Lactic acid bacteria
| Lactic acid bacteria | |
|---|---|
| |
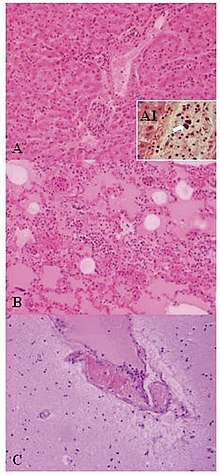
| Lesions of pulmonary alveoli (x10). C) encephalitis: congestion and marginalized neutrophils in nervous vessels (x10)
| |
| Scientific classification | |
| Domain: | Bacteria |
| Kingdom: | Bacillati |
| Phylum: | Bacillota |
| Class: | Bacilli |
| Order: | Lactobacillales Ludwig, Schleifer & Whitman 2010 |
| Families | |
| |
| Synonyms | |
| |
Lactobacillales are an order of
Production of lactic acid has linked LAB with
The genera that comprise the LAB are at its core Lactobacillus, Leuconostoc, Pediococcus, Lactococcus, and Streptococcus, as well as the more peripheral Aerococcus, Carnobacterium, Enterococcus, Oenococcus, Sporolactobacillus, Tetragenococcus, Vagococcus, and Weissella. All but Sporolactobacillus are members of the Lactobacillales order, and all are members of the Bacillota phylum.
Although lactic acid bacteria are generally associated with the order Lactobacillales, bacteria of the genus Bifidobacterium (phylum Actinomycetota) also produce lactic acid as the major product of carbohydrate metabolism.[1]
Characteristics
The lactic acid bacteria (LAB) are either rod-shaped (
Metabolism
LAB genera are classified in terms of two main pathways of hexose fermentation:
- Under conditions of excess
- glucose-6-phosphate is initially dehydrogenated to 6-phosphogluconate and subsequently decarboxylated to yield one mole of CO2. The resulting pentose-5-phosphate is cleaved into one mole glyceraldehyde phosphate (GAP) and one mole acetyl phosphate. GAP is further metabolized to lactate as in homofermentation, with the acetyl phosphate reduced to ethanol via acetyl-CoA and acetaldehyde intermediates. In theory, end products (including ATP) are produced in equimolar quantities from the catabolism of one mole of glucose. Obligate heterofermentative LAB include Leuconostoc, Oenococcus, Weissella, and group III lactobacilli [4]
Some members of Lactobacillus appear also able to perform
Streptococcus reclassification

In 1985, members of the diverse genus Streptococcus were reclassified into Lactococcus, Enterococcus, Vagococcus, and Streptococcus based on biochemical characteristics, as well as molecular features. Formerly, streptococci were segregated primarily based on serology, which has proven to correlate well with the current taxonomic definitions. Lactococci (formerly Lancefield group N streptococci) are used extensively as fermentation starters in dairy production, with humans estimated to consume 1018 (one billion billion) lactococci annually.[citation needed] Partly due to their industrial relevance, both L. lactis subspecies (L. l. lactis and L. l. cremoris) are widely used as generic LAB models for research. L. lactis ssp. cremoris, used in the production of hard cheeses, is represented by the laboratory strains LM0230 and MG1363. In similar manner, L. lactis ssp. lactis is employed in soft cheese fermentations, with the workhorse strain IL1403 ubiquitous in LAB research laboratories. In 2001, Bolotin et al. sequenced the genome of IL1403, which coincided with a significant shift of resources to understanding LAB genomics and related applications.
Phylogeny
The currently accepted taxonomy is based on the List of Prokaryotic names with Standing in Nomenclature (LPSN)[6] and National Center for Biotechnology Information (NCBI).[7]
| 16S rRNA based | 120 marker proteins based GTDB 09-RS220[11][12][13] | |||
|---|---|---|---|---|
|
Uses
Fermentation

Lactic acid bacteria are used in the food industry for a variety of reasons such as the production of cheese and yogurt products. Popular drinks such as kombucha are made using lactic acid bacteria, with kombucha having been known to have traces of Lactobacillus and Pediococcus once the drink is made.[14]
The beer and wine-making process utilizes certain lactic acid bacteria, mostly Lactobacillus. Lactic acid bacteria is used to start the wine-making process by starting the malolactic fermentation. After the malolactic fermentation, yeast cells are used to start the alcoholic fermentation process in grapes. The malolactic fermentation mechanism is mainly transformation of L-malic acid (dicarboxylic acid) to an lactic acid (monocarboxylic acid).[15] This change occurs due to the presence of malolactic and malic enzymes. All malic acid are degraded and this increase the pH levels which changes the taste of the wine.[15] Not only do they start the process but they are responsible for the different aromas produced in wine by the nutrients presence and the quality of the grapes. Also, the presence of different strains can change the desirability of aromas' presence. The different availability of enzymes that contribute to the vast spectrum of aromas in wine are associated with glycosidases, β-glucosidases, esterases, phenolic acid decarboxylases and citrate lyases.[16]
By using molecular biology, researchers can help pick out different desirable strains that help improve the quality of wine and help with the removable of the undesirable strains. The same can be said about brewing beer as well which uses yeast with some breweries using lactic acid bacteria to change the taste of their beer.[17]
Probiotics
Probiotics have been evaluated in research studies in animals and humans with respect to antibiotic-associated diarrhea, travellers' diarrhea, pediatric diarrhea,
Foods
The quest to find food ingredients with valuable
Some LAB produce bacteriocins which limit pathogens by interfering with cell wall synthesis or causing pore formation in the cell membrane.
Fertilizer
Researchers have studied the impact of lactic acid bacteria on indoleacetic acid production, phosphate solubilization, and nitrogen fixation on citrus. While most of the bacterial isolates were able to produce IAA, phosphate-solubilization was limited to only one of the eight LAB isolates.[26]
Management of bacteriophages in industry
A broad number of food products, commodity chemicals, and
Bacteriophage–host interaction
The first contact between an infecting phage and its bacterial host is the phage's attaching to the host cell. This attachment is mediated by the phage's receptor binding protein (RBP), which recognizes and binds to a receptor on the bacterial surface. RBPs are also referred to as host-specificity proteins, host determinants, and antireceptors. A variety of molecules have been suggested to act as host receptors for
Lactic acid bacteria and dental plaque
LAB are able to synthesize levans from sucrose, and dextrans from glucose.[28] Dextrans, like other glucan, enable bacteria to adhere to the surface of teeth, which in turn can cause tooth decay through the formation of dental plaque and production of lactic acid.[29] While the primary bacteria responsible for tooth decay is Streptococcus mutans, LAB do feature among the other most common oral bacteria that cause decay.[30]
Lactic acid bacteria genera
- Abiotrophia
- Aerococcus
- Carnobacterium
- Enterococcus
- Lactobacillus
- Lactococcus
- Leuconostoc
- Oenococcus
- Pediococcus
- Streptococcus
- Tetragenococcus
- Vagococcus
- Weissella
See also
- Gaffkaemia
- Malolactic fermentation
- Lactic acid fermentation
- Lacto-2 RNA motif
- List of bacterial orders
- List of bacteria genera
- Lacto-3 RNA motif
References
- PMID 25793197.
- ^ ISBN 978-1-904455-82-0.
- PMID 30169778.
- ^ .
- PMID 28063197.
- ^ J.P. Euzéby. "Lactobacillales". List of Prokaryotic names with Standing in Nomenclature (LPSN). Retrieved 2022-09-09.
- ^ Sayers; et al. "Lactobacillales". National Center for Biotechnology Information (NCBI) taxonomy database. Retrieved 2022-09-09.
- ^ "The LTP". Retrieved 10 December 2024.
- ^ "LTP_all tree in newick format". Retrieved 10 December 2024.
- ^ "LTP_10_2024 Release Notes" (PDF). Retrieved 10 December 2024.
- ^ "GTDB release 09-RS220". Genome Taxonomy Database. Retrieved 10 May 2024.
- ^ "bac120_r220.sp_labels". Genome Taxonomy Database. Retrieved 10 May 2024.
- ^ "Taxon History". Genome Taxonomy Database. Retrieved 10 May 2024.
- PMID 25763303.
- ^ S2CID 30267659.
- PMID 27940412.
- hdl:11250/2637117.
- ISBN 978-1-904455-01-1.
- ^ ISBN 978-1-904455-41-7.
- PMID 36271552.
- ISBN 978-1-904455-45-5.
- S2CID 25524132.
- ^ S2CID 20844172.
- ^ ISSN 0956-7135.
- PMID 24151598.
- PMID 27393998.
- ^ ISBN 978-1-904455-14-1.
- ISBN 978-0-19-539304-0.
- ISBN 978-0-13-144329-7.
- PMID 11699974.
Further reading
- Holzapfel WH, Wood BJ (1998). The genera of lactic acid bacteria (1st ed.). London Blackie Academic & Professional. ISBN 978-0-7514-0215-5.
- Salminen S, von Wright A, Ouwehand AC, eds. (2004). Lactic Acid Bacteria: Microbiological and Functional Aspects (3rd ed.). New York: Marcel Dekker, Inc. ISBN 978-0-8247-5332-0.
- Madigan MT, Martinko JM, Parker J (2004). Brock. Biología de los Microorganismos (10th ed.). Madrid: Pearson Educaciòn S.A. ISBN 978-84-205-3679-8.
External links
- "Lactic Acid Bacteria at MetaMicrobe: taxonomy, facts, probiotic properties, and references". Archived from the original on 2019-05-04.
- ^ "GTDB release 09-RS220". Genome Taxonomy Database. Retrieved 10 May 2024.
- ^ "bac120_r220.sp_labels". Genome Taxonomy Database. Retrieved 10 May 2024.
- ^ "Taxon History". Genome Taxonomy Database. Retrieved 10 May 2024.
- ^ "The LTP". Retrieved 10 December 2024.
- ^ "LTP_all tree in newick format". Retrieved 10 December 2024.
- ^ "LTP_10_2024 Release Notes" (PDF). Retrieved 10 December 2024.
