Propionyl-CoA

| |
| Names | |
|---|---|
| IUPAC name
3′-O-Phosphonoadenosine 5′-{(3R)-3-hydroxy-2,2-dimethyl-4-oxo-4-[(3-oxo-3-{[2-(propanoylsulfanyl)ethyl]amino}propyl)amino]butyl dihydrogen diphosphate}
| |
| Systematic IUPAC name
O1-{[(2R,3S,4R,5R)-5-(6-Amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)oxolan-2-yl]methyl} O3-{(3R)-3-hydroxy-2,2-dimethyl-4-oxo-4-[(3-oxo-3-{[2-(propanoylsulfanyl)ethyl]amino}propyl)amino]butyl} dihydrogen diphosphate | |
| Other names
Propionyl Coenzyme A; Propanoyl Coenzyme A
| |
| Identifiers | |
3D model (
JSmol ) |
|
| ChEBI | |
| ChemSpider | |
ECHA InfoCard
|
100.005.698 |
| MeSH | propionyl-coenzyme+A |
PubChem CID
|
|
| UNII | |
CompTox Dashboard (EPA)
|
|
| |
| |
| Properties | |
| C24H40N7O17P3S | |
| Molar mass | 823.60 g/mol |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |
Propionyl-CoA is a
Production
There are several different pathways through which propionyl-CoA can be produced:
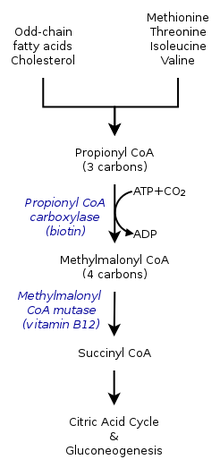
- Propionyl-CoA, a three-carbon structure, is considered to be a minor species of propionic acid. Therefore, odd-number chains of fatty acids are oxidized to yield both propionyl-CoA as well as acetyl-CoA. Propionyl-CoA is later converted into succinyl-CoA through biotin-dependant propionyl-CoA carboxylase (PCC) and b12-dependant methylmalonyl-CoA mutase (MCM), sequentially.[2]
- Propionyl-CoA is not only produced from the oxidation of odd-chain fatty acids, but also by the oxidation of amino acids including methionine, valine, isoleucine, and threonine. Furthermore, catabolism of amino acids can also be a result of the conversion of propionyl-CoA to methylmalonyl-CoA by propionyl-CoA carboxylase.[1]
- Cholesterol oxidation, which forms bile acids, also forms propionyl-CoA as a side product. In an experiment performed by Suld et al., when combining liver mitochondria and propionic acid with the addition of coenzyme A, labeled isotopes of psionic acid were degraded. However, following 5β-cholestane-3α,7α,12α,26-tetrol-26,27-C14 incubation, propionyl CoA was able to be rescued along with the formation of bile.[6]
Metabolic fate
The metabolic (catabolic fate) of propionyl-CoA depends on what environment it is being synthesized in. Therefore, propionyl-CoA in an anaerobic environment could have a different fate than that in an aerobic organism. The multiple pathways, either catabolism by propionyl-CoA carboxylase or methylcitrate synthase, also depend on the presence of various genes.[7]
Reaction with propionyl-CoA carboxylase
Within the citric acid cycle in humans, propionyl-CoA, which interacts with oxaloacetate to form methylcitrate, can also catalyzed into methylmalonyl-CoA through
Mechanism
In mammals, propionyl-CoA is converted to (S)-methylmalonyl-CoA by propionyl-CoA carboxylase, a biotin-dependent enzyme also requiring bicarbonate and ATP.
This product is converted to (R)-methylmalonyl-CoA by
(R)-Methylmalonyl-CoA is converted to

The methylmalonyl-CoA mutase mechanism begins with the cleavage of the bond between the 5' CH
2- of 5'-deoxyadenosyl and the cobalt, which is in its 3+ oxidation state (III), which produces a 5'-deoxyadenosyl radical and cobalamin in the reduced Co(II) oxidation state.
Next, this radical abstracts a hydrogen atom from the methyl group of methylmalonyl-CoA, which generates a methylmalonyl-CoA radical. It is believed that this radical forms a carbon-cobalt bond to the coenzyme, which is then followed by the rearrangement of the substrate's carbon skeleton, thus producing a succinyl-CoA radical. This radical then goes on to abstract a hydrogen from the previously produced 5'-deoxyadenosine, again creating a deoxyadenosyl radical, which attacks the coenzyme to reform the initial complex.
A defect in methylmalonyl-CoA mutase enzyme results in

Methylcitrate cycle
Propionyl-CoA accumulation can prove toxic to different organisms. Since different cycles have been proposed regarding how propionyl-CoA is transformed into pyruvate, one studied mechanism is the methylcitrate cycle. The initial reaction is beta-oxidation to form the propionyl-CoA which is further broken down by the cycle. This pathway involves the enzymes both related to the methylcitrate cycle as well as the citric acid cycle. These all contribute to the overall reaction to detoxify the bacteria from harmful propionyl-CoA. It is also attributed as a resulting pathway due to the catabolism of fatty acids in mycobacteria.[3] In order to proceed, the prpC gene codes for methylcitrate synthase, and if not present, the methylcitrate cycle will not occur. Instead, catabolism proceeds through propionyl-CoA carboxylase.[7] This mechanism is shown below to the left along with the participating reactants, products, intermediates, and enzymes.
Bacterial metabolism
Mycobacterium tuberculosis metabolism
The oxidation of propionyl-CoA to form pyruvate is influenced by its necessity in
Possible sequestration in R. sphaeroides
Propionyl-CoA has can have many adverse and toxic affects on different species, including bacterium. For example, inhibition of pyruvate dehydrogenase by an accumulation of propionyl-CoA in Rhodobacter sphaeroides can prove deadly. Furthermore, as with E. coli, an influx of propionyl-CoA in Myobacterial species can result in toxicity if not dealt with immediately. This toxicity is caused by a pathway involving the lipids that form the bacterial cell wall. Using esterification of long-chain fatty acids, excess propionyl-CoA can be sequestered and stored in the lipid, triacylglycerol (TAG), leading to regulation of elevated propionyl-CoA levels. Such a process of methyl branching of the fatty acids causes them to act as sinks for accumulating propion [4]
Escherichia coli metabolism
In an investigation performed by Luo et al., Escherichia coli strains were utilized to examine how the metabolism of propionyl-CoA could potentially lead to the production of 3-hydroxypropionic acid (3-HP). It was shown that a mutation in a key gene involved in the pathway, succinate CoA-transferase, led to a significant increase in 3-HP.[7] However, this is still a developing field and information on this topic is limited.[12]

Plant metabolism
Amino acid metabolism in plants has been deemed a controversial topic, due to the lack of concrete evidence for any particular pathway. However, it has been suggested that enzymes related to the production and use of propionyl-CoA are involved. Associated with this is the metabolism of isobutyryl-CoA. These two molecules are deemed to be intermediates in valine metabolism. As propionate consists in the form of propionyl-CoA, it was discovered that propionyl-CoA is converted to β-hydroxypropionate through a peroxisomal enzymatic β-oxidation pathway. Nevertheless, in the plant Arabidopsis, key enzymes in the conversion of valine to propionyl-CoA were not observed. Through different experiments performed by Lucas et al., it has been suggested that in plants, through peroxisomal enzymes, propionyl-CoA (and isobutyryl-CoA) are involved in the metabolism of many different substrates (currently being evaluated for identity), and not just valine.[13]

Fungi metabolism
Propionyl-CoA production through the
Protein Propionylation
Propionyl-CoA is also a substrate for post-translational modification of proteins by reacting with lysine residues on proteins, a reaction called protein propionylation.[15][16] Due to structural similarities of Acetyl-CoA and Propionyl-CoA, propionylation reaction are thought to use many of the same enzymes used for protein acetylation.[16] Although functional consequences of protein propionylation are currently not completely understood, in vitro propionylation of the Propionyl-CoA Synthetase enzyme controls its activity.[17]
Human and clinical significance

Gen5
Similar to how plant peroxisomal enzymes bind propionyl-CoA and isobutyryl-CoA, Gen5, an
Propionic acidemia
In the
References
- ^ ISBN 9780128164297.
- ^ PMID 29033250.
- ^ PMID 18048912.
- ^ PMID 29458664.
- ^ PMID 22593918. Retrieved 2019-06-13.
- PMID 13918291.
- ^ PMID 27227837.
- PMID 2647392.
- PMID 30745367.
- PMID 31032474.
- PMID 16689789.
- PMID 23435886.
- PMID 17580301.
- PMID 15514053.
- PMID 17267393.
- ^ PMID 18753126.
- PMID 17684016.
- PMID 27377381.
