Mitochondrion
| Mitochondrion | |
|---|---|
 Diagram of an animal mitochondrion | |
| Details | |
| Pronunciation | /ˌmaɪtəˈkɒndriən/[1] |
| Part of | Cell |
| Identifiers | |
| Latin | organella |
| MeSH | D008928 |
| FMA | 63835 |
| Anatomical terms of microanatomy | |
Animal cell diagram | |
|---|---|
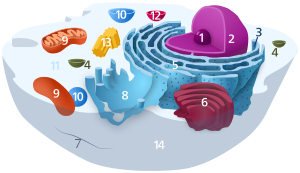 Components of a typical animal cell:
|
A mitochondrion (pl. mitochondria) is an
Some cells in some multicellular organisms lack mitochondria (for example, mature mammalian red blood cells). The multicellular animal Henneguya salminicola is known to have retained mitochondrion-related organelles in association with a complete loss of their mitochondrial genome.[5][6][7] A large number of unicellular organisms, such as microsporidia, parabasalids and diplomonads, have reduced or transformed their mitochondria into other structures,[8] e.g. hydrogenosomes and mitosomes.[9] The oxymonads Monocercomonoides, Streblomastix, and Blattamonas have completely lost their mitochondria.[5][10]
Mitochondria are commonly between 0.75 and 3 μm2 in cross section,[11] but vary considerably in size and structure. Unless specifically stained, they are not visible. In addition to supplying cellular energy, mitochondria are involved in other tasks, such as signaling, cellular differentiation, and cell death, as well as maintaining control of the cell cycle and cell growth.[12] Mitochondrial biogenesis is in turn temporally coordinated with these cellular processes.[13][14] Mitochondria have been implicated in several human disorders and conditions, such as mitochondrial diseases,[15] cardiac dysfunction,[16] heart failure[17] and autism.[18]
The number of mitochondria in a cell can vary widely by organism, tissue, and cell type. A mature red blood cell has no mitochondria,[19] whereas a liver cell can have more than 2000.[20][21] The mitochondrion is composed of compartments that carry out specialized functions. These compartments or regions include the outer membrane, intermembrane space, inner membrane, cristae, and matrix.
Although most of a eukaryotic cell's DNA is contained in the cell nucleus, the mitochondrion has its own genome ("mitogenome") that is substantially similar to bacterial genomes.[22] This finding has led to general acceptance of the endosymbiotic hypothesis - that free-living prokaryotic ancestors of modern mitochondria permanently fused with eukaryotic cells in the distant past, evolving such that modern animals, plants, fungi, and other eukaryotes are able to respire to generate cellular energy.[23]
Structure
| Cell biology | |
|---|---|
| mitochondrion | |
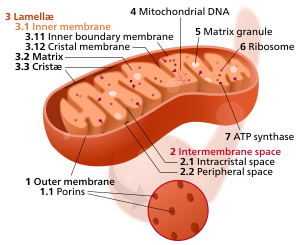 Components of a typical mitochondrion
1 Outer membrane
3 Lamella
4 Mitochondrial DNA |
Mitochondria may have a number of different shapes. The two membranes have different properties. Because of this double-membraned organization, there are five distinct parts to a mitochondrion:
- The outer mitochondrial membrane,
- The intermembrane space (the space between the outer and inner membranes),
- The inner mitochondrial membrane,
- The cristae space (formed by infoldings of the inner membrane), and
- The matrix (space within the inner membrane), which is a fluid.
Mitochondria have folding to increase surface area, which in turn increases ATP (adenosine triphosphate) production. Mitochondria stripped of their outer membrane are called mitoplasts.
Outer membrane
The outer mitochondrial membrane, which encloses the entire organelle, is 60 to 75
The outer membrane also contains
Intermembrane space
The mitochondrial intermembrane space is the space between the outer membrane and the inner membrane. It is also known as perimitochondrial space. Because the outer membrane is freely permeable to small molecules, the concentrations of small molecules, such as ions and sugars, in the intermembrane space is the same as in the cytosol.[20] However, large proteins must have a specific signaling sequence to be transported across the outer membrane, so the protein composition of this space is different from the protein composition of the cytosol. One protein that is localized to the intermembrane space in this way is cytochrome c.[29]
Inner membrane
The inner mitochondrial membrane contains proteins with three types of functions:[20]
- Those that perform the electron transport chain redox reactions
- ATP synthase, which generates ATP in the matrix
- Specific transport proteins that regulate metabolite passage into and out of the mitochondrial matrix
It contains more than 151 different
Cristae

The inner mitochondrial membrane is compartmentalized into numerous folds called cristae, which expand the surface area of the inner mitochondrial membrane, enhancing its ability to produce ATP. For typical liver mitochondria, the area of the inner membrane is about five times as large as that of the outer membrane. This ratio is variable and mitochondria from cells that have a greater demand for ATP, such as muscle cells, contain even more cristae. Mitochondria within the same cell can have substantially different crista-density, with the ones that are required to produce more energy having much more crista-membrane surface.[35] These folds are studded with small round bodies known as F1 particles or oxysomes.[36]
Matrix
The matrix is the space enclosed by the inner membrane. It contains about 2/3 of the total proteins in a mitochondrion.
Function
The most prominent roles of mitochondria are to produce the energy currency of the cell,
Energy conversion
A dominant role for the mitochondria is the production of ATP, as reflected by the large number of proteins in the inner membrane for this task. This is done by oxidizing the major products of
Pyruvate and the citric acid cycle
In the citric acid cycle, all the intermediates (e.g.
Acetyl-CoA, on the other hand, derived from pyruvate oxidation, or from the
In the liver, the
The enzymes of the citric acid cycle are located in the mitochondrial matrix, with the exception of
O2 and NADH: energy-releasing reactions

The electrons from NADH and FADH2 are transferred to oxygen (O2) and hydrogen (protons) in several steps via an electron transport chain. NADH and FADH2 molecules are produced within the matrix via the citric acid cycle and in the cytoplasm by
The major energy-releasing reactions
releasing a lot of
The
While the reactions are controlled by an electron transport chain, free electrons are not amongst the reactants or products in the three reactions shown and therefore do not affect the free energy released, which is used to pump protons (H+) into the intermembrane space. This process is efficient, but a small percentage of electrons may prematurely reduce oxygen, forming reactive oxygen species such as superoxide.[21] This can cause oxidative stress in the mitochondria and may contribute to the decline in mitochondrial function associated with aging.[45]
As the proton concentration increases in the intermembrane space, a strong electrochemical gradient is established across the inner membrane. The protons can return to the matrix through the ATP synthase complex, and their potential energy is used to synthesize ATP from ADP and inorganic phosphate (Pi).[21] This process is called chemiosmosis, and was first described by Peter Mitchell,[46][47] who was awarded the 1978 Nobel Prize in Chemistry for his work. Later, part of the 1997 Nobel Prize in Chemistry was awarded to Paul D. Boyer and John E. Walker for their clarification of the working mechanism of ATP synthase.[48]
Heat production
Under certain conditions, protons can re-enter the mitochondrial matrix without contributing to ATP synthesis. This process is known as proton leak or
Mitochondrial fatty acid synthesis
Mitochondrial fatty acid synthesis (mtFASII) is essential for cellular respiration and mitochondrial biogenesis.[50] It is also thought to play a role as a mediator in intracellular signaling due to its influence on the levels of bioactive lipids, such as lysophospholipids and sphingolipids.[51]
Other products of mtFASII play a role in the regulation of mitochondrial translation,
Furthermore, with the help of mtFASII and acylated ACP, acetyl-CoA regulates its consumption in mitochondria.[52]
Uptake, storage and release of calcium ions
The concentrations of free calcium in the cell can regulate an array of reactions and is important for
Ca2+ influx to the mitochondrial matrix has recently been implicated as a mechanism to regulate respiratory
Cellular proliferation regulation
The relationship between cellular proliferation and mitochondria has been investigated. Tumor cells require ample ATP to synthesize bioactive compounds such as lipids, proteins, and nucleotides for rapid proliferation.[65] The majority of ATP in tumor cells is generated via the oxidative phosphorylation pathway (OxPhos).[66] Interference with OxPhos cause cell cycle arrest suggesting that mitochondria play a role in cell proliferation.[66] Mitochondrial ATP production is also vital for cell division and differentiation in infection [67] in addition to basic functions in the cell including the regulation of cell volume, solute concentration, and cellular architecture.[68][69][70] ATP levels differ at various stages of the cell cycle suggesting that there is a relationship between the abundance of ATP and the cell's ability to enter a new cell cycle.[71] ATP's role in the basic functions of the cell make the cell cycle sensitive to changes in the availability of mitochondrial derived ATP.[71] The variation in ATP levels at different stages of the cell cycle support the hypothesis that mitochondria play an important role in cell cycle regulation.[71] Although the specific mechanisms between mitochondria and the cell cycle regulation is not well understood, studies have shown that low energy cell cycle checkpoints monitor the energy capability before committing to another round of cell division.[12]
Additional functions
Mitochondria play a central role in many other metabolic tasks, such as:
- Signaling through mitochondrial reactive oxygen species[72]
- Regulation of the membrane potential[21]
- Apoptosis-programmed cell death[73]
- Calcium signaling (including calcium-evoked apoptosis)[74]
- Regulation of cellular metabolism[12]
- Certain heme synthesis reactions[75] (see also: Porphyrin)
- Steroid synthesis[56]
- Hormonal signaling[76] – mitochondria are sensitive and responsive to hormones, in part by the action of mitochondrial estrogen receptors (mtERs). These receptors have been found in various tissues and cell types, including brain[77] and heart[78]
- Immune signaling[79]
- Neuronal mitochondria also contribute to cellular quality control by reporting neuronal status towards microglia through specialised somatic-junctions.[80]
- Mitochondria of developing neurons contribute to intercellular signaling towards microglia, which communication is indispensable for proper regulation of brain development.[81]
Some mitochondrial functions are performed only in specific types of cells. For example, mitochondria in
Mitochondrial proteins (proteins transcribed from mitochondrial DNA) vary depending on the tissue and the species. In humans, 615 distinct types of proteins have been identified from cardiac mitochondria,[82] whereas in rats, 940 proteins have been reported.[83] The mitochondrial proteome is thought to be dynamically regulated.[84]
Organization and distribution
Mitochondria (or related structures) are found in all eukaryotes (except the Oxymonad Monocercomonoides).[5] Although commonly depicted as bean-like structures they form a highly dynamic network in the majority of cells where they constantly undergo fission and fusion. The population of all the mitochondria of a given cell constitutes the chondriome.[85] Mitochondria vary in number and location according to cell type. A single mitochondrion is often found in unicellular organisms, while human liver cells have about 1000–2000 mitochondria per cell, making up 1/5 of the cell volume.[20] The mitochondrial content of otherwise similar cells can vary substantially in size and membrane potential,[86] with differences arising from sources including uneven partitioning at cell division, leading to extrinsic differences in ATP levels and downstream cellular processes.[87] The mitochondria can be found nestled between myofibrils of muscle or wrapped around the sperm flagellum.[20] Often, they form a complex 3D branching network inside the cell with the cytoskeleton. The association with the cytoskeleton determines mitochondrial shape, which can affect the function as well:[88] different structures of the mitochondrial network may afford the population a variety of physical, chemical, and signalling advantages or disadvantages.[89] Mitochondria in cells are always distributed along microtubules and the distribution of these organelles is also correlated with the endoplasmic reticulum.[90] Recent evidence suggests that vimentin, one of the components of the cytoskeleton, is also critical to the association with the cytoskeleton.[91]
Mitochondria-associated ER membrane (MAM)
The mitochondria-associated ER membrane (MAM) is another structural element that is increasingly recognized for its critical role in cellular physiology and
Purified MAM from subcellular fractionation is enriched in enzymes involved in phospholipid exchange, in addition to channels associated with Ca2+ signaling.[92][93] These hints of a prominent role for the MAM in the regulation of cellular lipid stores and signal transduction have been borne out, with significant implications for mitochondrial-associated cellular phenomena, as discussed below. Not only has the MAM provided insight into the mechanistic basis underlying such physiological processes as intrinsic apoptosis and the propagation of calcium signaling, but it also favors a more refined view of the mitochondria. Though often seen as static, isolated 'powerhouses' hijacked for cellular metabolism through an ancient endosymbiotic event, the evolution of the MAM underscores the extent to which mitochondria have been integrated into overall cellular physiology, with intimate physical and functional coupling to the endomembrane system.
Phospholipid transfer
The MAM is enriched in enzymes involved in lipid biosynthesis, such as phosphatidylserine synthase on the ER face and phosphatidylserine decarboxylase on the mitochondrial face.[94][95] Because mitochondria are dynamic organelles constantly undergoing fission and fusion events, they require a constant and well-regulated supply of phospholipids for membrane integrity.[96][97] But mitochondria are not only a destination for the phospholipids they finish synthesis of; rather, this organelle also plays a role in inter-organelle trafficking of the intermediates and products of phospholipid biosynthetic pathways, ceramide and cholesterol metabolism, and glycosphingolipid anabolism.[95][97]
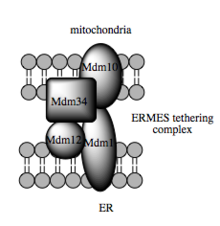
Such trafficking capacity depends on the MAM, which has been shown to facilitate transfer of lipid intermediates between organelles.[94] In contrast to the standard vesicular mechanism of lipid transfer, evidence indicates that the physical proximity of the ER and mitochondrial membranes at the MAM allows for lipid flipping between opposed bilayers.[97] Despite this unusual and seemingly energetically unfavorable mechanism, such transport does not require ATP.[97] Instead, in yeast, it has been shown to be dependent on a multiprotein tethering structure termed the ER-mitochondria encounter structure, or ERMES, although it remains unclear whether this structure directly mediates lipid transfer or is required to keep the membranes in sufficiently close proximity to lower the energy barrier for lipid flipping.[97][98]
The MAM may also be part of the secretory pathway, in addition to its role in intracellular lipid trafficking. In particular, the MAM appears to be an intermediate destination between the rough ER and the Golgi in the pathway that leads to
The MAM thus serves as a critical metabolic and trafficking hub in lipid metabolism.Calcium signaling
A critical role for the ER in calcium signaling was acknowledged before such a role for the mitochondria was widely accepted, in part because the low affinity of Ca2+ channels localized to the outer mitochondrial membrane seemed to contradict this organelle's purported responsiveness to changes in intracellular Ca2+ flux.
The fate of these puffs—in particular, whether they remain restricted to isolated locales or integrated into Ca2+ waves for propagation throughout the cell—is determined in large part by MAM dynamics. Although reuptake of Ca2+ by the ER (concomitant with its release) modulates the intensity of the puffs, thus insulating mitochondria to a certain degree from high Ca2+ exposure, the MAM often serves as a firewall that essentially buffers Ca2+ puffs by acting as a sink into which free ions released into the cytosol can be funneled.[92][100][101] This Ca2+ tunneling occurs through the low-affinity Ca2+ receptor VDAC1, which recently has been shown to be physically tethered to the IP3R clusters on the ER membrane and enriched at the MAM.[92][30][102] The ability of mitochondria to serve as a Ca2+ sink is a result of the electrochemical gradient generated during oxidative phosphorylation, which makes tunneling of the cation an exergonic process.[102] Normal, mild calcium influx from cytosol into the mitochondrial matrix causes transient depolarization that is corrected by pumping out protons.
But transmission of Ca2+ is not unidirectional; rather, it is a two-way street.
Regulating ER release of Ca2+ at the MAM is especially critical because only a certain window of Ca2+ uptake sustains the mitochondria, and consequently the cell, at homeostasis. Sufficient intraorganelle Ca2+ signaling is required to stimulate metabolism by activating dehydrogenase enzymes critical to flux through the citric acid cycle.[103][104] However, once Ca2+ signaling in the mitochondria passes a certain threshold, it stimulates the intrinsic pathway of apoptosis in part by collapsing the mitochondrial membrane potential required for metabolism.[92] Studies examining the role of pro- and anti-apoptotic factors support this model; for example, the anti-apoptotic factor Bcl-2 has been shown to interact with IP3Rs to reduce Ca2+ filling of the ER, leading to reduced efflux at the MAM and preventing collapse of the mitochondrial membrane potential post-apoptotic stimuli.[92] Given the need for such fine regulation of Ca2+ signaling, it is perhaps unsurprising that dysregulated mitochondrial Ca2+ has been implicated in several neurodegenerative diseases, while the catalogue of tumor suppressors includes a few that are enriched at the MAM.[102]
Molecular basis for tethering
Recent advances in the identification of the tethers between the mitochondrial and ER membranes suggest that the scaffolding function of the molecular elements involved is secondary to other, non-structural functions. In yeast, ERMES, a multiprotein complex of interacting ER- and mitochondrial-resident membrane proteins, is required for lipid transfer at the MAM and exemplifies this principle. One of its components, for example, is also a constituent of the protein complex required for insertion of transmembrane beta-barrel proteins into the lipid bilayer.[97] However, a homologue of the ERMES complex has not yet been identified in mammalian cells. Other proteins implicated in scaffolding likewise have functions independent of structural tethering at the MAM; for example, ER-resident and mitochondrial-resident mitofusins form heterocomplexes that regulate the number of inter-organelle contact sites, although mitofusins were first identified for their role in fission and fusion events between individual mitochondria.[92] Glucose-related protein 75 (grp75) is another dual-function protein. In addition to the matrix pool of grp75, a portion serves as a chaperone that physically links the mitochondrial and ER Ca2+ channels VDAC and IP3R for efficient Ca2+ transmission at the MAM.[92][30] Another potential tether is Sigma-1R, a non-opioid receptor whose stabilization of ER-resident IP3R may preserve communication at the MAM during the metabolic stress response.[105][106]
Perspective
The MAM is a critical signaling, metabolic, and trafficking hub in the cell that allows for the integration of ER and mitochondrial physiology. Coupling between these organelles is not simply structural but functional as well and critical for overall cellular physiology and homeostasis. The MAM thus offers a perspective on mitochondria that diverges from the traditional view of this organelle as a static, isolated unit appropriated for its metabolic capacity by the cell.[107] Instead, this mitochondrial-ER interface emphasizes the integration of the mitochondria, the product of an endosymbiotic event, into diverse cellular processes. Recently it has also been shown, that mitochondria and MAM-s in neurons are anchored to specialised intercellular communication sites (so called somatic-junctions). Microglial processes monitor and protect neuronal functions at these sites, and MAM-s are supposed to have an important role in this type of cellular quality-control.[80]
Origin and evolution
There are two hypotheses about the origin of mitochondria:
A
Proteobacteria
|
|
The ribosomes coded for by the mitochondrial DNA are similar to those from bacteria in size and structure.[122] They closely resemble the bacterial 70S ribosome and not the 80S cytoplasmic ribosomes, which are coded for by nuclear DNA.
The

A few groups of unicellular eukaryotes have only vestigial mitochondria or derived structures: The
Monocercomonoides and other oxymonads appear to have lost their mitochondria completely and at least some of the mitochondrial functions seem to be carried out by cytoplasmic proteins now.[5][130][10]
Mitochondrial genetics
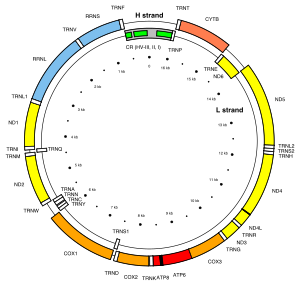
Mitochondria contain their own genome. The
As in prokaryotes, there is a very high proportion of coding DNA and an absence of repeats. Mitochondrial genes are
In animals, the mitochondrial genome is typically a single circular chromosome that is approximately 16 kb long and has 37 genes. The genes, while highly conserved, may vary in location. Curiously, this pattern is not found in the human body louse (
Human population genetic studies
The near-absence of
However, mitochondrial DNA reflects only the history of the females in a population. This can be partially overcome by the use of paternal genetic sequences, such as the
Recent measurements of the molecular clock for mitochondrial DNA[147] reported a value of 1 mutation every 7884 years dating back to the most recent common ancestor of humans and apes, which is consistent with estimates of mutation rates of autosomal DNA (10−8 per base per generation).[148]
Alternative genetic code
| Organism | Codon | Standard | Mitochondria |
|---|---|---|---|
| Mammals | AGA, AGG | Arginine | Stop codon |
| Invertebrates | AGA, AGG | Arginine | Serine |
| Fungi | CUA | Leucine | Threonine |
| All of the above | AUA | Isoleucine | Methionine |
| UGA | Stop codon | Tryptophan |
While slight variations on the standard genetic code had been predicted earlier,[149] none was discovered until 1979, when researchers studying human mitochondrial genes determined that they used an alternative code.[150] Nonetheless, the mitochondria of many other eukaryotes, including most plants, use the standard code.[151] Many slight variants have been discovered since,[151] including various alternative mitochondrial codes.[152] Further, the AUA, AUC, and AUU codons are all allowable start codons.
Some of these differences should be regarded as pseudo-changes in the genetic code due to the phenomenon of RNA editing, which is common in mitochondria. In higher plants, it was thought that CGG encoded for tryptophan and not arginine; however, the codon in the processed RNA was discovered to be the UGG codon, consistent with the standard genetic code for tryptophan.[153] Of note, the arthropod mitochondrial genetic code has undergone parallel evolution within a phylum, with some organisms uniquely translating AGG to lysine.[154]
Replication and inheritance
Mitochondria divide by
The hypothesis of mitochondrial binary fission has relied on the visualization by fluorescence microscopy and conventional
An individual's mitochondrial genes are inherited only from the mother, with rare exceptions.
Uniparental inheritance leads to little opportunity for genetic recombination between different lineages of mitochondria, although a single mitochondrion can contain 2–10 copies of its DNA.[132] What recombination does take place maintains genetic integrity rather than maintaining diversity. However, there are studies showing evidence of recombination in mitochondrial DNA. It is clear that the enzymes necessary for recombination are present in mammalian cells.[166] Further, evidence suggests that animal mitochondria can undergo recombination.[167] The data are more controversial in humans, although indirect evidence of recombination exists.[168][169]
Entities undergoing uniparental inheritance and with little to no recombination may be expected to be subject to Muller's ratchet, the accumulation of deleterious mutations until functionality is lost. Animal populations of mitochondria avoid this buildup through a developmental process known as the mtDNA bottleneck. The bottleneck exploits stochastic processes in the cell to increase the cell-to-cell variability in mutant load as an organism develops: a single egg cell with some proportion of mutant mtDNA thus produces an embryo where different cells have different mutant loads. Cell-level selection may then act to remove those cells with more mutant mtDNA, leading to a stabilization or reduction in mutant load between generations. The mechanism underlying the bottleneck is debated,[170][171][172] with a recent mathematical and experimental metastudy providing evidence for a combination of random partitioning of mtDNAs at cell divisions and random turnover of mtDNA molecules within the cell.[173]
DNA repair
Mitochondria can repair oxidative DNA damage by mechanisms analogous to those occurring in the cell nucleus. The proteins employed in mtDNA repair are encoded by nuclear genes, and are translocated to the mitochondria. The DNA repair pathways in mammalian mitochondria include base excision repair, double-strand break repair, direct reversal and mismatch repair.[174][175] Alternatively, DNA damage may be bypassed, rather than repaired, by translesion synthesis.
Of the several DNA repair process in mitochondria, the base excision repair pathway has been most comprehensively studied.
Double-strand breaks can be repaired by homologous recombinational repair in both mammalian mtDNA[177] and plant mtDNA.[178] Double-strand breaks in mtDNA can also be repaired by microhomology-mediated end joining.[179] Although there is evidence for the repair processes of direct reversal and mismatch repair in mtDNA, these processes are not well characterized.[175]
Lack of mitochondrial DNA
Some organisms have lost mitochondrial DNA altogether. In these cases, genes encoded by the mitochondrial DNA have been lost or transferred to the nucleus.[131] Cryptosporidium have mitochondria that lack any DNA, presumably because all their genes have been lost or transferred.[180] In Cryptosporidium, the mitochondria have an altered ATP generation system that renders the parasite resistant to many classical mitochondrial inhibitors such as cyanide, azide, and atovaquone.[180] Mitochondria that lack their own DNA have been found in a marine parasitic dinoflagellate from the genus Amoebophyra. This microorganism, A. cerati, has functional mitochondria that lack a genome.[181] In related species, the mitochondrial genome still has three genes, but in A. cerati only a single mitochondrial gene — the cytochrome c oxidase I gene (cox1) — is found, and it has migrated to the genome of the nucleus.[182]
Dysfunction and disease
Mitochondrial diseases
Damage and subsequent dysfunction in mitochondria is an important factor in a range of human diseases due to their influence in cell metabolism. Mitochondrial disorders often present as neurological disorders, including
It has also been reported that drug tolerant cancer cells have an increased number and size of mitochondria which suggested an increase in mitochondrial biogenesis.[185] A 2022 study in Nature Nanotechnology has reported that cancer cells can hijack the mitochondria from immune cells via physical tunneling nanotubes.[186]
In other diseases, defects in nuclear genes lead to dysfunction of mitochondrial proteins. This is the case in
Mitochondria-mediated oxidative stress plays a role in cardiomyopathy in
Mitochondria also modulate processes such as testicular somatic cell development, spermatogonial stem cell differentiation, luminal acidification, testosterone production in testes, and more. Thus, dysfunction of mitochondria in spermatozoa can be a cause for infertility.[195]
In efforts to combat mitochondrial disease, mitochondrial replacement therapy (MRT) has been developed. This form of in vitro fertilization uses donor mitochondria, which avoids the transmission of diseases caused by mutations of mitochondrial DNA.[196] However, this therapy is still being researched and can introduce genetic modification, as well as safety concerns. These diseases are rare but can be extremely debilitating and progressive diseases, thus posing complex ethical questions for public policy.[197]
Relationships to aging
There may be some leakage of the electrons transferred in the respiratory chain to form reactive oxygen species. This was thought to result in significant oxidative stress in the mitochondria with high mutation rates of mitochondrial DNA.[198] Hypothesized links between aging and oxidative stress are not new and were proposed in 1956,[199] which was later refined into the mitochondrial free radical theory of aging.[200] A vicious cycle was thought to occur, as oxidative stress leads to mitochondrial DNA mutations, which can lead to enzymatic abnormalities and further oxidative stress.
A number of changes can occur to mitochondria during the aging process.
Since mitochondria cover a pivotal role in the ovarian function, by providing ATP necessary for the development from germinal vesicle to mature oocyte, a decreased mitochondria function can lead to inflammation, resulting in premature ovarian failure and accelerated ovarian aging. The resulting dysfunction is then reflected in quantitative (such as mtDNA copy number and mtDNA deletions), qualitative (such as mutations and strand breaks) and oxidative damage (such as dysfunctional mitochondria due to ROS), which are not only relevant in ovarian aging, but perturb oocyte-cumulus crosstalk in the ovary, are linked to genetic disorders (such as Fragile X) and can interfere with embryo selection.[207]
History
The first observations of intracellular structures that probably represented mitochondria were published in 1857, by the physiologist
In 1939, experiments using minced muscle cells demonstrated that cellular respiration using one
The first high-resolution electron
The popular term "powerhouse of the cell" was coined by Philip Siekevitz in 1957.[4][217]
In 1967, it was discovered that mitochondria contained
See also
References
- ^ "mitochondrion". Lexico UK English Dictionary. Oxford University Press. Archived from the original on January 2, 2020.
- ISBN 978-0132508827. Archivedfrom the original on November 2, 2014. Retrieved January 6, 2009.
- ^ "Mighty Mitochondria and Neurodegenerative Diseases". Science in the News. February 1, 2012. Archived from the original on April 6, 2022. Retrieved April 24, 2022.
- ^ .
- ^ PMID 27185558.
- ^ Le Page M. "Animal that doesn't need oxygen to survive discovered New Scientist". New Scientist. Archived from the original on February 26, 2020. Retrieved February 25, 2020.
- ^ PMID 32094163.
- ^ S2CID 862398.
- PMID 28474007.
- ^ PMID 38060519.
- PMID 26777473.
- ^ S2CID 16252290.
- PMID 24606795.
Mitochondrial biogenesis is therefore defined as the process via which cells increase their individual mitochondrial mass [3]. ... Mitochondrial biogenesis occurs by growth and division of pre-existing organelles and is temporally coordinated with cell cycle events [1].
- PMID 24606801.
Mitochondrial biogenesis (MB) is the essential mechanism by which cells control the number of mitochondria
- .
- PMID 11444914.
- PMID 26443844.
- ^ PMID 28630658.
- PMID 21423015.
- ^ ISBN 978-0815341055.
- ^ ISBN 978-0471214953.
- PMID 12594925.
- ^ S2CID 236916203.
- ^ "Mitochondrion – much more than an energy converter". British Society for Cell Biology. Archived from the original on April 4, 2019. Retrieved August 19, 2013.
- S2CID 38314888.
- PMID 17524423.
- PMID 20450883.
- ^ (PDF) from the original on August 16, 2022. Retrieved July 10, 2022.
- ^ PMID 16710362.
- ^ PMID 19144519.
- PMID 24578708.
- PMID 12531542.
- S2CID 253211400.
- PMID 22936770.
- PMID 29383328.
- PMID 16730811.
- PMID 22142616.
- PMID 14641005.
- S2CID 8963850.
- PMID 9242927.
- ^ ISBN 0716720094.
- S2CID 26263513.
- ^ ISBN 047119350X
- ^ a b Atkins, P.; de Paula, J. (2006) "Physical Chemistry", 8th ed.; pp. 225-229, Freeman: New York, 2006.
- S2CID 2278219. Archived from the original(PDF) on March 3, 2019.
- S2CID 4149605.
- S2CID 4160146.
- ^ Nobel Foundation. "Chemistry 1997". Archived from the original on July 8, 2007. Retrieved December 16, 2007.
- ^ S2CID 164450.
- PMID 27553474.
- PMID 26963735.
- ^ PMID 30352195.
- S2CID 199404906.
- PMID 26217001.
- ^ ISBN 978-0397518203.
- ^ PMID 16759697.
- PMID 4134194.
- ^
Brighton CT, Hunt RM (July 1978). "The role of mitochondria in growth plate calcification as demonstrated in a rachitic model". The Journal of Bone and Joint Surgery. American Volume. 60 (5): 630–639. PMID 681381.
- ^ PMID 25966694.
- PMID 17851078.
- ^ S2CID 5193821.
- PMID 25844899.
- PMID 22395486.
- PMID 23746507.
- S2CID 29827252.
- ^ S2CID 7748115.
- PMID 31727843.
- S2CID 10279742.
- S2CID 22259833.
- S2CID 29715383.
- ^ S2CID 25906279.
- PMID 23442817.
- S2CID 16654082.
- PMID 17074387.
- S2CID 13602877.
- PMID 18846505.
- S2CID 30841790.
- S2CID 24794040.
- PMID 31247505.
- ^ from the original on May 2, 2023. Retrieved February 10, 2022.
- S2CID 252416407.
- S2CID 27329521.
- PMID 18348319.
- PMID 18484766.
- PMID 20491666.
- PMID 21179497.
- PMID 22412363.
- S2CID 28165195.
- PMID 25847815.
- PMID 1363623.
- PMID 17983357.
- ^ PMID 19341702.
- ^ PMID 20717141.
- ^ PMID 8694499.
- ^ PMID 19703651.
- PMID 18200046.
- ^ PMID 21220505.
- PMID 19556461.
- PMID 7961664.
- ^ PMID 17889347.
- PMID 11243933.
- ^ PMID 21146562.
- PMID 27913206.
- PMID 12543096.
- PMID 22923735.
- S2CID 18885068.
- . Epub 2018 Mar 24.
- S2CID 235786110.
- from the original on August 15, 2022. Retrieved August 18, 2022.
- ^ ISBN 978-0300033403.
- ^ Martin WF, Müller M (2007). Origin of mitochondria and hydrogenosomes. Heidelberg, DE: Springer Verlag.
- PMID 12694174.
- PMID 10376009.
- PMID 10066161.
- PMID 22355532.
- S2CID 13740626.
- S2CID 220507452.
- PMID 34083540.
- S2CID 248836055.
- (PDF) from the original on November 18, 2023. Retrieved December 30, 2023.
- PMID 35798888.
- S2CID 43415449.
- PMID 11541392.
- S2CID 10144020.
- PMID 9371794.
- PMID 1854912.
- PMID 20370908.
- ^ Coghaln A (April 7, 2010). "The mud creature that lives without oxygen". Zoologger. New Scientist. Archived from the original on February 26, 2020. Retrieved February 27, 2020.
- PMID 20528687.
- from the original on April 12, 2024. Retrieved April 14, 2024.
- ^ S2CID 8551160.
- ^ PMID 1550563.
- PMID 29880722.
- ^ S2CID 4355527.
- PMID 8455612.
- S2CID 4314378.
- PMID 1579468.
- PMID 9461442.
- PMID 15568984.
- PMID 25339973.
- PMID 19336451.
- PMID 10943382.
- S2CID 4285418.
- PMID 16678300.
- ^ S2CID 176541.
- S2CID 13581775.
- PMID 19500773.
- PMID 10978293.
- doi:10.1016/0019-1035(73)90110-3. Archived (PDF) from the original on October 29, 2011. Retrieved October 21, 2014..
p. 344: It is a little surprising that organisms with somewhat different codes do not coexist.
Further discussion Archived September 3, 2011, at the Wayback Machine - S2CID 4335828.
- ^ a b Elzanowski A, Ostell J (January 7, 2019). "The Genetic Codes". www.ncbi.nlm.nih.gov. Archived from the original on May 13, 2011. Retrieved February 10, 2023.
- S2CID 19264964.
- PMID 2480644.
- PMID 16620150.
- ISBN 978-1439807149.
- PMID 20940129.
- S2CID 7440669.
- PMID 30643304.
- ^ Kimball, J.W. (2006) "Sexual Reproduction in Humans: Copulation and Fertilization" Archived October 2, 2015, at the Wayback Machine, Kimball's Biology Pages (based on Biology, 6th ed., 1996)
- .
- JSTOR 2446172.
- PMID 11280005.
- PMID 9475744.
- ^ Male and Female Mitochondrial DNA Lineages in the Blue Mussel (Mytilus edulis) Species Group Archived May 18, 2013, at the Wayback Machine by Donald T. Stewart, Carlos Saavedra, Rebecca R. Stanwood, Amy 0. Ball, and Eleftherios Zouros
- S2CID 90422.
- PMID 8910339.
- S2CID 4318161.
- PMID 10189711.
- S2CID 27685285.
- S2CID 205344980.
- S2CID 10686347.
- S2CID 225349.
- PMID 26035426.
- PMID 23050036.
- ^ PMID 27915046.
- PMID 26912170.
- S2CID 4779650.
- PMID 17396019.
- PMID 26609070.
- ^ PMID 15664529.
- PMID 31032404.
- ^ "Veritable powerhouse – even without DNA: Parasitic algae from the dinoflagellate lineage have organized their genetic material in an unprecedented way". ScienceDaily. Archived from the original on June 24, 2019. Retrieved May 8, 2019.
- ^ PMID 15358637.
- PMID 15861210.
- PMID 31431543.
- S2CID 244349825.
- PMID 12933917.
- S2CID 24318009.
- PMID 17127363.
- S2CID 33077667.
- PMID 24931469.
- S2CID 27039617.
- PMID 17239370.
- PMID 20639213.
- PMID 35269419.
- PMID 33498182.
- ^ Committee on the Ethical and Social Policy Considerations of Novel Techniques for Prevention of Maternal Transmission of Mitochondrial DNA Diseases; Board on Health Sciences Policy; Institute of Medicine; National Academies of Sciences, Engineering, and Medicine (March 17, 2016), Claiborne A, English R, Kahn J (eds.), "Introduction", Mitochondrial Replacement Techniques: Ethical, Social, and Policy Considerations, National Academies Press (US), retrieved December 5, 2023
- PMID 3413108.
- PMID 13332224.
- S2CID 396830.
- ^ "Mitochondria and Aging". circuitblue.co. Archived from the original on September 29, 2017. Retrieved October 23, 2006.
- PMID 8155742.
- PMID 15588517.
- S2CID 13956928.
- S2CID 201168692.
- PMID 33398403.
- S2CID 221472721.
- ^ Kölliker A (1857). "Einige Bemerkungen über die Endigungen der Hautnerven und den Bau der Muskeln" [Some remarks about the terminations of the cutaneous nerves and the structure of muscles]. Zeitschrift für wissenschaftliche Zoologie (in German). 8: 311–325. Archived from the original on June 22, 2023. Retrieved June 22, 2023. On p. 316, Kölliker described mitochondria which he observed in fresh frog muscles: " ... sehr blasse rundliche Körnchen, welche in langen linienförmigen Zügen [...] wenn man einmal auf dieselben aufmerksam geworden ist." ( ... [they are] very faint round granules, which are embedded in the [muscle's] contractile substance in long linear trains. These granules are located in the whole thickness of the muscle fiber, on the surface as in the interior, and [they] are so numerous that they appear as a not unimportant element of the muscle fibers, once one has become alert to them.) Kölliker said (p. 321) that he had found mitochondria in the muscles of other animals. In Figure 3 of Table XIV Archived June 22, 2023, at the Wayback Machine, Kölliker depicted mitochondria in frog muscles.
- ^ PMID 7033239.
- ^ Altmann R (1890). Die Elementarorganismen und ihre Beziehungen zu den Zellen [Elementary Organisms and Their Relations to Cells] (in German). Leipzig, Germany: Veit & Co. p. 125. Archived from the original on June 23, 2023. Retrieved June 23, 2023. From p. 125: "Da auch sonst mancherlei Umstände dafür sprechen, dass Mikroorganismen und Granula einander gleichwerthig sind und Elementarorganismen vorstellen, welche sich überall finden, wo lebendige Kräfte ausgelöst werden, so wollen wir sie mit dem gemeinschaftlichen Namen der Bioblasten bezeichnen." (Since otherwise some circumstances indicate that microorganisms and granula are equivalent to each other and suggest elementary organisms, which are to be found wherever living forces are unleashed, we will designate them with the collective name of "bioblasts".)
- ^ "mitochondria". Online Etymology Dictionary. Archived from the original on March 4, 2016. Retrieved May 23, 2013.
- ^ Benda C (1898). "Ueber die Spermatogenese der Vertebraten und höherer Evertebraten. II. Theil: Die Histiogenese der Spermien" [On spermatogenesis in vertebrates and higher invertebrates. Part II: The histogenesis of sperm.]. Archiv für Physiologie (in German). 1898: 393–398. Archived from the original on February 24, 2019. Retrieved January 14, 2018. From p. 397: After Brenda states that " ... ich bereits in vielen Zellarten aller möglichen Thierclassen gefunden habe, ... " ( ... I have already found [them (mitochondria)] in many types of cells of all possible classes of animals, ... ), he suggests: "Ich möchte vorläufig vorschlagen, ihnen als Mitochondria eine besondere Stellung vorzubehalten, die ich in weiteren Arbeiten begründen werde." (I would like to suggest provisionally reserving for them, as "mitochondria", a special status which I will justify in further work.)
- Benda C (1899). "Weitere Mitteilungen über die Mitochondria" [Further reports on mitochondria]. Archiv für Physiologie: Verhandlungen der Berliner Physiologischen Gesellschaft (in German). 1899: 376–383. Archived from the original on June 23, 2023. Retrieved June 23, 2023.
- from the original on June 23, 2023. Retrieved June 23, 2023.
- S2CID 257822147., cited in Meves' 1908 paper and in Schmidt EW (1913). "Pflanzliche Mitochondrien". Progressus Rei Botanicae. 4: 164–183. Retrieved September 21, 2012., with confirmation of Nymphaea alba
- from the original on June 23, 2023. Retrieved June 23, 2023. From p. 47: " ... the mitochondria are the structural expression thereof [i.e., of the chemical reducing processes in the cytoplasm], ... "
- ^ Warburg O (1913). "Über sauerstoffatmende Körnchen aus Leberzellen und über Sauerstoffatmung in Berkefeld-Filtraten wässriger Leberextrake" [On respiring granules from liver cells and on respiration in Berkefeld filtrates of aqueous liver extracts]. Pflügers Archiv für die gesamte Physiologie des Menschen und der Tiere (Pfluger's Archive for All Physiology of Humans and Animals) (in German). 154: 599–617. Archived from the original on June 23, 2023. Retrieved June 23, 2023.
- PMID 25841699.
- PMID 26323761.
General
 This article incorporates NCBI. Archived from the originalon December 8, 2009.
This article incorporates NCBI. Archived from the originalon December 8, 2009.
External links
- Lane N (2016). The Vital Question: Energy, Evolution, and the Origins of Complex Life. WW Norton & Company. ISBN 978-0393352979.
- Powering the Cell Mitochondria Archived August 17, 2022, at the Wayback Machine – XVIVO Scientific Animation
- Mitodb.com Archived July 3, 2013, at the Wayback Machine – The mitochondrial disease database.
- Mitochondria Atlas Archived June 29, 2012, at the Wayback Machine at University of Mainz
- Mitochondria Research Portal at mitochondrial.net
- Mitochondria: Architecture dictates function at cytochemistry.net
- Mitochondria links at University of Alabama
- MIP Archived May 23, 2016, at the Portuguese Web Archive Mitochondrial Physiology Society
- 3D structures of proteins from inner mitochondrial membrane Archived September 13, 2011, at the Wayback Machine at University of Michigan
- 3D structures of proteins associated with outer mitochondrial membrane Archived September 13, 2011, at the Wayback Machine at University of Michigan
- Mitochondrial Protein Partnership at University of Wisconsin
- MitoMiner – A mitochondrial proteomics database[permanent dead link] at MRC Mitochondrial Biology Unit
- Mitochondrion – Cell Centered Database Archived October 7, 2011, at the Wayback Machine
- Mitochondrion Reconstructed by Electron Tomography Archived November 14, 2012, at the Wayback Machine at San Diego State University
- Video Clip of Rat-liver Mitochondrion from Cryo-electron Tomography






