Systems biology
| Complex systems |
|---|
| Topics |

Systems biology is the
Particularly from the year 2000 onwards, the concept has been used widely in biology in a variety of contexts. The
Overview
This section is written like a personal reflection, personal essay, or argumentative essay that states a Wikipedia editor's personal feelings or presents an original argument about a topic. (December 2022) |
Systems biology can be considered from a number of different aspects.
As a field of study, particularly, the study of the interactions between the components of biological systems, and how these interactions give rise to the function and behavior of that system (for example, the
As a
As a series of operational
As the application of dynamical systems theory to molecular biology. Indeed, the focus on the dynamics of the studied systems is the main conceptual difference between systems biology and bioinformatics.[11]
As a socioscientific phenomenon defined by the strategy of pursuing integration of complex data about the interactions in biological systems from diverse experimental sources using interdisciplinary tools and personnel.[12]
History
Although the concept of a systems view of cellular function has been well understood since at least the 1930s,[13] technological limitations made it difficult to make systems wide measurements. The advent of microarray technology in the 1990s opened up an entire new visa for studying cells at the systems level. In 2000, the Institute for Systems Biology was established in Seattle in an effort to lure "computational" type people who it was felt were not attracted to the academic settings of the university. The institute did not have a clear definition of what the field actually was: roughly bringing together people from diverse fields to use computers to holistically study biology in new ways.[14] A Department of Systems Biology at Harvard Medical School was launched in 2003.[15] In 2006 it was predicted that the buzz generated by the "very fashionable" new concept would cause all the major universities to need a systems biology department, thus that there would be careers available for graduates with a modicum of ability in computer programming and biology.[14] In 2006 the National Science Foundation put forward a challenge to build a mathematical model of the whole cell.[citation needed] In 2012 the first whole-cell model of Mycoplasma genitalium was achieved by the Karr Laboratory at the Mount Sinai School of Medicine in New York. The whole-cell model is able to predict viability of M. genitalium cells in response to genetic mutations.[16]
An earlier precursor of systems biology, as a distinct discipline, may have been by systems theorist
According to
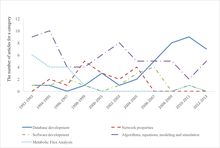
An important milestone in the development of systems biology has become the international project Physiome.[citation needed]
Associated disciplines
According to the interpretation of systems biology as using large data sets using interdisciplinary tools, a typical application is
Items that may be a computer database include:
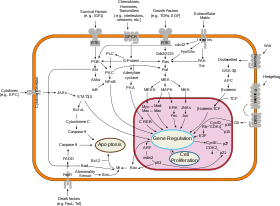
The molecular interactions within the cell are also studied, this is called
In approaching a systems biology problem there are two main approaches. These are the top down and bottom up approach. The top down approach takes as much of the system into account as possible and relies largely on experimental results. The RNA-Seq technique is an example of an experimental top down approach. Conversely, the bottom up approach is used to create detailed models while also incorporating experimental data. An example of the bottom up approach is the use of circuit models to describe a simple gene network.[31]
Various technologies utilized to capture dynamic changes in mRNA, proteins, and post-translational modifications. Mechanobiology, forces and physical properties at all scales, their interplay with other regulatory mechanisms;[32] biosemiotics, analysis of the system of sign relations of an organism or other biosystems; Physiomics, a systematic study of physiome in biology.
The systems biology approach often involves the development of
Bioinformatics and data analysis
Other aspects of computer science,
Creating biological models
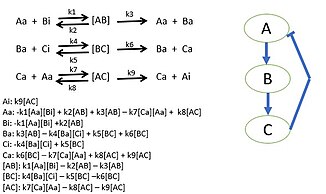
Researchers begin by choosing a biological pathway and diagramming all of the protein, gene, and/or metabolic pathways. After determining all of the interactions,
The use of constraint-based reconstruction and analysis (COBRA) methods has become popular among systems biologists to simulate and predict the metabolic phenotypes, using genome-scale models. One of the methods is the flux balance analysis (FBA) approach, by which one can study the biochemical networks and analyze the flow of metabolites through a particular metabolic network, by optimizing the objective function of interest (e.g. maximizing biomass production to predict growth).[46]

See also
- Biochemical systems equation
- Biological computation
- BioSystems (journal)
- Computational biology
- Exposome
- Interactome
- List of omics topics in biology
- List of systems biology modeling software
- Living systems
- Metabolic Control Analysis
- Metabolic network modelling
- Modelling biological systems
- Molecular pathological epidemiology
- Network biology
- Network medicine
- Synthetic biology
- Systems biomedicine
- Systems immunology
- Systems medicine
- TIARA (database)
References
- ^ S2CID 52922135.
- ^ Zewail, Ahmed (2008). Physical Biology: From Atoms to Medicine. Imperial College Press. p. 339.
- S2CID 27653540.
- PMID 21570668.
- ISBN 978-3-540-22968-1.
- ^ "Systems Biology: the 21st Century Science". Institute for Systems Biology. Retrieved 15 June 2011.
- ^ ISBN 978-0-19-929573-9.
- S2CID 42448991.
- ISBN 978-3-540-22968-1.
- ISBN 978-1-59745-440-7.
- ISBN 9780815344674.
- PMID 19623486.
- ^ Wright, Sewall (1934). "Physiological and Evolutionary Theories of Dominance". The American Naturalist. pp. 24–53.
- ^ a b c Kling, Jim (3 March 2006). "Working the Systems". Science. Retrieved 15 June 2011.
- ^ "HMS launches new department to study systems biology". Harvard Gazette. September 23, 2003.
- PMID 22817898.
- Mesarovic, Mihajlo D.(1968). Systems Theory and Biology. Berlin: Springer-Verlag.
- ^ JSTOR 1724368.
- PMID 4148886.
- PMID 7672373.
- PMID 4830198.
- )
- )
- )
- ^ "Systems Biology - National Institute of General Medical Sciences". Archived from the original on 19 October 2013. Retrieved 12 December 2012.
- PMID 22491028.
- PMID 30044828.
- ^ PMID 18793124.
- PMID 16162640.
- S2CID 12355069.
- PMID 23372424.
- PMID 30105089.
- ISBN 978-1439831441.
- ^
S2CID 24616792.
- S2CID 8356492.
- S2CID 16270018.
- ^ S2CID 89307028.
- PMID 26284497.
- ^ PMID 27429455.
- PMID 17045684.
- PMID 22123829.
- PMID 28855977.
- ^ PMID 27187545.
- S2CID 12122032.
- PMID 14525003.
- PMID 20212490.
Further reading
- Klipp, Edda; Liebermeister, Wolfram; Wierling, Christoph; Kowald, Axel (2016). Systems Biology - A Textbook, 2nd edition. Wiley. ISBN 978-3-527-33636-4.
- Asfar S. Azmi, ed. (2012). Systems Biology in Cancer Research and Drug Discovery. Springer. ISBN 978-94-007-4819-4.
- ISBN 978-0-262-11266-6.
- Werner, Eric (29 March 2007). "All systems go". Nature. 446 (7135): 493–494. doi:10.1038/446493a. provides a comparative review of three books:
- ISBN 978-1-58488-642-6.
- Kaneko, Kunihiko (15 September 2006). Life: An Introduction to Complex Systems Biology. Springer-Verlag. ISBN 978-3-540-32666-3.
- Palsson, Bernhard O. (16 January 2006). Systems Biology: Properties of Reconstructed Networks. Cambridge University Press. ISBN 978-0-521-85903-5.
- Werner Dubitzky; Olaf Wolkenhauer; Hiroki Yokota; Kwan-Hyun Cho, eds. (13 August 2013). Encyclopedia of Systems Biology. Springer-Verlag. ISBN 978-1-4419-9864-4.
External links
 Media related to Systems biology at Wikimedia Commons
Media related to Systems biology at Wikimedia Commons- Biological Systems in bio-physics-wiki

