 28535337.
- ^ PMID 25972535.
- ^ PMID 21238945.
- PMID 21297162.
- ^ PMID 27933783.
- ^ PMID 25384189.
- PMID 24639329.
- ^ PMID 21795470.
- ^ PMID 24621191.
- ^ PMID 23431163.
- PMID 24768676.
- PMID 26863439.
- PMID 25046163.
- ^ PMID 26904396.
- PMID 24733478.
- PMID 27348813.
- ^ PMID 26778412.
- PMID 19263475.
- ^ Archer, Melissa; Steinvoort, Carin; Oderda, Gary. "UPDATE: New Hepatitis C Combination Agents" (PDF). Utah Department of Health. Archived from the original (PDF) on 27 December 2016. Retrieved 8 September 2016.
- PMID 11110665.
- ^ PMID 19216075.
- PMID 10390360.
- PMID 12584326.
- S2CID 207488399.
- ^ "FDA approves new treatment for chronic hepatitis C genotype 3 infections". FDA. 27 July 2015. Retrieved 29 September 2016.
- ^ PMID 24320933.
- S2CID 31943736.
- PMID 24659881.
- PMID 26099532.
- ^ PMID 24900306.
- PMID 22507961.
- PMID 22851970.
- PMID 27503644.
- ^ PMID 27641983.