Phospholipase D
| Phospholipase D | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | PLDc | ||||||||
SCOP2 | 1byr / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 118 | ||||||||
| OPM protein | 3rlh | ||||||||
| CDD | cd00138 | ||||||||
| Membranome | 306 | ||||||||
| |||||||||
| phospholipase D | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Phospholipase D (EC 3.1.4.4, lipophosphodiesterase II, lecithinase D, choline phosphatase, PLD; systematic name phosphatidylcholine phosphatidohydrolase) is an enzyme of the phospholipase superfamily that catalyses the following reaction
- a phosphatidylcholine + H2O = choline + a phosphatidate
Phospholipases occur widely, and can be found in a wide range of organisms, including bacteria, yeast, plants, animals, and viruses.
Discovery
PLD-type
Function
Strictly speaking, phospholipase D is a
Phosphatidic acid
Many of phospholipase D's
It is possible that, though PA and DAG are interconvertible, separate pools of signalling and non-signalling
A lysophospholipase D called autotaxin was recently identified as having an important role in cell-proliferation through its product, lysophosphatidic acid (LPA).
Structure
Plant and animal PLDs have a consistent
Human proteins containing this motif include:
PLD
Mechanism of catalysis
PLD-
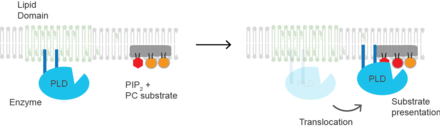
Mechanism of activation
Substrate presentation For mammalian PLD2, the molecular basis of activation is substrate presentation. The enzyme resides inactive in lipid micro-domains rich in sphingomyelin and depleted of PC substrate.[23] Increased PIP2 or a decrease in cholesterol causes the enzyme to translocate to PIP2 micro domains near its substrate PC. Hence PLD can is primarily activated by localization within the plasma membrane rather than a protein conformational change. Disruption of lipid domains by anesthetics.[24] or mechanical force.[23] The protein may also undergo a conformational change upon PIP2 binding, but this has not been shown experimentally and would constitute a mechanism of activation distinct from substrate presentation.
Isoforms
Two major
PLD1
PLD2In contrast, PLD2 is a 106 |
| ||||||||||||||||||||
Regulation
The activity of phospholipase D is extensively
Specific
C2 domain
PX domain
The
PH domain
The highly conserved
Interactions with small GTPases
In
Physiological and pathophysiological roles
Alcohol Intoxication
Phospholipase D metabolizes ethanol into phosphatidylethanol (PEtOH) in a process termed transphosphatidylation. Using fly genetics the PEtOH was shown to mediates alcohol's hyperactive response in fruit flies.[27] And ethanol transphosphatidylation was shown to be up-regulated in alcoholics and the family members of alcoholic.s[28] This ethanol transphosphatidylation mechanism recently emerged as an alternative theory for alcohol's effect on ion channels. Many ion channels are regulated by anionic lipids.[29] and the competition of PEtOH with endogenous signaling lipids is thought to mediate the effect of ethanol on ion channels in some instances and not direct binding of the free ethanol to the channel.[27]
Mechanosensation
PLD2 is a mechanosensor and directly sensitive to mechanical disruption of clustered GM1 lipids.[3] Mechanical disruption (fluid shear) then signals for the cell to differentiate. PLD2 also activates TREK-1 channels, a potassium channel in the analgesic pathway.[30]
In cancer
Phospholipase D is a regulator of several critical cellular processes, including
In neurodegenerative diseases
Phospholipase D may also play an important
Abnormal PLD activity has also been suspected in
Gallery
-
Phosphatidyl choline
-
Phosphatidate
-
Phospholipase cleavage sites
References
- ^ S2CID 26447185.
- ^ PMID 11987824.
- ^ PMID 27976674.
- ^ S2CID 14815405.
- ^ PMID 21447092.
- ^ PMID 14517341.
- doi:10.1101/758896.
- PMID 32467161.
- ^ S2CID 24389113.
- ^ PMID 15052340.
- ^ PMID 8144636.
- ^ PMID 10873862.
- S2CID 28348537.
- PMID 3117799.
- S2CID 10674790.
- PMID 9644971.
- S2CID 21937549.
- ^ Nowicki M (2006). Characterization of the Cardiolipin Synthase from Arabidopsis thaliana (Ph.D. thesis). RWTH-Aachen University. Archived from the original on 2011-10-05. Retrieved 2011-07-11.
- PMID 8732763.
- PMID 8755242.
- PMID 8051126.
- PMID 9242915.
- ^ PMID 27976674.
- PMID 32467161.
- ^ PMID 22157867.
- PMID 27976674.
- ^ PMID 30529033.
- PMID 3200856.
- PMID 25633344.
- PMID 25197053.
- PMID 24336208.
External links
- Phospholipase+D at the U.S. National Library of Medicine Medical Subject Headings (MeSH)



