Metagenomics
| Part of a series on |
| Genetics |
|---|
 |

Metagenomics is the study of genetic material recovered directly from environmental or clinical samples by a method called sequencing. The broad field may also be referred to as environmental genomics, ecogenomics, community genomics or microbiomics.
While traditional
Because of its ability to reveal the previously hidden diversity of microscopic life, metagenomics offers a powerful way of understanding the microbial world that might revolutionize understanding of biology.[3] As the price of DNA sequencing continues to fall, metagenomics now allows microbial ecology to be investigated at a much greater scale and detail than before. Recent studies use either "shotgun" or PCR directed sequencing to get largely unbiased samples of all genes from all the members of the sampled communities.[4]
Etymology
The term "metagenomics" was first used by Jo Handelsman, Robert M. Goodman, Michelle R. Rondon, Jon Clardy, and Sean F. Brady, and first appeared in publication in 1998.[5] The term metagenome referenced the idea that a collection of genes sequenced from the environment could be analyzed in a way analogous to the study of a single genome. In 2005, Kevin Chen and Lior Pachter (researchers at the University of California, Berkeley) defined metagenomics as "the application of modern genomics technique without the need for isolation and lab cultivation of individual species".[6]
History
| Part of a series on |
| DNA barcoding |
|---|
 |
| By taxa |
| Other |
|
Conventional
In the 1980s early
In 2002,
Beginning in 2003, Craig Venter, leader of the privately funded parallel of the Human Genome Project, has led the Global Ocean Sampling Expedition (GOS), circumnavigating the globe and collecting metagenomic samples throughout the journey. All of these samples were sequenced using shotgun sequencing, in hopes that new genomes (and therefore new organisms) would be identified. The pilot project, conducted in the Sargasso Sea, found DNA from nearly 2000 different species, including 148 types of bacteria never before seen.[15] Venter thoroughly explored the West Coast of the United States, and completed a two-year expedition in 2006 to explore the Baltic, Mediterranean, and Black Seas. Analysis of the metagenomic data collected during this journey revealed two groups of organisms, one composed of taxa adapted to environmental conditions of 'feast or famine', and a second composed of relatively fewer but more abundantly and widely distributed taxa primarily composed of plankton.[16]
In 2005 Stephan C. Schuster at
Sequencing
Recovery of DNA sequences longer than a few thousand base pairs from environmental samples was very difficult until recent advances in molecular biological techniques allowed the construction of libraries in bacterial artificial chromosomes (BACs), which provided better vectors for molecular cloning.[20]
Shotgun metagenomics
Advances in
High-throughput sequencing
An advantage to high throughput sequencing is that this technique does not require cloning the DNA before sequencing, removing one of the main biases and bottlenecks in environmental sampling. The first metagenomic studies conducted using
An emerging approach combines shotgun sequencing and chromosome conformation capture (Hi-C), which measures the proximity of any two DNA sequences within the same cell, to guide microbial genome assembly.[25] Long read sequencing technologies, including PacBio RSII and PacBio Sequel by Pacific Biosciences, and Nanopore MinION, GridION, PromethION by Oxford Nanopore Technologies, is another choice to get long shotgun sequencing reads that should make ease in assembling process.[26]
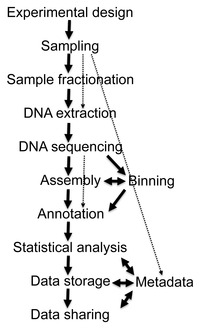
Bioinformatics
This section is missing information about quality assessment: on assembly (N50, MetaQUAST), on genome (universal single-copy marker genes – CheckM and BUSCO). (February 2022) |
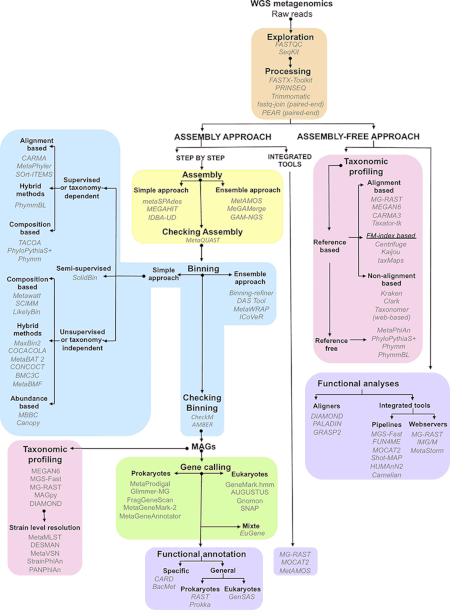
The data generated by metagenomics experiments are both enormous and inherently noisy, containing fragmented data representing as many as 10,000 species.
Sequence pre-filtering
The first step of metagenomic data analysis requires the execution of certain pre-filtering steps, including the removal of redundant, low-quality sequences and sequences of probable
Assembly
DNA sequence data from genomic and metagenomic projects are essentially the same, but genomic sequence data offers higher
There are several assembly programs, most of which can use information from
Gene prediction
Metagenomic analysis pipelines use two approaches in the annotation of coding regions in the assembled contigs.[37] The first approach is to identify genes based upon homology with genes that are already publicly available in sequence databases, usually by BLAST searches. This type of approach is implemented in the program MEGAN4.[41] The second, ab initio, uses intrinsic features of the sequence to predict coding regions based upon gene training sets from related organisms. This is the approach taken by programs such as GeneMark[42] and GLIMMER. The main advantage of ab initio prediction is that it enables the detection of coding regions that lack homologs in the sequence databases; however, it is most accurate when there are large regions of contiguous genomic DNA available for comparison.[1]
Species diversity
Gene annotations provide the "what", while measurements of
Data integration
The massive amount of exponentially growing sequence data is a daunting challenge that is complicated by the complexity of the
Several tools have been developed to integrate metadata and sequence data, allowing downstream comparative analyses of different datasets using a number of ecological indices. In 2007, Folker Meyer and Robert Edwards and a team at Argonne National Laboratory and the University of Chicago released the Metagenomics Rapid Annotation using Subsystem Technology server (MG-RAST) a community resource for metagenome data set analysis.[51] As of June 2012 over 14.8 terabases (14x1012 bases) of DNA have been analyzed, with more than 10,000 public data sets freely available for comparison within MG-RAST. Over 8,000 users now have submitted a total of 50,000 metagenomes to MG-RAST. The Integrated Microbial Genomes/Metagenomes (IMG/M) system also provides a collection of tools for functional analysis of microbial communities based on their metagenome sequence, based upon reference isolate genomes included from the Integrated Microbial Genomes (IMG) system and the Genomic Encyclopedia of Bacteria and Archaea (GEBA) project.[52]
One of the first standalone tools for analysing high-throughput metagenome shotgun data was MEGAN (MEta Genome ANalyzer).[41][44] A first version of the program was used in 2005 to analyse the metagenomic context of DNA sequences obtained from a mammoth bone.[17] Based on a BLAST comparison against a reference database, this tool performs both taxonomic and functional binning, by placing the reads onto the nodes of the NCBI taxonomy using a simple lowest common ancestor (LCA) algorithm or onto the nodes of the SEED or KEGG classifications, respectively.[53]
With the advent of fast and inexpensive sequencing instruments, the growth of databases of DNA sequences is now exponential (e.g., the NCBI GenBank database [54]). Faster and efficient tools are needed to keep pace with the high-throughput sequencing, because the BLAST-based approaches such as MG-RAST or MEGAN run slowly to annotate large samples (e.g., several hours to process a small/medium size dataset/sample [55]). Thus, ultra-fast classifiers have recently emerged, thanks to more affordable powerful servers. These tools can perform the taxonomic annotation at extremely high speed, for example CLARK [56] (according to CLARK's authors, it can classify accurately "32 million metagenomic short reads per minute"). At such a speed, a very large dataset/sample of a billion short reads can be processed in about 30 minutes.
With the increasing availability of samples containing ancient DNA and due to the uncertainty associated with the nature of those samples (ancient DNA damage),[57] a fast tool capable of producing conservative similarity estimates has been made available. According to FALCON's authors, it can use relaxed thresholds and edit distances without affecting the memory and speed performance.
Comparative metagenomics
Comparative analyses between metagenomes can provide additional insight into the function of complex microbial communities and their role in host health.[58] Pairwise or multiple comparisons between metagenomes can be made at the level of sequence composition (comparing GC-content or genome size), taxonomic diversity, or functional complement. Comparisons of population structure and phylogenetic diversity can be made on the basis of 16S rRNA and other phylogenetic marker genes, or—in the case of low-diversity communities—by genome reconstruction from the metagenomic dataset.[59] Functional comparisons between metagenomes may be made by comparing sequences against reference databases such as COG or KEGG, and tabulating the abundance by category and evaluating any differences for statistical significance.[53] This gene-centric approach emphasizes the functional complement of the community as a whole rather than taxonomic groups, and shows that the functional complements are analogous under similar environmental conditions.[59] Consequently, metadata on the environmental context of the metagenomic sample is especially important in comparative analyses, as it provides researchers with the ability to study the effect of habitat upon community structure and function.[1]
Additionally, several studies have also utilized oligonucleotide usage patterns to identify the differences across diverse microbial communities. Examples of such methodologies include the dinucleotide relative abundance approach by Willner et al.[60] and the HabiSign approach of Ghosh et al.[61] This latter study also indicated that differences in tetranucleotide usage patterns can be used to identify genes (or metagenomic reads) originating from specific habitats. Additionally some methods as TriageTools[62] or Compareads[63] detect similar reads between two read sets. The similarity measure they apply on reads is based on a number of identical words of length k shared by pairs of reads.
A key goal in comparative metagenomics is to identify microbial group(s) which are responsible for conferring specific characteristics to a given environment. However, due to issues in the sequencing technologies artifacts need to be accounted for like in metagenomeSeq.
Data analysis
Community metabolism
In many bacterial communities, natural or engineered (such as
Metatranscriptomics
Metagenomics allows researchers to access the functional and metabolic diversity of microbial communities, but it cannot show which of these processes are active.
Viruses
Metagenomic sequencing is particularly useful in the study of viral communities. As viruses lack a shared universal phylogenetic marker (as
Applications
Metagenomics has the potential to advance knowledge in a wide variety of fields. It can also be applied to solve practical challenges in medicine, engineering, agriculture, sustainability and ecology.[31][76]
Agriculture
The
Biofuel
The
Biotechnology
Microbial communities produce a vast array of biologically active chemicals that are used in competition and communication.
Two types of analysis are used in the bioprospecting of metagenomic data: function-driven screening for an expressed trait, and sequence-driven screening for DNA sequences of interest.[88] Function-driven analysis seeks to identify clones expressing a desired trait or useful activity, followed by biochemical characterization and sequence analysis. This approach is limited by availability of a suitable screen and the requirement that the desired trait be expressed in the host cell. Moreover, the low rate of discovery (less than one per 1,000 clones screened) and its labor-intensive nature further limit this approach.[89] In contrast, sequence-driven analysis uses conserved DNA sequences to design PCR primers to screen clones for the sequence of interest.[88] In comparison to cloning-based approaches, using a sequence-only approach further reduces the amount of bench work required. The application of massively parallel sequencing also greatly increases the amount of sequence data generated, which require high-throughput bioinformatic analysis pipelines.[89] The sequence-driven approach to screening is limited by the breadth and accuracy of gene functions present in public sequence databases. In practice, experiments make use of a combination of both functional and sequence-based approaches based upon the function of interest, the complexity of the sample to be screened, and other factors.[89][90] An example of success using metagenomics as a biotechnology for drug discovery is illustrated with the malacidin antibiotics.[91]
Ecology

Metagenomics can provide valuable insights into the functional ecology of environmental communities.[92] Metagenomic analysis of the bacterial consortia found in the defecations of Australian sea lions suggests that nutrient-rich sea lion faeces may be an important nutrient source for coastal ecosystems. This is because the bacteria that are expelled simultaneously with the defecations are adept at breaking down the nutrients in the faeces into a bioavailable form that can be taken up into the food chain.[93]
DNA sequencing can also be used more broadly to identify species present in a body of water,[94] debris filtered from the air, sample of dirt, or animal's faeces,[95] and even detect diet items from blood meals.[96] This can establish the range of invasive species and endangered species, and track seasonal populations.
Environmental remediation
Metagenomics can improve strategies for monitoring the impact of pollutants on ecosystems and for cleaning up contaminated environments. Increased understanding of how microbial communities cope with pollutants improves assessments of the potential of contaminated sites to recover from pollution and increases the chances of bioaugmentation or biostimulation trials to succeed.[97]
Gut microbe characterization
Another medical study as part of the MetaHit (Metagenomics of the Human Intestinal Tract) project consisted of 124 individuals from Denmark and Spain consisting of healthy, overweight, and irritable bowel disease patients.[100] The study attempted to categorize the depth and phylogenetic diversity of gastrointestinal bacteria. Using Illumina GA sequence data and SOAPdenovo, a de Bruijn graph-based tool specifically designed for assembly short reads, they were able to generate 6.58 million contigs greater than 500 bp for a total contig length of 10.3 Gb and a N50 length of 2.2 kb.
The study demonstrated that two bacterial divisions, Bacteroidetes and Firmicutes, constitute over 90% of the known phylogenetic categories that dominate distal gut bacteria. Using the relative gene frequencies found within the gut these researchers identified 1,244 metagenomic clusters that are critically important for the health of the intestinal tract. There are two types of functions in these range clusters: housekeeping and those specific to the intestine. The housekeeping gene clusters are required in all bacteria and are often major players in the main metabolic pathways including central carbon metabolism and amino acid synthesis. The gut-specific functions include adhesion to host proteins and the harvesting of sugars from globoseries glycolipids. Patients with irritable bowel syndrome were shown to exhibit 25% fewer genes and lower bacterial diversity than individuals not suffering from irritable bowel syndrome indicating that changes in patients' gut biome diversity may be associated with this condition.[100]
While these studies highlight some potentially valuable medical applications, only 31–48.8% of the reads could be aligned to 194 public human gut bacterial genomes and 7.6–21.2% to bacterial genomes available in GenBank which indicates that there is still far more research necessary to capture novel bacterial genomes.[101]
In the Human Microbiome Project (HMP), gut microbial communities were assayed using high-throughput DNA sequencing. HMP showed that, unlike individual microbial species, many metabolic processes were present among all body habitats with varying frequencies. Microbial communities of 649 metagenomes drawn from seven primary body sites on 102 individuals were studied as part of the human microbiome project. The metagenomic analysis revealed variations in niche specific abundance among 168 functional modules and 196 metabolic pathways within the microbiome. These included glycosaminoglycan degradation in the gut, as well as phosphate and amino acid transport linked to host phenotype (vaginal pH) in the posterior fornix. The HMP has brought to light the utility of metagenomics in diagnostics and evidence-based medicine. Thus metagenomics is a powerful tool to address many of the pressing issues in the field of personalized medicine.[102]
In animals, metagenomics can be used to profile their gut microbiomes and enable detection of antibiotic-resistant bacteria.[103] This can have implications in monitoring the spread of diseases from wildlife to farmed animals and humans.
Infectious disease diagnosis
Differentiating between infectious and non-infectious illness, and identifying the underlying etiology of infection, can be challenging. For example, more than half of cases of encephalitis remain undiagnosed, despite extensive testing using state-of-the-art clinical laboratory methods. Clinical metagenomic sequencing shows promise as a sensitive and rapid method to diagnose infection by comparing genetic material found in a patient's sample to databases of all known microscopic human pathogens and thousands of other bacterial, viral, fungal, and parasitic organisms and databases on antimicrobial resistances gene sequences with associated clinical phenotypes.[104]
Arbovirus surveillance
Metagenomics has been an invaluable tool to help characterise the diversity and ecology of pathogens that are vectored by
See also
References
- ^ a b c d e f g
Wooley JC, Godzik A, Friedberg I (February 2010). Bourne PE (ed.). "A primer on metagenomics". PLOS Computational Biology. 6 (2): e1000667. PMID 20195499.
- ^ a b
Hugenholtz P, Goebel BM, Pace NR (September 1998). "Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity". Journal of Bacteriology. 180 (18): 4765–74. PMID 9733676.
- ^
Marco, D, ed. (2011). Metagenomics: Current Innovations and Future Trends. ISBN 978-1-904455-87-5.
- ^
Eisen JA (March 2007). "Environmental shotgun sequencing: its potential and challenges for studying the hidden world of microbes". PLOS Biology. 5 (3): e82. PMID 17355177.
- ^
Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM (October 1998). "Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products". Chemistry & Biology. 5 (10): R245-9. PMID 9818143..
- ^
Chen K, Pachter L (July 2005). "Bioinformatics for whole-genome shotgun sequencing of microbial communities". PLOS Computational Biology. 1 (2): 106–12. PMID 16110337.
- ^
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (October 1985). "Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses". Proceedings of the National Academy of Sciences of the United States of America. 82 (20): 6955–9. PMID 2413450.
- ISBN 978-1-4757-0611-6.
- ^
PMID 2066334.
- ^
Healy FG, Ray RM, Aldrich HC, Wilkie AC, Ingram LO, Shanmugam KT (1995). "Direct isolation of functional genes encoding cellulases from the microbial consortia in a thermophilic, anaerobic digester maintained on lignocellulose". Applied Microbiology and Biotechnology. 43 (4): 667–74. S2CID 31384119.
- ^
Stein JL, Marsh TL, Wu KY, Shizuya H, DeLong EF (February 1996). "Characterization of uncultivated prokaryotes: isolation and analysis of a 40-kilobase-pair genome fragment from a planktonic marine archaeon". Journal of Bacteriology. 178 (3): 591–9. PMID 8550487.
- ^
Breitbart M, Salamon P, Andresen B, Mahaffy JM, Segall AM, Mead D, et al. (October 2002). "Genomic analysis of uncultured marine viral communities". Proceedings of the National Academy of Sciences of the United States of America. 99 (22): 14250–5. PMID 12384570.
- ^ a b
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM, et al. (March 2004). "Community structure and metabolism through reconstruction of microbial genomes from the environment". Nature. 428 (6978): 37–43. S2CID 4420754.(subscription required)
- ^
Hugenholtz P (2002). "Exploring prokaryotic diversity in the genomic era". Genome Biology. 3 (2): REVIEWS0003. PMID 11864374.
- ^
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA, et al. (April 2004). "Environmental genome shotgun sequencing of the Sargasso Sea". Science. 304 (5667): 66–74. S2CID 1454587.
- ^
Yooseph S, Nealson KH, Rusch DB, McCrow JP, Dupont CL, Kim M, et al. (November 2010). "Genomic and functional adaptation in surface ocean planktonic prokaryotes". Nature. 468 (7320): 60–6. PMID 21048761.(subscription required)
- ^ a b c
Poinar HN, Schwarz C, Qi J, Shapiro B, Macphee RD, Buigues B, et al. (January 2006). "Metagenomics to paleogenomics: large-scale sequencing of mammoth DNA". Science. 311 (5759): 392–4. S2CID 11238470.
- ^
Edwards RA, Rodriguez-Brito B, Wegley L, Haynes M, Breitbart M, Peterson DM, et al. (March 2006). "Using pyrosequencing to shed light on deep mine microbial ecology". BMC Genomics. 7: 57. PMID 16549033.
- PMID 22587947.
- ^
Béjà O, Suzuki MT, Koonin EV, Aravind L, Hadd A, Nguyen LP, et al. (October 2000). "Construction and analysis of bacterial artificial chromosome libraries from a marine microbial assemblage". Environmental Microbiology. 2 (5): 516–29. S2CID 8267748.
- ^ a b
Segata N, Boernigen D, Tickle TL, Morgan XC, Garrett WS, Huttenhower C (May 2013). "Computational meta'omics for microbial community studies". Molecular Systems Biology. 9 (666): 666. PMID 23670539.
- PMID 20676378.
- S2CID 1465786.
- .
- PMID 29491419.
- PMID 27383682.
- PMID 32706331.
- ^ a b
Hess M, Sczyrba A, Egan R, Kim TW, Chokhawala H, Schroth G, et al. (January 2011). "Metagenomic discovery of biomass-degrading genes and genomes from cow rumen". Science. 331 (6016): 463–7. S2CID 36572885.
- ^
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, et al. (March 2010). "A human gut microbial gene catalogue established by metagenomic sequencing". Nature. 464 (7285): 59–65. PMID 20203603.(subscription required)
- ^ a b
Paulson JN, Stine OC, Bravo HC, Pop M (December 2013). "Differential abundance analysis for microbial marker-gene surveys". Nature Methods. 10 (12): 1200–2. PMID 24076764.
- ^ a b c d e f g
Committee on Metagenomics: Challenges and Functional Applications, National Research Council (2007). The New Science of Metagenomics: Revealing the Secrets of Our Microbial Planet. Washington, D.C.: The National Academies Press. PMID 21678629.
- PMID 25983555.
- PMID 22384016.
- PMID 23376350.
- S2CID 25857874.
- PMID 21408061.
- ^ PMID 19052320.
- PMID 22821567.
- PMID 18349386.
- PMID 24855317.
- ^ a b
Huson DH, Mitra S, Ruscheweyh HJ, Weber N, Schuster SC (September 2011). "Integrative analysis of environmental sequences using MEGAN4". Genome Research. 21 (9): 1552–60. PMID 21690186.
- ^
Zhu W, Lomsadze A, Borodovsky M (July 2010). "Ab initio gene identification in metagenomic sequences". Nucleic Acids Research. 38 (12): e132. PMID 20403810.
- PMID 19657372.
- ^ a b
Huson DH, Auch AF, Qi J, Schuster SC (March 2007). "MEGAN analysis of metagenomic data". Genome Research. 17 (3): 377–86. PMID 17255551.
- ^
Segata N, Waldron L, Ballarini A, Narasimhan V, Jousson O, Huttenhower C (June 2012). "Metagenomic microbial community profiling using unique clade-specific marker genes". Nature Methods. 9 (8): 811–4. PMID 22688413.
- ^
Sunagawa S, Mende DR, Zeller G, Izquierdo-Carrasco F, Berger SA, Kultima JR, et al. (December 2013). "Metagenomic species profiling using universal phylogenetic marker genes". Nature Methods. 10 (12): 1196–9. S2CID 7728395.
- ^ a b
Milanese A, Mende DR, Paoli L, Salazar G, Ruscheweyh HJ, Cuenca M, et al. (March 2019). "Microbial abundance, activity and population genomic profiling with mOTUs2". Nature Communications. 10 (1): 1014. PMID 30833550.
- ^
Liu B, Gibbons T, Ghodsi M, Treangen T, Pop M (2011). "Accurate and fast estimation of taxonomic profiles from metagenomic shotgun sequences". BMC Genomics. 12 (Suppl 2): S4. PMID 21989143.
- ^
Dadi TH, Renard BY, Wieler LH, Semmler T, Reinert K (2017). "SLIMM: species level identification of microorganisms from metagenomes". PeerJ. 5: e3138. PMID 28367376.
- ^
Pagani I, Liolios K, Jansson J, Chen IM, Smirnova T, Nosrat B, et al. (January 2012). "The Genomes OnLine Database (GOLD) v.4: status of genomic and metagenomic projects and their associated metadata". Nucleic Acids Research. 40 (Database issue): D571-9. PMID 22135293.
- ^
Meyer F, Paarmann D, D'Souza M, Olson R, Glass EM, Kubal M, et al. (September 2008). "The metagenomics RAST server - a public resource for the automatic phylogenetic and functional analysis of metagenomes". BMC Bioinformatics. 9: 386. PMID 18803844.
- PMID 22086953.
- ^ PMID 21342551.
- ^
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (January 2013). "GenBank". Nucleic Acids Research. 41 (Database issue): D36-42. PMID 23193287.
- ^
Bazinet AL, Cummings MP (May 2012). "A comparative evaluation of sequence classification programs". BMC Bioinformatics. 13: 92. PMID 22574964.
- ^
Ounit R, Wanamaker S, Close TJ, Lonardi S (March 2015). "CLARK: fast and accurate classification of metagenomic and genomic sequences using discriminative k-mers". BMC Genomics. 16 (1): 236. PMID 25879410.
- bioRxiv 10.1101/267179.
- ^
Kurokawa K, Itoh T, Kuwahara T, Oshima K, Toh H, Toyoda A, et al. (August 2007). "Comparative metagenomics revealed commonly enriched gene sets in human gut microbiomes". DNA Research. 14 (4): 169–81. PMID 17916580.
- ^ PMID 21169428.
- PMID 19302541.
- PMID 22373355.
- ^
Fimereli D, Detours V, Konopka T (April 2013). "TriageTools: tools for partitioning and prioritizing analysis of high-throughput sequencing data". Nucleic Acids Research. 41 (7): e86. PMID 23408855.
- PMID 23282463.
- PMID 23978768.
- ^
Werner JJ, Knights D, Garcia ML, Scalfone NB, Smith S, Yarasheski K, et al. (March 2011). "Bacterial community structures are unique and resilient in full-scale bioenergy systems". Proceedings of the National Academy of Sciences of the United States of America. 108 (10): 4158–63. PMID 21368115.
- ^
McInerney MJ, Sieber JR, Gunsalus RP (December 2009). "Syntrophy in anaerobic global carbon cycles". Current Opinion in Biotechnology. 20 (6): 623–32. PMID 19897353.
- PMID 21592777.
- S2CID 4380804.
- S2CID 4466854.
- PMID 27799466.
- PMID 30407573.
- S2CID 2127494.
- ^
Kristensen DM, Mushegian AR, Dolja VV, Koonin EV (January 2010). "New dimensions of the virus world discovered through metagenomics". Trends in Microbiology. 18 (1): 11–9. PMID 19942437.
- ^
Kerepesi C, Grolmusz V (March 2016). "Giant viruses of the Kutch Desert". Archives of Virology. 161 (3): 721–4. S2CID 13145926.
- ^
Kerepesi C, Grolmusz V (June 2017). "The "Giant Virus Finder" discovers an abundance of giant viruses in the Antarctic dry valleys". Archives of Virology. 162 (6): 1671–1676. S2CID 1925728.
- ^ Copeland CS (September–October 2017). "The World Within Us" (PDF). Healthcare Journal of New Orleans: 21–26.
- ^ Jansson J (2011). "Towards "Tera-Terra": Terabase Sequencing of Terrestrial Metagenomes Print E-mail". Microbe. Vol. 6, no. 7. p. 309. Archived from the original on 31 March 2012.
- .
- ^ "TerraGenome Homepage". TerraGenome international sequencing consortium. Retrieved 30 December 2011.
- ^ a b Committee on Metagenomics: Challenges and Functional Applications, National Research Council (2007). Understanding Our Microbial Planet: The New Science of Metagenomics (PDF). The National Academies Press. Archived from the original (PDF) on 30 October 2012. Retrieved 30 December 2011.
- ISBN 978-1-904455-54-7.
- PMID 26052316.
- ^
Li LL, McCorkle SR, Monchy S, Taghavi S, van der Lelie D (May 2009). "Bioprospecting metagenomes: glycosyl hydrolases for converting biomass". Biotechnology for Biofuels. 2: 10. PMID 19450243.
- ^
Jaenicke S, Ander C, Bekel T, Bisdorf R, Dröge M, Gartemann KH, et al. (January 2011). Aziz RK (ed.). "Comparative and joint analysis of two metagenomic datasets from a biogas fermenter obtained by 454-pyrosequencing". PLOS ONE. 6 (1): e14519. PMID 21297863.
- ^
Suen G, Scott JJ, Aylward FO, Adams SM, Tringe SG, Pinto-Tomás AA, et al. (September 2010). Sonnenburg J (ed.). "An insect herbivore microbiome with high plant biomass-degrading capacity". PLOS Genetics. 6 (9): e1001129. PMID 20885794.
- PMID 19760178.
- ISBN 978-1-904455-54-7.
- ^ PMID 12849784. Archived from the original(PDF) on 4 March 2016. Retrieved 20 January 2012.
- ^ a b c
Kakirde KS, Parsley LC, Liles MR (November 2010). "Size Does Matter: Application-driven Approaches for Soil Metagenomics". Soil Biology & Biochemistry. 42 (11): 1911–1923. PMID 21076656.
- ^
Parachin NS, Gorwa-Grauslund MF (May 2011). "Isolation of xylose isomerases by sequence- and function-based screening from a soil metagenomic library". Biotechnology for Biofuels. 4 (1): 9. PMID 21545702.
- PMID 29434326.
- PMID 21407210.
- PMID 22606263.
- ^ "What's Swimming in the River? Just Look For DNA". NPR.org. 24 July 2013. Retrieved 10 October 2014.
- PMID 33971086.
- S2CID 248041252.
- ^
George I, Stenuit B, Agathos SN (2010). "Application of Metagenomics to Bioremediation". In Marco D (ed.). Metagenomics: Theory, Methods and Applications. Caister Academic Press. ISBN 978-1-904455-54-7.
- ^ Zimmer C (13 July 2010). "How Microbes Defend and Define Us". New York Times. Retrieved 29 December 2011.
- ^
Nelson KE and White BA (2010). "Metagenomics and Its Applications to the Study of the Human Microbiome". Metagenomics: Theory, Methods and Applications. Caister Academic Press. ISBN 978-1-904455-54-7.
- ^ PMID 20203603.
- PMID 20203603.
- PMID 22719234.
- S2CID 248739527.
- PMID 30918369.
- PMID 29549363.
- PMID 32878948.
- PMID 34442732.
- PMID 31852942.
- PMID 28855093.
External links
- Focus on Metagenomics at Nature Reviews Microbiology journal website
- The “Critical Assessment of Metagenome Interpretation” (CAMI) initiative to evaluate methods in metagenomics
