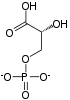
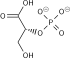
User:UndercoverGeologicalAgent/3-Phosphoglyceric acid

| |
| Names | |
|---|---|
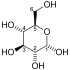
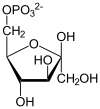
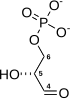
| IUPAC name
(2R)-2-Hydroxy-3-phosphonooxypropanoic acid
| |
| Identifiers | |
3D model (
JSmol ) |
|
| 3DMet | |
| ChEBI | |
| ChEMBL | |
| ChemSpider | |
| DrugBank | |
| KEGG | |
PubChem CID
|
|
| |
| |
| Properties | |
| C3H7O7P | |
| Molar mass | 186.06 g/mol |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |
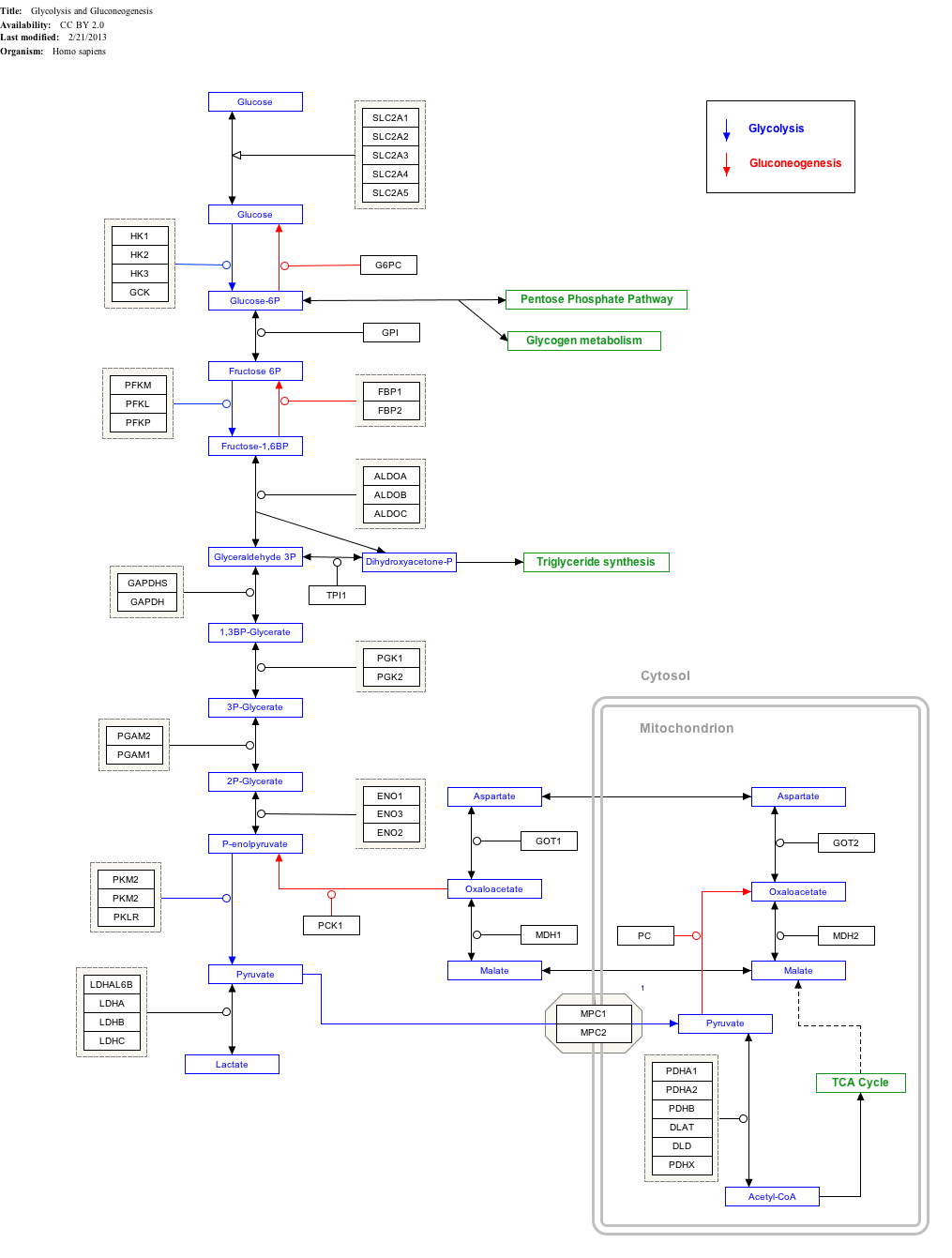
3-Phosphoglyceric acid (3PG, 3-PGA, or PGA) is the conjugate acid of 3-phosphoglycerate or glycerate 3-phosphate (GP or G3P).
Glycolysis
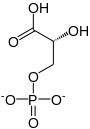
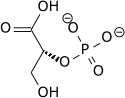
In the glycolytic pathway, 1,3-bisphosphoglycerate is dephosphorylated to form 3-phosphoglyceric acid in a coupled reaction producing two ATP via substrate-level phosphorylation.[5] The single phosphate group left on the 3-PGA molecule then moves from an end carbon to a central carbon, producing 2-phosphoglycerate.[5][a] This phosphate group relocation is catalyzed by phosphoglycerate mutase, an enzyme that also catalyzes the reverse reaction.[6]
1,3-bisphospho-D-glycerate
|
3-phosphoglycerate kinase
|
3-phospho-D-glycerate
|
Phosphoglyceromutase
|
2-phospho-D-glycerate
| ||

|
|
| ||||
| ADP | ATP | |||||

|

| |||||
| ADP | ATP | |||||
3-phosphoglycerate kinase
|
Phosphoglyceromutase
|
|||||
Compound C00236 at KEGG Pathway Database. Enzyme 2.7.2.3 at KEGG Pathway Database. Compound C00197 at KEGG Pathway Database. Enzyme 5.4.2.1 at KEGG Pathway Database. Compound C00631 at KEGG Pathway Database.
Click on genes, proteins and metabolites below to link to respective articles.[§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "GlycolysisGluconeogenesis_WP534".
Calvin-Benson cycle
In the
Amino acid synthesis
Glycerate 3-phosphate (formed from 3-phosphoglycerate) is also a precursor for serine, which, in turn, can create cysteine and glycine through the homocysteine cycle.[11][12][13]
Measurement
3-phosphoglycerate can be separated and measured using paper chromatography[14] as well as with column chromatography and other chromatographic separation methods.[15] It can be identified using both gas-chromatography and liquid-chromatography mass spectrometry and has been optimized for evaluation using tandem MS techniques.[1][16][17]
See also
References
- ^ a b "3-Phosphoglyceric acid (HMDB0000807)". Human Metabolome Database. The Metabolomics Innovation Centre. Retrieved 23 May 2021.
{{cite web}}: CS1 maint: url-status (link) - ISBN 0-7167-3051-0.
- ISBN 1-57259-153-6.
- ^ ISBN 978-0-7923-6143-5.
- ^ a b Rye, Connie; Wise, Robert; Jurukovski, Vladimir; DeSaix, Jean; Choi, Jung; Avissar, Yael (2016). "Glycolysis". Biology. OpenStax College.
- PMID 8447.
- PMID 18417482.
- ^ a b Moran, L. (2007). "The Calvin Cycle: Regeneration". Sandwalk. Retrieved 11 May 2021.
{{cite web}}: CS1 maint: url-status (link) - ^ PMID 3137030.
- PMID 10482420.
- PMID 29593770.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 16589713.
- doi:10.1038/182532a0.
- PMID 13385236.
- PMID 4278264.
- PMID 25790351.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - .
- ^ Note that 3-phosphoglycerate and 2-phosphoglycerate are isomers of one another