Peripheral membrane protein
Peripheral membrane proteins, or extrinsic membrane proteins,
The reversible attachment of proteins to biological membranes has shown to regulate
Binding to the lipid bilayer

Peripheral membrane proteins may interact with other proteins or directly with the lipid bilayer. In the latter case, they are then known as amphitropic proteins.[4] Some proteins, such as
The
Some water-soluble proteins associate with lipid bilayers irreversibly and can form transmembrane alpha-helical or
Membrane binding mechanisms

The association of a protein with a
Typical amphitropic proteins must interact strongly with the lipid bilayer in order to perform their biological functions. These include the enzymatic processing of lipids and other hydrophobic substances, membrane anchoring, and the binding and transfer of small nonpolar compounds between different cellular membranes. These proteins may be anchored to the bilayer as a result of hydrophobic interactions between the bilayer and exposed nonpolar residues at the surface of a protein,
It has been shown that the membrane binding affinities of many peripheral proteins depend on the specific lipid composition of the membrane with which they are associated.[14]

Non-specific hydrophobic association
Amphitropic proteins associate with lipid bilayers via various
Covalently bound lipid anchors
Specific protein–lipid binding

Some
Protein–lipid electrostatic interactions
Any positively charged protein will be attracted to a negatively charged membrane by nonspecific
Electrostatic interactions are strongly dependent on the
Spatial position in membrane
Orientations and penetration depths of many amphitropic proteins and peptides in membranes are studied using
Two distinct membrane-association modes of proteins have been identified. Typical water-soluble proteins have no exposed nonpolar residues or any other hydrophobic anchors. Therefore, they remain completely in aqueous solution and do not penetrate into the lipid bilayer, which would be energetically costly. Such proteins interact with bilayers only electrostatically, for example,
Categories
Enzymes
Peripheral enzymes participate in
| Class | Function | Physiology | Structure |
|---|---|---|---|
Alpha/beta hydrolase fold |
Catalyzes the hydrolysis of chemical bonds.[37] | Includes fungal, gastric and pancreatic lipases, palmitoyl protein thioesterases, cutinase, and cholinesterases |
central beta sheet inserted in between two layers of alpha helices[38] |
| Phospholipase A2 (secretory and cytosolic) | Hydrolysis of sn-2 fatty acid bond of phospholipids.[39] | Lipid digestion, membrane disruption, and lipid signaling. | contains catalytic amino acid triad: aspartic acid, serine, and histidine[40] |
| Phospholipase C | Hydrolyzes PIP2, a inositol triphosphate and diacylglycerol.[41] |
Lipid signaling | core structure composed of a split triosephosphate isomerase (TIM) barrel which has an active site, catalytic residues, and a Ca2+ binding site [42] |
| Cholesterol oxidases | Oxidizes and isomerizes cholesterol to cholest-4-en-3-one.[43] |
Depletes cellular membranes of cholesterol, used in bacterial pathogenesis . |
two loops of residue which act as a lid on the active site[44] |
| Carotenoid oxygenase | Cleaves carotenoids.[45] |
Carotenoids function in both plants and animals as flavors , floral scents and defense compounds. |
composed of multiple enzymes attached together forming branch-like structures[46] |
| Lipoxygenases | In animals lipoxygenases are involved in the synthesis of leukotrienes . |
hundreds of amino acids that makes up a protein are organized into two domains: beta-sheet N terminal and helical C terminal[48] | |
| Alpha toxins | Cleave phospholipids in the cell membrane, similar to Phospholipase C.[49] | Bacterial pathogenesis, particularly by Clostridium perfringens. | soluble monomer with oligomeric pre-pore complexes[50] |
Sphingomyelinase C |
A phosphodiesterase, cleaves phosphodiester bonds.[51] | Processing of lipids such as sphingomyelin. | saposin domain and connector regions with a metallophosphate catalytic domain [52] |
| Glycosyltransferases: MurG and Transglycosidases | Catalyzes the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds.[53] |
Biosynthesis of disaccharides, oligosaccharides and polysaccharides (glycoconjugates), MurG is involved in bacterial peptidoglycan biosynthesis. | three glycine rich loops: one in the C terminal and two in the N terminal [54] |
| Ferrochelatase | Converts protoporphyrin IX into heme.[55] | Involved in egg shells . |
helices[56]
|
| Myotubularin-related protein family | Lipid PtdIns(3,5)P2.[57] |
Required for muscle cell differentiation. | contains a GRAM domain, SET interacting domain, and a PDZ binding domain[58] |
| Dihydroorotate dehydrogenases | Oxidation of dihydroorotate (DHO) to orotate.[59] |
Biosynthesis of eukaryotic cells. |
composed of two domains: alpha-helical domain that forms the opening tunnel to the active site [60]
|
Glycolate oxidase |
Catalyses the ketoacids.[61] |
In green plants, the enzyme participates in photorespiration. In animals, the enzyme participates in production of oxalate. | β8/α8 fold containing alpha helices, beta strands, and loops and turns[62] |
Membrane-targeting domains (“lipid clamps")
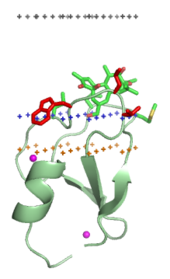
Membrane-targeting domains associate specifically with head groups of their lipid ligands embedded into the membrane. These lipid ligands are present in different concentrations in distinct types of biological membranes (for example,
Hence, each domain is targeted to a specific membrane.- C1 domains and phorbol esters.
- PtdIns(4,5)P2.
- Pleckstrin homology domains, PX domains, and Tubby domains bind different phosphoinositides
- FYVE domains are more specific for PtdIns3P.
- PtdIns(4,5)P2.
- ANTHdomain binds PtdIns(4,5)P2.
- Proteins from ERM (ezrin/radixin/moesin) family bind PtdIns(4,5)P2.
- Other phosphotyrosine-binding domain and certain PDZ domains. They bind PtdIns(4,5)P2.
- Discoidin domains of blood coagulation factors
- ANTHdomains
Structural domains
Structural domains mediate attachment of other proteins to membranes. Their binding to membranes can be mediated by calcium ions (Ca2+) that form bridges between the acidic protein residues and phosphate groups of lipids, as in annexins or GLA domains.
| Class | Function | Physiology | Structure |
|---|---|---|---|
| Annexins | Calcium-dependent intracellular membrane/ phospholipid binding.[63] | Functions include vesicle trafficking, membrane fusion and ion channel formation. |
|
| Synapsin I | Coats cytoskeletal elements.[64] |
Functions in the regulation of neurotransmitter release. | |
| Synuclein | Unknown cellular function.[65] | Thought to play a role in regulating the stability and/or turnover of the plasma membrane. Associated with both Parkinson's disease and Alzheimer's disease . |
|
| GLA-domains of the coagulation system | Gamma-carboxyglutamate (GLA) domains are responsible for the high-affinity binding of calcium ions.[66] |
Involved in function of clotting factors in the blood coagulation cascade. | |
| Spectrin and α-actinin-2 | Found in several cytoskeletal and microfilament proteins.[67] | Maintenance of plasma membrane integrity and cytoskeletal structure. |
Transporters of small hydrophobic molecules
These peripheral proteins function as carriers of non-polar compounds between different types of cell membranes or between membranes and cytosolic protein complexes. The transported substances are phosphatidylinositol, tocopherol, gangliosides, glycolipids, sterol derivatives, retinol, fatty acids, water, macromolecules, red blood cells, phospholipids, and nucleotides.
- Glycolipid transfer proteins
- retinol binding proteins and fatty acid-binding proteins
- Polyisoprenoid-binding protein, such as YceI protein domain
- Ganglioside GM2 activator proteins
- CRAL-TRIO domain (α-Tocopherol and phosphatidylinositol sec14p transfer proteins)
- Sterol carrier proteins
- Phosphatidylinositol transfer proteins and STAR domains
- Oxysterol-binding protein
Electron carriers
These proteins are involved in
Polypeptide hormones, toxins, and antimicrobial peptides
Many hormones,
Some water-soluble proteins and peptides can also form
| Class | Proteins | Physiology |
|---|---|---|
| Venom toxins |
|
Well known types of biotoxins include |
| Sea anemone toxins |
|
Inhibition of sodium and carnivorous animals and use toxins in predation and defense; anemone toxin is of similar toxicity as the most toxic organophosphate chemical warfare agents.[69]
|
Bacterial toxins |
|
second messenger pathways causing dramatic alterations to signal transduction pathways critical in maintaining a variety of cellular functions. Several bacterial toxins can act directly on the immune system, by acting as superantigens and causing massive T cell proliferation, which overextends the immune system. Botulinum toxin is a neurotoxin that prevents neuro-secretory vesicles from docking/fusing with the nerve synapse plasma membrane, inhibiting neurotransmitter release.[70]
|
Fungal toxins |
|
These peptides are characterized by the presence of an unusual amino acid, α-aminoisobutyric acid, and exhibit antibiotic and antifungal properties due to their membrane channel-forming activities.[71] |
Antimicrobial peptides |
|
The modes of action by which antimicrobial peptides kill bacteria is varied and includes disrupting membranes, interfering with bacteriostatic .
|
| Defensins |
|
Defensins are a type of antimicrobial peptide; and are an important component of virtually all innate host defenses against microbial invasion. Defensins penetrate microbial cell membranes by way of electrical attraction, and form a pore in the membrane allowing efflux, which ultimately leads to the lysis of microorganisms.[72] |
Neuronal peptides |
|
These proteins excite neurons, evoke behavioral responses, are potent vasodilatators, and are responsible for contraction in many types of smooth muscle.[73]
|
| Apoptosis regulators | Members of the Bcl-2 family govern neuronal cells .
|
See also
References
- ^ "extrinsic protein | biology | Britannica". www.britannica.com. Retrieved 2022-07-04.
- ISBN 3-527-31151-3.
- PMID 16814865.
- ^ PMID 10503244.
- PMID 16650855.
- S2CID 36940600.
- ISBN 978-0-12-643871-0.
- PMID 15016920.
- PMID 11438731.
- S2CID 36212541.
- PMID 11063882.
- S2CID 20892603.
- PMID 12719487.
- PMID 16689633.
- ^ PMID 8780505.
- ISBN 978-0-12-643871-0.
- ISBN 978-0-444-51139-3.
- ^ PMID 15869386.
- PMID 9336168.
- ISBN 0-444-81575-9.
- PMID 15869384.
- PMID 10551860.
- PMID 11258945.
- PMID 11230696.
- PMID 16055150.
- ^ PMID 10388560.
- PMID 11804593.
- PMID 15379706.
- PMID 16731967.
- ^ Lomize A, Lomize M, Pogozheva I. "Comparison with experimental data". Orientations of Proteins in Membranes. University of Michigan. Retrieved 2007-02-08.
- PMID 52374.
- PMID 15519307.
- .
- PMID 15315949.
- PMID 12829487.
- PMID 12670959.
- ^ "Pfam entry Abhydrolase 1". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ISSN 1742-464X.
- ^ "Pfam entry: Phospholipase A2". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- PMID 30521272, retrieved 2023-11-29
- ^ "Pfam entry: Phosphatidylinositol-specific phospholipase C, X domain". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ^ "Phospholipase C", Wikipedia, 2023-08-16, retrieved 2023-11-29
- ^ "Pfam entry: Cholesterol oxidase". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ISSN 0006-2960.
- ^ "Pfam entry: Retinal pigment epithelial membrane protein". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ^ "Carotenoid oxygenase", Wikipedia, 2023-11-29, retrieved 2023-11-29
- ^ "Pfam entry: Lipoxygenase". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ISSN 0300-9084.
- ^ PDBsum entry: Alpha Toxin
- ^ "Alpha Toxin - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 2023-11-29.
- ^ "Pfam entry: Type I phosphodiesterase". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ISSN 0022-2836.
- ^ "Pfam entry: Glycosyl transferases group 1". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ISSN 0959-440X.
- ^ "Pfam entry: Ferrochelatase". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ISSN 0969-2126.
- ^ "Pfam entry:Myotubularin-related". Archived from the original on 2007-09-26. Retrieved 2007-01-25.
- ^ "Emery and Rimoin's Principles and Practice of Medical Genetics". ScienceDirect. Retrieved 2023-11-29.
- ^ "Pfam entry:Dihydroorotate dehydrogenase". Archived from the original on 2007-09-26. Retrieved 2007-01-25.
- ISSN 0969-2126.
- ^ "Pfam entry: FMN-dependent dehydrogenase". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ^ "Glycolate oxidase - Proteopedia, life in 3D". proteopedia.org. Retrieved 2023-11-28.
- ^ "Pfam entry: Annexin". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ^ "Pfam entry Synapsin N". Archived from the original on 2007-09-26. Retrieved 2007-01-25.
- ^ "Pfam entry Synuclein". Archived from the original on 2007-09-26. Retrieved 2007-01-25.
- ^ "Pfam entry: Gla". Archived from the original on 2007-09-29. Retrieved 2007-01-25.
- ^ "Pfam entry Spectrin". Archived from the original on 2007-09-26. Retrieved 2007-01-25.
- ISBN 3-7643-6020-8.
- ^ Patocka J, Strunecka A (1999). "Sea Anemone Toxins". The ASA Newsletter. Archived from the original on 15 June 2013.
- PMID 10221874.
- PMID 11498029.
- PMID 14532141.
- ^ "Pfam entry Tachykinin". Archived from the original on 2007-09-26. Retrieved 2007-01-25.
Further reading
- Tamm LK, ed. (2005). Protein-Lipid Interactions: From Membrane Domains to Cellular Networks. Chichester: John Wiley & Sons. ISBN 3-527-31151-3.
- Cho W, Stahelin RV (June 2005). "Membrane-protein interactions in cell signaling and membrane trafficking". Annual Review of Biophysics and Biomolecular Structure. 34 (1): 119–151. PMID 15869386.
- Goñi FM (2002). "Non-permanent proteins in membranes: when proteins come as visitors (Review)". Molecular Membrane Biology. 19 (4): 237–245. S2CID 20892603.
- Johnson JE, Cornell RB (1999). "Amphitropic proteins: regulation by reversible membrane interactions (review)". Molecular Membrane Biology. 16 (3): 217–235. PMID 10503244.
- Seaton BA, Roberts MF (1996). "Peripheral membrane proteins". In Mertz K, Roux B (eds.). Biological Membranes. Boston, MA: Birkhauser. pp. 355–403.
- Benga G (1985). "Protein-lipid interactions in biological membranes". In Benga G (ed.). Structure and Properties of Biological Membranes. Vol. 1. Boca Raton, FL: CRC Press. pp. 159–188.
- Kessel A, Ben-Tal N (January 2002). "Free energy determinants of peptide association with lipid bilayers". Current Topics in Membranes. 52: 205–253. ISBN 9780121533526.
- Malmberg NJ, Falke JJ (2005). "Use of EPR power saturation to analyze the membrane-docking geometries of peripheral proteins: applications to C2 domains". Annual Review of Biophysics and Biomolecular Structure. 34 (1): 71–90. PMID 15869384.
- McIntosh TJ, Simon SA (2006). "Roles of bilayer material properties in function and distribution of membrane proteins". Annual Review of Biophysics and Biomolecular Structure. 35 (1): 177–198. PMID 16689633.
External links
- Peripheral membrane proteins in OPM database
- DOLOP Genomics-oriented database of bacterial lipoproteins
- Peptaibol database
- Antimicrobial Peptide Database Archived 2011-07-20 at the Wayback Machine
