PETase
| PETase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
PETases are an esterase class of enzymes that catalyze the breakdown (via hydrolysis) of polyethylene terephthalate (PET) plastic to monomeric mono-2-hydroxyethyl terephthalate (MHET). The idealized chemical reaction is:
- (ethylene terephthalate)n + H2O → (ethylene terephthalate)n-1 + MHET,
where n is the number of
Whereas the degradation of PET by natural (non-enzymatic) means will take hundreds of years, PETases can degrade it in a matter of days.[3]
History
The first PETase was discovered in 2016 from Ideonella sakaiensis strain 201-F6 bacteria found from sludge samples collected close to a Japanese PET bottle recycling site.[1][4] There were other types of hydrolases previously known to degrade PET,[2] including lipases, esterases, and cutinases.[5] For comparison, enzymes that degrade polyester have been known to exist at least as far back as 1975 (in the case of α-chymotrypsin)[6] and 1977 (lipase).[7]
PET plastic came into widespread use in the 1970s and it has been suggested that PETases in bacteria evolved only recently.[2] PETase may have had past enzymatic activity associated with degradation of a waxy coating on plants.[8]
Structure
As of April 2019, there were 17 known three-dimensional crystal structures of PETases: 6QGC, 6ILX, 6ILW, 5YFE, 6EQD, 6EQE, 6EQF, 6EQG, 6EQH, 6ANE, 5XJH, 5YNS, 5XFY, 5XFZ, 5XG0, 5XH2 and 5XH3.
PETase exhibits shared qualities with both lipases and cutinases in that it possesses an α/β-hydrolase fold; although, the active-site cleft observed in PETase is more open than in cutinases.[2] The Ideonella sakaiensis PETase is similar to dienelactone hydrolase, according to Pfam. According to ESTHER, it falls into the Polyesterase-lipase-cutinase family.
There are approximately 69 PETase-like enzymes comprising a variety of diverse organisms, and there are two classifications of these enzymes including type I and type II. It is suggested that 57 enzymes fall into the type I category whereas the rest fall into the type II group, including the PETase enzyme found in the Ideonella sakaiensis. Within all 69 PETase-like enzymes, there exists the same three residues within the active site, suggesting that the catalytic mechanism is the same in all forms of PETase-like enzymes.[9]
-
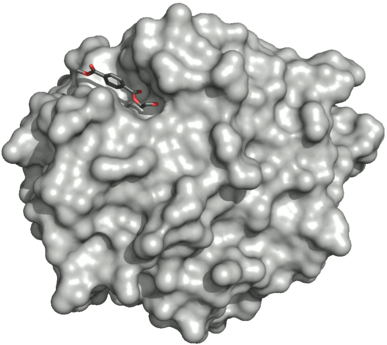
Surface of the PETase double mutant (R103G and S131A) with HEMT (1-(2-hydroxyethyl) 4-methyl terephthalate) bound to its active site. HEMT is an analogue of MHET, and has an additional methanol esterified to it. PDBID: 5XH3.
-
Ribbon diagram of PETase with three residues Ser160, Asp206, and His237. The catalytic triad is represented by cyan-colored sticks. The active site is shown in orange to represent stimulation by a 2-HE(MHET)4 molecule.[9]
Mutations
The discovery of PETase from I. sakaiensis provides a potential solution to the world’s amassing plastic; however, naturally occurring enzymes are limited in their degradation abilities due to instability, low activity, and expression levels, which ultimately drive the need for improvement if they are to be used for large-scale industrial applications.
Biological pathway

In I. sakaiensis, the resultant MHET is further broken down by the action of MHETase enzyme to terephthalic acid and ethylene glycol.[1] Laboratory experiments showed that chimeric proteins that artificially link a MHETase and a PETase outperform similar mixtures of free enzymes.[15]
See also
- Plastivore
- Galleria mellonella, a caterpillar that can digest polyethylene.
- Aspergillus tubingensis, a fungus that can digest polyurethane.
- Pestalotiopsis microspora, an endophytic fungus species able to break down polyurethane.
- Cutinase, an esterase enzyme of similar geometric shape
References
- ^ S2CID 31146235.
- "Discovery of a Bacterium that Degrades and Assimilates Poly(ethylene terephthalate) could Serve as a Degradation and/or Fermentation Platform for Biological Recycling of PET Waste Products" (PDF). Kyoto Institute of Technology (Press release). 2016-03-30.
- ^ PMID 29666242.
- ^ Dockrill, Peter. "Scientists Have Accidentally Created a Mutant Enzyme That Eats Plastic Waste". ScienceAlert. Retrieved 2018-11-27.
- PMID 27045688.
- PMID 29235460.
- .
- S2CID 4145159.
- ^ "Lab 'Accident' Becomes Mutant Enzyme That Devours Plastic". Live Science. Retrieved 2018-11-27.
- ^ PMID 29374183.
- ^ PMID 35056486.
- ^ PMID 34917598.
- PMID 29666242.
- S2CID 233620229.
- ^ Allison Chan (2016). "The Future of Bacteria Cleaning Our Plastic Waste" (PDF).
- PMID 32989159.

![Ribbon diagram of PETase with three residues Ser160, Asp206, and His237. The catalytic triad is represented by cyan-colored sticks. The active site is shown in orange to represent stimulation by a 2-HE(MHET)4 molecule.[9]](http://upload.wikimedia.org/wikipedia/commons/thumb/d/db/PETase_active_site.png/739px-PETase_active_site.png)