Genetic history of the Iberian Peninsula

The ancestry of modern Iberians (comprising the
Modern Iberians' genetic inheritance largely derives from the pre-Roman inhabitants of the Iberian Peninsula:
There are also minor genetic influences from the
Population Genetics: Methods and Limitations
The foremost pioneer of the study of population genetics was Luigi Luca Cavalli-Sforza. Cavalli-Sforza used classical genetic markers to analyse DNA by proxy. This method studies differences in the frequencies of particular allelic traits, namely polymorphisms from proteins found within human blood (such as the ABO blood groups, Rhesus blood antigens, HLA loci, immunoglobulins, G-6-P-D isoenzymes, among others). Subsequently, his team calculated genetic distances between populations, based on the principle that two populations that share similar frequencies of a trait are more closely related than populations that have more divergent frequencies of the trait.[20]
Since then, population genetics has progressed significantly and studies using direct DNA analysis are now abundant and may use mitochondrial DNA (mtDNA), the non-recombining portion of the Y chromosome (NRY) or autosomal DNA. MtDNA and NRY DNA share some similar features which have made them particularly useful in genetic anthropology. These properties include the direct, unaltered inheritance of mtDNA and NRY DNA from mother to offspring and father to son, respectively, without the 'scrambling' effects of genetic recombination. We also presume that these genetic loci are not affected by natural selection and that the major process responsible for changes in base pairs has been mutation (which can be calculated).[21]
Whereas Y-DNA and mtDNA haplogroups represent but a small component of a person's DNA pool, autosomal DNA has the advantage of containing hundreds and thousands of examinable genetic loci, thus giving a more complete picture of genetic composition. Descent relationships can only to be determined on a statistical basis, because autosomal DNA undergoes recombination. A single chromosome can record a history for each gene. Autosomal studies are much more reliable for showing the relationships between existing populations but do not offer the possibilities for unraveling their histories in the same way as mtDNA and NRY DNA studies promise, despite their many complications.[citation needed]
Analyses of nuclear and ancient DNA

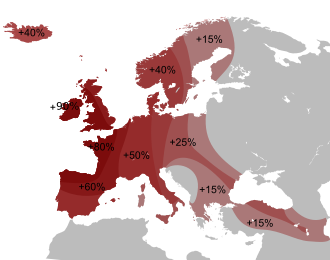
Nuclear DNA analysis shows that Spanish and Portuguese populations are most closely related to other populations of western Europe.[22][23][24] There is an axis of significant genetic differentiation along the east–west direction, in contrast to remarkable genetic similarity in the north–south direction. North African admixture, associated with the
A study published in 2019 using samples of 271 iberians spanning prehistoric and historic times proposes the following inflexion points in Iberian genomic history:[26]
- Mesolithic: hunter-gatherers from the European Steppes of Western Russia, Georgia and Ukraine are the first humans to settle the northwest of the Iberian Peninsula.
- neolithic farmers settle the entire Iberian Peninsula from Anatolia.
- Chalcolithic: Inflow of Central European hunter-gatherers and some gene inflow from sporadic contact with North Africa.
- Bronze Age: Steppe inflow from Central Europe.
- Basque peopleremains mostly intact from this point on.
- Mediterranean. Some additional inflow of North African genes detected in Southern Iberia.
- Visigothic period: no detectable inflows.
- Muslim period: Inflow from Northern Africa. Following the Reconquista, there is further genetic convergence between North and South Iberia.
North African influence

A number of studies have focused on ascertaining the genetic impact of historical North African population movements into Iberia on the genetic composition of modern Spanish and Portuguese populations. Initial studies pointed to the
In terms of autosomal DNA, the most recent study regarding African admixture in Iberian populations was conducted in April 2013 by Botigué et al. using genome-wide SNP data for over 2000 European, Maghreb, Qatar and Sub-Saharan individuals of which 119 were Spaniards and 117 Portuguese, concluding that Spain and Portugal hold significant levels of North African ancestry. Estimates of shared ancestry averaged from 4% in some places to 10% in the general population; the populations of the Canary Islands yielded from 0% to 96% of shared ancestry with north Africans, although the Canary islands are a Spanish exclave located in the African continent, and thus this output is not representative of the Iberian population; these same results did not exceed 2% in other western or southern European populations.[37][38][39][40] However, contrary to past autosomal studies and to what is inferred from Y-Chromosome and Mitochondrial Haplotype frequencies (see below), it does not detect significant levels of Sub-Saharan ancestry in any European population outside the Canary Islands. Indeed, a prior 2011 autosomal study by Moorjani et al. found Sub-Saharan ancestry in many parts of southern Europe at ranges of between 1-3%, "the highest proportion of African ancestry in Europe is in Iberia (Portugal 4.2±0.3% and Spain 1.4±0.3%), consistent with inferences based on mitochondrial DNA and Y chromosomes and the observation by Auton et al. that within Europe, the Southwestern Europeans have the highest haplotype-sharing with North Africans."[30][34][35]
Recent studies show minor relationships between some Iberian regions and North African populations as a result of the Al-Andalus historical period which in Portugal lasted between the 8th and 12th centuries AD, and in southern Spain continued until the late 15th century AD. Iberia is the European region that has a more prominent presence of haplogroup E3b of the human Y chromosome (E-M81),[41] of haplogroup U (U6) and Haplotype Va, and this may be the result of some original common western Mediterranean population. In Portugal, North African Y-chromosome haplogroups (especially those typically North-West African) are at a frequency of 7.1%.[42] Some studies of mitochondrial DNA also find evidence of the North African haplogroup U6, especially in northern Portugal.[43] Although the frequency of U6 is low (4–6%), it was estimated that approximately 27% of the population of northern Portugal had some North African ancestry, as U6 is also not a common lineage in North Africa.[44] According to some studies, the North African and Arab elements in the ancestry of today's Iberians are more than trivial when compared to the basis of pre-Islamic ancestry, and the Strait of Gibraltar seems to function more as a genetic bridge than a barrier.[45][46][47]
However, a study that has used different genetic markers has reached different conclusions. In an autosomal study by Spínola et al. (2005), which analyzed the

According to a study published in the American Journal of Human Genetics in December 2008, 30% of modern Portuguese (23.6% in the north and 36.3% in the south) have DNA that shows they have male Sephardic Jewish ancestry and 14% (11.8 in the North and 16.1% in the South) have Moorish ancestry.[49] Despite the possible alternative sources for lineages attributed to a Sephardic Jewish origin, these proportions were testimony to the importance of religious conversion (voluntary or forced), shown by historical episodes of social and religious intolerance.

In terms of paternal Y-Chromosome DNA, recent studies coincide in that Iberia has the greatest presence of the typically
Other studies of the Iberian gene-pool have estimated significantly lower levels of North African Ancestry. According to Bosch et al. 2000 "NW African populations may have contributed 7% of Iberian Y chromosomes".[28] A wide-ranging study by Cruciani et al. 2007, using 6,501 unrelated Y-chromosome samples from 81 populations found that: "Considering both these E-M78 sub-haplogroups (E-V12, E-V22, E-V65) and the E-M81 haplogroup, the contribution of northern African lineages to the entire male gene pool of Iberia (barring Pasiegos), continental Italy and Sicily can be estimated as 5.6 percent, 3.6 percent and 6.6 percent, respectively".[52] A 2007 study estimated the contribution of northern African lineages to the entire male gene pool of Iberia as 5.6%."[53] In general aspects, according to (Bosch et al. 2007) "...the origins of the Iberian Y-chromosome pool may be summarized as follows: 5% recent NW African, 78% Upper Paleolithic and later local derivatives (group IX), and 10% Neolithic" (H58, H71).[54]
Mitochondrial DNA studies of 2003, coincide in that the Iberian Peninsula holds higher levels of typically North African Haplotype U6,
The most recent and comprehensive genomic studies establish that
Current debates revolve around whether U6 presence is due to Islamic expansion into the Iberian peninsula or prior population movements[34][35][36] and whether Haplogroup L is linked to the slave trade or prior population movements linked to Islamic expansion. A majority of Haplogroup L lineages in Iberia being North African in origin points to the latter.[56][58][30][35][63] In 2015, Hernández et al. concluded that "the estimated entrance of the North African U6 lineages into Iberia at 10 ky correlates well with other L African clades, indicating that some U6 and L lineages moved together from Africa to Iberia in the Early Holocene while a majority were introduced during historic times."[64]
Haplogroups

Y-Chromosome haplogroups
Like other Western Europeans, among Spaniards and Portuguese the Y-DNA Haplogroup R1b is the most frequent, occurring at over 70% throughout most of Spain.[65] R1b is particularly dominant in the Basque Country and Catalonia, occurring at rate of over 80%. In Iberia, most men with R1b belong to the subclade R-P312 (R1b1a1a2a1a2; as of 2017). The distribution of haplogroups other than R1b varies widely from one region to another.
In Portugal as a whole the R1b haplogroups rate 70%, with some areas in the Northwest regions reaching over 90%.[66]
Although R1b prevails in much of Western Europe, a key difference is found in the prevalence in Iberia of R-DF27 (R1b1a1a2a1a2a). This subclade is found in over 60% of the male population in the Basque Country and 40-48% in Madrid, Alicante, Barcelona, Cantabria, Andalucia, Asturias and Galicia. Sephardic Jews
Mitochondrial DNA

There have been a number of studies about the
The subhaplogroups H1 and H3 have been subject to a more detailed study and would be associated to the Magdalenian expansion from Iberia c. 13,000 years ago:[56]
- H1 encompasses an important fraction of Western European mtDNA, reaching its local peak among contemporary Basques (27.8%) and appearing at a high frequency among other Iberians and North Africans. Its frequency is above 10% in many other parts of Europe (France, Sardinia, British Isles, Alps, large portions of Eastern Europe), and above 5% in nearly all the continent. Its subclade H1b is most common in eastern Europe and NW Siberia.[74] So far, the highest frequency of H1 - 61%- has been found among the Tuareg of the Fezzan region in Libya.[75]
- H3 represents a smaller fraction of European genome than H1 but has a somewhat similar distribution with peak among Basques (13.9%), Sardinians (8.5%). Its frequency decreases towards the northeast of the continent, though. Studies have suggested haplogroup H3 is highly protective against AIDS progression.[76]
A 2007 European-wide study including Spanish Basques and Valencian Spaniards found Iberian populations to cluster the furthest from other continental groups, implying that Iberia holds the most ancient European ancestry. In this study, the most prominent genetic stratification in Europe was found to run from the north to the south-east, while another important axis of differentiation runs east–west across the continent. It also found, despite the differences, that all Europeans are closely related.[77]
Subregions
Spain
| Region | Sample size | C | E | G | I | J2
|
JxJ2 | R1a
|
R1b
|
Notes |
|---|---|---|---|---|---|---|---|---|---|---|
| Aragon | 34 | 6% | 0% | 18% | 12% | 0% | 3% | 56% | ||
| Andalusia East | 95 | 4% | 3% | 6% | 9% | 3% | 1% | 72% | ||
| Andalusia West | 73 | 15% | 4% | 5% | 14% | 1% | 4% | 54% | ||
| Asturias | 20 | 15% | 5% | 10% | 15% | 0% | 0% | 50% | ||
| Basques | 116 | 1% | 0% | 8% | 3% | 1% | 0% | 87% | ||
Castilla La Mancha
|
63 | 4% | 10% | 2% | 6% | 2% | 2% | 72% | ||
| Castile North-East | 31 | 9% | 3% | 3% | 3% | 0% | 0% | 77% | ||
| Castile North-West | 100 | 19% | 5% | 3% | 8% | 1% | 2% | 60% | ||
| Catalonia | 80 | >0%[79] | 3% | 6% | 3% | 6% | 0% | 0% | 81% | |
| Extremadura | 52 | 18% | 4% | 10% | 12% | 0% | 0% | 50% |
||
| Galicia | 88 | 17% | 6% | 10% | 7% | 1% | 0% | 57% |
||
| Valencia | 73 | >0%[80] | 10% | 1% | 10% | 5% | 3% | 3% | 64% |
|
Majorca
|
62 | 9% | 6% | 8% | 8% | 2% | 0% | 66% |
||
| Menorca | 37 | 19% | 0% | 3% | 3% | 0% | 3% | 73% |
||
| Ibiza | 54 | 8% | 13% | 2% | 4% | 0% | 0% | 57% |
||
| Seville | 155 | 7% | 4% | 12% | 8% | 3% | 1% | 60% |
||
| Huelva | 22 | 14% | 0% | 9% | 14% | 0% | 0% | 59% |
||
Cadiz
|
28 | 4% | 0% | 14% | 14% | 4% | 0% | 51% |
||
Cordoba
|
27 | 11% | 0% | 15% | 15% | 0% | 0% | 56% |
||
| Málaga | 26 | 31% | 4% | 0% | 15% | 0% | 8% | 43% |
||
| Leon | 60 | 10% | 7% | 3% | 5% | 2% | 7% | 62% |
||
| Cantabria | 70 | 13% | 9% | 6% | 3% | 3% | 4% | 58% |
Portugal
Excerpts from the Abstract of a study published[81] in 2015:
"[...] In the case of Portugal, previous population genetics studies have already revealed the general portrait of HVS-I and HVS-II mitochondrial diversity, becoming now important to update and expand the mitochondrial region analysed. Accordingly, a total of 292 complete control region sequences from continental Portugal were obtained, under a stringent experimental design to ensure the quality of data through double sequencing of each target region.* Furthermore, H-specific coding region SNPs were examined to detail haplogroup classification and complete mitogenomes were obtained for all sequences belonging to haplogroups U4 and U5. In general, a typical
Western European haplogroup or Atlantic modal haplotype(AMH) composition was found in mainland Portugal, associated to high level of mitochondrial genetic diversity. Within the country, no signs of substructure were detected. The typing of extra coding region SNPs has provided the refinement or confirmation of the previous classification obtained with EMMA tool in 96% of the cases. Finally, it was also possible to enlarge haplogroup U phylogeny with 28 new U4 and U5 mitogenomes."
The AMH reaches the highest frequencies in the Iberian Peninsula, in Great Britain and Ireland. In the Iberian Peninsula it reaches 70% in Portugal as a whole, with more than 90% in NW Portugal and nearly 90% in Galicia (NW Spain), while the highest value is to be found among the Basques (NE Spain).
The
The AMH is the most frequently occurring haplotype amongst human males in Atlantic Europe. It is characterized by the following marker alleles:
- DYS388 12
- DYS390 24
- DYS391 11
- DYS392 13
- DYS393 13
- DYS394 14 (also known as DYS19)
See also
- Genetic history of Europe
- Genetic history of North Africa
- Moroccan genetics
- Genetic studies on Jews
- Genetic studies on Arabs
References
- PMID 31127131.
- PMID 19424496.
- PMID 18758442.
- ^ "Iberians - MSN Encarta". Encarta.msn.com. Archived from the original on 30 October 2009. Retrieved 12 January 2022.
- ^ Álvarez-Sanchís, Jesús (28 February 2005). "Oppida and Celtic society in western Spain". E-Keltoi: Journal of Interdisciplinary Celtic Studies. 6 (1).
- ^ a b "Ethnographic Map of Pre-Roman Iberia (Circa 200 B.C.)". Arqueotavira.com. Archived from the original on 11 June 2004. Retrieved 12 January 2022.
- ^ "Spain - History". Britannica.com.
- ^ PMID 30710075.
- ^ PMID 30872528.
- ISBN 9788490960981.
- ^ https://alpha.sib.uc.pt/?q=content/o-património-visigodo-da-l%C3%ADngua-portuguesa [dead link]
- ^ Quiroga, Jorge López (January 2017). "(PDF) IN TEMPORE SUEBORUM. The time of the Suevi in Gallaecia (411–585 AD)". Jorge López Quiroga-Artemio M. Martínez Tejera (Coord.): In Tempore Sueborum. The Time of the Sueves in Gallaecia (411–585 Ad). The First Medieval Kingdom of the West, Ourense. Academia.edu. Retrieved 21 January 2020.
- ^ James S. Amelang. "The Expulsion of the Moriscos: Still more Questions than Answers" (PDF). Intransitduke.org. Universidad Autónoma, Madrid. Retrieved 22 January 2022.
- ^ Jónsson 2007, p. 195.
- PMID 19061982.
- ^ Torres, Gabriela (31 December 2008). "El español "puro" tiene de todo". BBC Mundo.
- PMID 19061982.
- ^ PMID 30710075.
- ^ PMID 30872528.
- ISBN 978-0-691-08750-4. Retrieved 2009-07-22.
- ISBN 978-0-306-46793-6.
- PMID 19424496.
- ^ Wade, Nicholas (13 August 2008). "The Genetic Map of Europe". The New York Times. Retrieved 17 October 2009.
- PMID 18758442.
- "Genetic map of Europe again". Gene Expression. 31 August 2008.
- PMID 30710075.
- PMID 30872528.
- S2CID 17665038.
- ^ PMID 11254456.
- S2CID 9618737.
- ^ PMID 21533020.
- ^ PMID 19156170.
- ^ PMID 15069642.
- ^ S2CID 13347549.
- ^ PMID 19061982.
- Tina Hesman Saey (3 January 2009). "Spanish Inquisition Couldn'T Quash Moorish, Jewish Genes". Science News. Vol. 175, no. 1. Archived from the original on 2009-02-21.
- ^ S2CID 6430969.
- ^ S2CID 18482536.
- PMID 23733930.
- ^ Estimating gene flow from North Africa to southern Europe Archived 2015-04-30 at archive.today, David Comas, one of the authors of the study
- ^ "La cifra del 20% sólo se da en Canarias, para el resto del país oscila entre el 10% y 12%", explica Comas.", David Comas, one of the authors of the study, Los españoles somos los europeos con más genes magrebíes, Huffington post, June 2013
- PMID 21479138.
- S2CID 13347549.
- S2CID 3229760.
- S2CID 20901589.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - )
- )
- )
- S2CID 9618737.
- PMID 15982254.
- ^ "The Genetic Legacy of Religious Diversity and Intolerance: Paternal Lineages of Christians, Jews, and Muslims in the Iberian Peninsula".
- PMID 31816048.
- ^ Lucotte, Gérard; Gérard, Nathalie; Mercier, Géraldine (2011-04-05). "North African Genes in Iberia Studied by Y-Chromosome DNA Haplotype 5". Human Biology. 73 (5).
- PMID 17351267.
- PMID 17351267.
- PMID 11254456.
- S2CID 11201992.
- ^ S2CID 20901589.
- PMID 22454235.
- ^ S2CID 8870699.
- ^ PMID 24460736.
- ISBN 9780190622183.
- ^ "Tracing Past Human Male Movements in Northern/Eastern Africa and Western Eurasia". Academic.oup.com. Retrieved 2020-01-21.
- ^ https://reich.hms.harvard.edu/sites/reich.hms.harvard.edu/files/inline-files/2019_Olalde_Science_IberiaTransect_2.pdf [bare URL PDF]
- PMID 20127843.
- PMID 26509580.
- ^ PMID 20736979.
- PMID 25457629.
- PMID 26081640.
- PMID 29466337.
- S2CID 4399103.
- S2CID 13670282.
- ^ "Eupedia".
- PMID 27441366.
- ^ Rosser et al. (2000)
- PMID 15254257.
- PMID 20975840.
- PMID 19005266.
- PMID 17436249.
- PMID 15280900.
- ^ Haplogroup C* (C-M130) has been found among males with the surname Llach and originating from Garrotxa, Catalonia, Spain. It was not found among males with the same surname from other areas, or males with other surnames of Catalan origin (Cognoms Catalans, n.d., Resultat; access 15 September 2015). The Cognoms Catalans project, which researches "genetic surnames" in Catalonia, Valencia and the Balearic Islands, is based at Universitat Pompeu Fabra, Barcelona.
- ^ C Haplogroup – Y-DNA Classic Chart (21 January 2017).
- PMID 25457629.
Works cited
- Jónsson, Már (2007). "The expulsion of the Moriscos from Spain in 1609–1614: the destruction of an Islamic periphery". S2CID 154793596.
- Rosser, Z; Zerjal, T; Hurles, M; Adojaan, M; Alavantic, D; Amorim, A; Amos, W; Armenteros, M; et al. (2000), "Y-Chromosomal Diversity in Europe Is Clinal and Influenced Primarily by Geography, Rather than by Language", PMID 11078479, archived from the originalon 2008-05-06
