Microbiome
| Part of a series on |
| Microbiomes |
|---|
 |
A microbiome (from
The
All animals and plants form associations with microorganisms, including protists, bacteria, archaea, fungi, and viruses. In the ocean, animal–microbial relationships were historically explored in single host–symbiont systems. However, new explorations into the diversity of microorganisms associating with diverse marine animal hosts is moving the field into studies that address interactions between the animal host and the multi-member microbiome. The potential for microbiomes to influence the health, physiology, behaviour, and ecology of marine animals could alter current understandings of how marine animals adapt to change. This applies to especially the growing climate-related and anthropogenic-induced changes already impacting the ocean. The
Microbiome research originated in microbiology back in the seventeenth century. The development of new techniques and equipment boosted microbiological research and caused paradigm shifts in understanding health and disease.
Background
History
Microbiome research originated in microbiology and started back in the seventeenth century. The development of new techniques and equipment has boosted microbiological research and caused paradigm shifts in understanding health and disease. Since infectious diseases have affected human populations throughout most of history, medical microbiology was the earliest focus of research and public interest. Additionally, food microbiology is an old field of empirical applications. The development of the first microscopes allowed the discovery of a new, unknown world and led to the identification of microorganisms.[2]
- Paradigm shift
-
Shift of paradigm from microbes as unsocial organisms causing diseases to a holistic view of microorganisms as the centre of theOne Health Concept interconnecting all areas of human lives.[2]
Access to the previously invisible world opened the eyes and the minds of the researchers of the seventeenth century.
However, comprehensive research over the past century has shown only a small proportion of microorganisms are associated with disease or pathogenicity. The overwhelming majority of
Subsequently, the concept that microorganisms exist as single cells began to change as it became increasingly obvious that microbes occur within complex assemblages in which species interactions and communication are critical to population dynamics and functional activities.
A further important step was the introduction of
Another major paradigm shift was initiated at the beginning of this century and continues through today, as new sequencing technologies and accumulated sequence data have highlighted both the ubiquity of
Etymology
The word microbiome (from the Greek micro meaning "small" and bíos meaning "life") was first used by J.L. Mohr in 1952 in The Scientific Monthly to mean the microorganisms found in a specific environment.[59][60]
Definitions
Microbial communities have commonly been defined as the collection of microorganisms living together. More specifically, microbial communities are defined as multi-species assemblages, in which (micro) organisms interact with each other in a contiguous environment.[61] In 1988, Whipps and colleagues working on the ecology of rhizosphere microorganisms provided the first definition of the term microbiome.[62] They described the microbiome as a combination of the words micro and biome, naming a "characteristic microbial community" in a "reasonably well-defined habitat which has distinct physio-chemical properties" as their "theatre of activity". This definition represents a substantial advancement of the definition of a microbial community, as it defines a microbial community with distinct properties and functions and its interactions with its environment, resulting in the formation of specific ecological niches.[2]
However, many other microbiome definitions have been published in recent decades. By 2020 the most cited definition was by
| Microbiome definitions[2] | |
|---|---|
| Definition type | Examples |
| Ecological | Definitions based on ecology describe the microbiome following the concepts derived from the ecology of multicellular organisms. The main issue here is that the theories from the macro-ecology do not always fit the rules in the microbial world. |
| |
| Organisms/host-dependent | The host-dependent definitions are based on the microbial interactions with the host. The main gaps here concern the question whether the microbial-host interaction data gained from one host can be transferred to another. The understanding of coevolution and selection in the host-dependent definitions is also underrepresented. |
| |
| Genomic/ method-driven | There is a variety of microbiome definitions available that are driven by the methods applied. Mostly, these definitions rely on DNA sequence-based analysis and describe microbiome as a collective genome of microorganisms in a specific environment. The main bottleneck here is that every new available technology will result in a need for a new definition. |
| |
| Combined | There are some microbiome definitions available that fit several categories with their advantages and disadvantages. |
| |
In 2020, a panel of international experts, organised by the EU-funded MicrobiomeSupport project,[76] published the results of their deliberations on the definition of the microbiome.[2] The panel was composed of about 40 leaders from diverse microbiome areas, and about one hundred further experts from around the world contributed through an online survey. They proposed a definition of the microbiome based on a revival of what they characterised as the "compact, clear, and comprehensive description of the term" as originally provided by Whipps et al. in 1988,[62] amended with a set of recommendations considering subsequent technological developments and research findings. They clearly separate the terms microbiome and microbiota and provide a comprehensive discussion considering the composition of microbiota, the heterogeneity and dynamics of microbiomes in time and space, the stability and resilience of microbial networks, the definition of core microbiomes, and functionally relevant keystone species as well as co-evolutionary principles of microbe-host and inter-species interactions within the microbiome.[2]
The panel extended the Whipps et al. definition, which contains all important points that are valid even 30 years after its publication in 1988, by two explanatory paragraphs differentiating the terms microbiome and microbiota and pronouncing its dynamic character, as follows:
- The microbiome is defined as a characteristic microbial community occupying a reasonable well-defined habitat which has distinct physio-chemical properties. The microbiome not only refers to the microorganisms involved but also encompass their theatre of activity, which results in the formation of specific ecological niches. The microbiome, which forms a dynamic and interactive micro-ecosystem prone to change in time and scale, is integrated in macro-ecosystems including eukaryotic hosts, and here crucial for their functioning and health.[2]
- The microbiota consists of the assembly of microorganisms belonging to different kingdoms (prokaryotes (bacteria, archaea), eukaryotes (algae, protozoa, fungi etc), while "their theatre of activity" includes microbial structures, metabolites, mobile genetic elements (such as transposons, phages, and viruses), and relic DNA embedded in the environmental conditions of the habitat.[2]
Membership
Microbiota
The microbiota comprises all living members forming the microbiome. Most microbiome researchers agree bacteria, archaea, fungi, algae, and small protists should be considered as members of the microbiome.
When it comes to the use of specific terms, a clear differentiation between microbiome and microbiota helps to avoid the controversy concerning the members of a microbiome.[2] Microbiota is usually defined as the assemblage of living microorganisms present in a defined environment.[64] As phages, viruses, plasmids, prions, viroids, and free DNA are usually not considered as living microorganisms,[80] they do not belong to the microbiota.[2]
The term microbiome, as it was originally postulated by Whipps and coworkers,
Microbiome studies sometimes focus on the behaviour of a specific group of microbiota, generally in relation to or justified by a clear hypothesis. More and more terms like
Networks
-
Co-occurrence networks help visualising microbial interactions
Nodes usually represent taxa of microorganisms, and edges represent statistically significant associations between nodes.[2]
–––––––––––––––––––––––––––
Testing of the hypotheses resulted from the network analyses is required for a comprehensive study of microbial interactions.[2]
Microbes interact with one another, and these symbiotic interactions have diverse consequences for microbial fitness, population dynamics, and functional capacities within the microbiome.
Microbiomes exhibit different adaptive strategies.[2] Oligotrophs are organisms that can live in an environment offering very low levels of nutrients, particularly carbon. They are characterised by slow growth, low rates of metabolism, and generally low population density. Oligotrophic environments include deep oceanic sediments, caves, glacial and polar ice, deep subsurface soil, aquifers, ocean waters, and leached soils. In contrast are the copiotrophs, which thrive in much higher carbon concentrations, and do well in high organic substrate conditions such as sewage lagoons.[85][86]
In addition to oligotrophic and copiotrophic strategists, the
Coevolution
- Shift in the understanding of the microbial-host coevolution
-
from "separation" theories to a holistic approachIn a holistic approach, the hosts and their associated microbiota are assumed to have coevolved with each other [2]
According to the "separation" approach, the microorganisms can be divided into pathogens, neutral, and symbionts, depending on their interaction with their host. The coevolution between host and its associated microbiota may be accordingly described as antagonistic (based on negative interactions) or mutualistic (based on positive interactions).[2][89]
As of 2020, the emergence in publications about
Types
Marine
- Marine animal host-microbiome relationship
-
Relationships are generally thought to exist in a symbiotic state, and are normally exposed to environmental and animal-specific factors that may cause natural variations. Some events may change the relationship into a functioning but altered symbiotic state, whereas extreme stress events may cause dysbiosis or a breakdown of the relationship and interactions.[90]
All animals on Earth form associations with microorganisms, including protists, bacteria, archaea, fungi, and viruses. In the ocean, animal–microbial relationships were historically explored in single host–symbiont systems. However, new explorations into the diversity of microorganisms associating with diverse marine animal hosts is moving the field into studies that address interactions between the animal host and a more multi-member microbiome. The potential for microbiomes to influence the health, physiology, behavior, and ecology of marine animals could alter current understandings of how marine animals adapt to change, and especially the growing climate-related and anthropogenic-induced changes already impacting the ocean environment.[90]
The microbiomes of diverse marine animals are currently under study, from simplistic organisms including sponges[91] and ctenophores [92] to more complex organisms such as sea squirts[93] and sharks.[94][90]
The relationship between the
The gutless marine

Sponges are common members of the ocean's diverse benthic habitats and their abundance and ability to filter large volumes of seawater have led to the awareness that these organisms play critical roles in influencing benthic and pelagic processes in the ocean.[109] They are one of the oldest lineages of animals, and have a relatively simple body plan that commonly associates with bacteria, archaea, algal protists, fungi, and viruses.[110] Sponge microbiomes are composed of specialists and generalists, and complexity of their microbiome appears to be shaped by host phylogeny.[111] Studies have shown that the sponge microbiome contributes to nitrogen cycling in the oceans, especially through the oxidation of ammonia by archaea and bacteria.[112][113] Most recently, microbial symbionts of tropical sponges were shown to produce and store polyphosphate granules,[114] perhaps enabling the host to survive periods of phosphate depletion in oligotrophic marine environments.[115] The microbiomes of some sponge species do appear to change in community structure in response to changing environmental conditions, including temperature[116] and ocean acidification,[117][118] as well as synergistic impacts.[119]
-
Collecting a sample of blow from a blue whale using a helicopter drone [120]
-
Relative abundance of bacterial classes from whale blow, air and seawater samples.[121]
Terrestrial
Plant
-
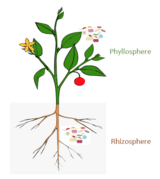
Microbiomes in the plant ecosystem [127]
The
Plant microbiomes are shaped by both factors related to the plant itself, such as genotype, organ, species and health status, as well as factors related to the plant's environment, such as management, land use and climate.[132] The health status of a plant has been reported in some studies to be reflected by or linked to its microbiome.[133][128][134][129]
Plant and plant-associated microbiota colonise different niches on and inside the plant tissue. All the above-ground plant parts together, called the phyllosphere, are a continuously evolving habitat due to ultraviolet (UV) radiation and altering climatic conditions. It is primarily composed of leaves. Below-ground plant parts, mainly roots, are generally influenced by soil properties. Harmful interactions affect the plant growth through pathogenic activities of some microbiota members. On the other hand, beneficial microbial interactions promote plant growth.[127]
Animal
-
Principal coordinate analysis of animal gut microbiome data [135]
The mammalian gut microbiome has emerged as a key regulator of host
The importance of phylogeny-correlated factors to the diversity of vertebrate microbiomes more generally is still poorly understood.
Without broader evolutionary context, it is unclear how universally conserved patterns of host-microbe phylosymbiosis actually are. Growing evidence indicates that the strong patterns identified in mammals are the exception rather than the rule in vertebrates.
Human
The
Humans are colonised by many microorganisms, with approximately the same order of magnitude of non-human cells as human cells.
The Human Microbiome Project (HMP) took on the project of sequencing the genome of the human microbiota, focusing particularly on the microbiota that normally inhabit the skin, mouth, nose, digestive tract, and vagina.[155] It reached a milestone in 2012 when it published its initial results.[160]
Assessment
Currently available methods for studying microbiomes, so-called
- Methods for assessing microbial functioning
-
Methods for assessing microbial functioningComplex microbiome studies cover various areas, starting from the level of complete microbial cells (single cell genomics, metabarcoding, metagenomics), RNA (metatranscriptomics), protein (metaproteomics), and metabolites (metabolomics). In that order, the focus of the studies shifts from the microbial potential (learning about available microbiota in the given habitat) over the metabolic potential (deciphering available genetic material) towards microbial functioning (e.g., the discovery of the active metabolic pathways).[2]
As of 2020, understanding remains limited due to missing links between the massive availability of microbiome
The number of prokaryotic phyla may reach hundreds, and archaeal ones are among the least studied.
Each microbiome system is suited to address different types of questions based on the culturability of microbes, genetic tractability of microbes and host (where relevant), ability to maintain system in laboratory setting, and ability to make host/environment germfree.[172]
- Underlying complexity
-
Tradeoffs between experimental questions and complexity of microbiome systemsgnotobiotic mice is crucial for making links between host diet and the effects on specific microbial taxa in a community.[172]
See also
- Earth Microbiome Project
- Human microbiome
- Initial acquisition of microbiota
- Microbial population biology
- Microbiomes of the built environment
- Mycobiome
References
- PMID 37258874, retrieved 24 November 2023
- ^ PMID 32605663..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - ^ Merchak A, Gaultier A. Microbial metabolites and immune regulation: New targets for major depressive disorder. Brain Behav Immun Health. 2020 Nov 2;9:100169. doi: 10.1016/j.bbih.2020.100169. PMID 34589904; PMCID: PMC8474524.
- ^ Hiltner L. (1902) "Die Keimungsverhältnisse der Leguminosensamen und ihre Beeinflussung durch Organismenwirkung". In: Parey P and Springer J (Eds.) Arb Biol Abt Land u Forstw K Gsndhtsamt, 3, Berlin. Pages 1–545.
- ^ Metchnikoff E. The prolongation of life: optimistic studies. GP Putnam's Sons; 1908.
- .
- .
- ^ PMID 270744.
- .
- .
- .
- .
- .
- .
- .
- .
- PMID 25750239.
- S2CID 88308313.
- PMID 16200144.
- .
- PMID 22882766.
- ^ Evans, A.S. (1976) "Causation and disease: the Henle-Koch postulates revisited. The Yale journal of biology and medicine, 49(2): 175.
- PMID 22092289.
- S2CID 4419735.
- The Nobel Foundation. Retrieved 28 September 2008.
- ^ Borman, S., Russell, H. and Siuzdak, G., (2003) "A Mass Spec Timeline Developing techniques to measure mass has been a Nobel pursuit. Todays Chemist at Work, 12(9): 47–50.
- PMID 13076173.
- PMID 20474956.
- ^ Hayes, W. (1966) "Genetic Transformation: a Retrospective Appreciation", First Griffith Memorial Lecture. Microbiology, 45(3): 385–397.
- ^ American Chemical Society (1999) Discovery and Development of Penicillin, 1928–1945. International Historic Chemical Landmarks, The Alexander Fleming Laboratory Museum, London.
- .
- PMID 33226.
- S2CID 4055566.
- S2CID 6089989.
- PMID 24747923.
- S2CID 4164029.
- OCLC 864917338.
- PMID 1105573.
- PMID 271968.
- S2CID 27846422.
- S2CID 8666687.
- S2CID 4289674.
- PMID 6801762.
- ^ Mullis, K.B. (1990) "The unusual origin of the polymerase chain reaction". Scientific American, 262(4): 56–65.
- S2CID 5714001.
- S2CID 17565963.
- ISSN 2590-0072.
- OCLC 22597587.
- S2CID 206934494.
- .
- OCLC 769756150.
- PMID 7535888.
- PMID 7542800.
- S2CID 86041893.
- PMID 22732228.
- PMID 20858600.
- S2CID 2144462.
- PMID 21304728.
- ^ "BioConcepts". www.biological-concepts.com. Archived from the original on 4 June 2023. Retrieved 18 December 2020.
- ^ "microbiome". Oxford English Dictionary (Online ed.). Oxford University Press. Retrieved 18 December 2020. (Subscription or participating institution membership required.)
- .
- ^ ISBN 9780719019791.
- ^ a b Lederberg, J. and McCray, A.T. (2001) "'Ome Sweet'Omics—A genealogical treasury of words". The Scientist, 15(7): 8.
- ^ .
- .
- .
- ^ a b Merriam-Webster Dictionary – microbiome.
- ^ Human Microbiome Project. Accessed 25 Aug 2020.
- ^ Nature.com: Microbiome. Accessed 25 August 2020.
- ^ ScienceDirect: Microbiome Accessed 25 August 2020.
- .
- .
- ISBN 9780127999227.
- .
- .
- ^ MicrobiomeSupport project
- ^ Carini, Paul (2016) A census of the dead: the story behind microbial 'relic DNA' in soil Archived 28 September 2021 at the Wayback Machine Nature Research: Microbiology.
- .
- ^ .
- ISBN 9789400724457.
- .
- S2CID 46895123.
- PMID 34365152.
- PMID 33499972..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - S2CID 39126203.
- PMID 28115400.
- PMID 28115400.
- S2CID 46895123.
- PMID 32873208.
- ^ doi:10.3389/fmars.2017.00222..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - .
- .
- .
- .
- .
- .
- doi:10.1038/35077067.
- ^ .
- .
- .
- .
- .
- ^ .
- ^ Dubinsky, Z. and Jokiel, P.L. (1994) "Ratio of energy and nutrient fluxes regulates symbiosis between zooxanthellae and corals". Pacific Science, 48(3): 313–324.
- .
- .
- .
- .
- .
- .
- .
- .
- .
- .
- .
- .
- .
- .
- .
- ^ S2CID 86518859.
- .
- PMID 31210263.
- PMID 23761254.
- PMID 31260051.
- PMID 32614926.
- PMID 29865228.
- ^ doi:10.3390/microorganisms7080269..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - ^ PMID 23805896.
- ^ PMID 30464766...
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - doi:10.1016/j.cpb.2020.100161..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - .
- PMID 26052316.
- S2CID 32900768.
- PMID 25278934.
- ^ PMID 31911491..
 Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Modified text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - PMID 23391737.
- ^ PMID 18497261.
- PMID 21596990.
- PMID 29995839.
- PMID 28230052.
- PMID 26393325.
- S2CID 9273396.
- S2CID 13850805.
- PMID 27861590.
- S2CID 8854653.
- PMID 30467310.
- PMID 29290090.
- S2CID 12320060.
- PMID 26042099.
- PMID 22486918.
- PMID 26191057.
- PMID 31097702.
- PMID 31132110.
- ^ PMID 26229597., which combined with the environment constitutes the microbiome.
Microbiome
This term refers to the entire habitat, including the microorganisms (bacteria, archaea, lower and higher eurkaryotes, and viruses), their genomes (i.e., genes), and the surrounding environmental conditions. This definition is based on that of "biome," the biotic and abiotic factors of given environments. Others in the field limit the definition of microbiome to the collection of genes and genomes of members of a microbiota. It is argued that this is the definition of metagenome - ^ OCLC 886600661.
- PMID 26824647.
- PMID 24729765.
- PMID 26274026.
we review literature on trimethylamine (TMA), a microbiota-generated metabolite linked to atherosclerosis development.
- PMID 25473158.
Trimethylamine is exclusively a microbiota-derived product of nutrients (lecithin, choline, TMAO, L-carnitine) from normal diet, from which seems originate two diseases, trimethylaminuria (or Fish-Odor Syndrome) and cardiovascular disease through the proatherogenic property of its oxidized liver-derived form.
- ^ "NIH Human Microbiome Project defines normal bacterial makeup of the body". NIH News. 13 June 2012.
- PMID 27774985.
- PMID 30661755.
- S2CID 198193092.
- PMID 22589385.
- ISSN 2452-3100.
- PMID 26109480.
- PMID 19528653.
- S2CID 36033561.
- ^ S2CID 52176496.
- PMID 28731467.
- PMID 30098831.
- ^ .

![Shift of paradigm from microbes as unsocial organisms causing diseases to a holistic view of microorganisms as the centre of the One Health Concept interconnecting all areas of human lives.[2]](http://upload.wikimedia.org/wikipedia/commons/3/33/Microbiome_paradigm_shifts.png)
![Co-occurrence networks help visualising microbial interactions Nodes usually represent taxa of microorganisms, and edges represent statistically significant associations between nodes.[2] ––––––––––––––––––––––––––– Testing of the hypotheses resulted from the network analyses is required for a comprehensive study of microbial interactions.[2]](http://upload.wikimedia.org/wikipedia/commons/thumb/2/21/Microbial_interactions_visualized_through_microbial_co-occurrence_networks.webp/615px-Microbial_interactions_visualized_through_microbial_co-occurrence_networks.webp.png)
![Co-occurrence networks show difference in gut microbiota between herbivorous and carnivorous cichlids Nodes coloured according to phylum. The herbivore network has higher complexity (156 nodes and 339 edges) compared to the carnivore network (21 nodes and 70 edges).[84]](http://upload.wikimedia.org/wikipedia/commons/thumb/d/d0/Co-occurrence_networks_showing_difference_in_gut_microbiota_between_herbivorous_and_carnivorous_cichlids.webp/562px-Co-occurrence_networks_showing_difference_in_gut_microbiota_between_herbivorous_and_carnivorous_cichlids.webp.png)

![Relationships are generally thought to exist in a symbiotic state, and are normally exposed to environmental and animal-specific factors that may cause natural variations. Some events may change the relationship into a functioning but altered symbiotic state, whereas extreme stress events may cause dysbiosis or a breakdown of the relationship and interactions.[90]](http://upload.wikimedia.org/wikipedia/commons/b/be/Marine_animal_host-microbiome_relationships.jpg)

![Relative abundance of bacterial classes from whale blow, air and seawater samples.[121]](http://upload.wikimedia.org/wikipedia/commons/thumb/b/b5/Cetacean_blow%27s_bacteria.png/426px-Cetacean_blow%27s_bacteria.png)


![Methods for assessing microbial functioning Complex microbiome studies cover various areas, starting from the level of complete microbial cells (microscopy, culturomics), followed by the DNA (single cell genomics, metabarcoding, metagenomics), RNA (metatranscriptomics), protein (metaproteomics), and metabolites (metabolomics). In that order, the focus of the studies shifts from the microbial potential (learning about available microbiota in the given habitat) over the metabolic potential (deciphering available genetic material) towards microbial functioning (e.g., the discovery of the active metabolic pathways).[2]](http://upload.wikimedia.org/wikipedia/commons/thumb/1/13/Methods_for_assessing_microbial_functioning.webp/972px-Methods_for_assessing_microbial_functioning.webp.png)
![Tradeoffs between experimental questions and complexity of microbiome systems [172] (A) Pairwise interactions between the soil bacteria Bacillus subtilis and Streptomyces spp. are well-suited for characterizing the functions of secondary metabolites in microbial interactions. (B) The symbiosis between bobtail squid and the marine bacterium Aliivibrio fischeri is fundamental to understanding host and microbial factors that influence colonization. (C) The use of gnotobiotic mice is crucial for making links between host diet and the effects on specific microbial taxa in a community.[172]](http://upload.wikimedia.org/wikipedia/commons/thumb/0/03/Tradeoffs_between_experimental_questions_and_complexity_of_microbiome_systems.jpg/1096px-Tradeoffs_between_experimental_questions_and_complexity_of_microbiome_systems.jpg)
