User:Nhattrongdo/sandbox
| Dihydrofolate reductase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Dihydrofolate reductase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | DHFR_1 | ||||||||
SCOP2 | 1dhi / SCOPe / SUPFAM | ||||||||
| |||||||||
| R67 dihydrofolate reductase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
An Error has occurred retrieving Wikidata item for infobox Dihydrofolate reductase, or DHFR, is an
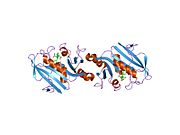
Structure
A central eight-stranded
Function
Dihydrofolate reductase converts
Found in all organisms, DHFR has a critical role in regulating the amount of tetrahydrofolate in the cell. Tetrahydrofolate and its derivatives are essential for
Mechanism

Since DHFR serves as an important model for mechanistic studies in enzymology, its catalytic mechanisms with different substrates have been investigated for a while using a wide range of methods including X-ray, NMR structures, molecular dynamics simulation, enzyme kinetic measurements, Raman spectroscopy analysis, and ensemble and single-molecule kinetics.[13][14][15][16][17] [18]
DHFR catalyzes the transfer of a hydride from
The catalytic cycle of the reaction catalyzed by DHFR incorporates five important intermediate: holoenzyme (E:NADPH), Michaelis complex (E:NADPH:DHF), ternary product complex (E:NADP+:THF), tetrahydrofolate binary complex (E:THF), and THF‚NADPH complex (E:NADPH:THF). The kinetic studies showed that the product (THF) dissociation step from E:NADPH:THF to E:NADPH is the rate determining step during steady-state turnover.[19]
Conformational changes are critical in DHFR's catalytic mechanism.[13] The Met20 loop of DHFR is able to open, close or occlude the active site.[20][21] Correspondingly, three different conformations classified as the opened, closed and occluded states are assigned to Met20. In addition, an extra distorted conformation of Met20 was defined due to its indistinct characterization results.[21] The Met20 loop is observed in its occluded conformation in the three product ligating intermediates, where the nicotinamide ring is occluded from the active site. This conformational feature accounts for the fact that the substitution of NADP+ by NADPH is prior to product dissociation. Thus, the next round of reaction can occur upon the binding of substrate.[19]

Studies on the kinetic mechanism of dehydrofolate reductase from Mycobacterium tuberculosis and Escherichia coli indicate that the mechanism of this enzyme is stepwise and steady-state random. Specifically, the catalytic reaction begins with the NADPH and the substrate attaching to the binding site of the enzyme, followed by the protonation and the hydride transfer from the cofactor NADPH to the substrate. However, two latter steps do not take place simultaneously in a same transition state.[20][22] In a study using computational and experimental approaches, Liu et al conclude that the protonation step precedes the hydride transfer.[23]
DHFR's enzymatic mechanism is shown to be pH dependent, particularly the hydride transfer step, since pH changes are shown to have remarkable influence on the electrostatics of the active site and the ionization state of its residues.[23] The acidity of the targeted nitrogen on the substrate is important in the binding of the substrate to the enzyme's binding site which is proved to be hydrophobic even though it has direct contact to water.[20][24] Asp27 is the only charged hydrophilic residue in the binding site, and neutralization of the charge on Asp27 may alter the pKa of the enzyme. Mutagenesis studies on important side chains show that Asp27 may play a critical role in the catalytic mechanism by helping with protonation of the substrate and restraining the substrate in the conformation favorable for the hydride transfer.[19][24] [20] The protonation step is shown to be associated with enol tautomerization even though this conversion is not considered favorable for the proton donation.[22] In some other studies, a water molecule is proved to be involved in the protonation step[21][17][25]. Analysis of DHFR crystal structures as well as simulation studies have proved that the entry of the water molecule to the active site of the enzyme is facilitated by the Met20 loop.[15]
Clinical significance
Dihydrofolate reductase deficiency has been linked to megaloblastic anemia.[9] Treatment is with reduced forms of folic acid. Because tetrahydrofolate, the product of this reaction, is the active form of folate in humans, inhibition of DHFR can cause functional folate deficiency. DHFR is an attractive pharmaceutical target for inhibition due to its pivotal role in DNA precursor synthesis. Trimethoprim, an antibiotic, inhibits bacterial DHFR while methotrexate, a chemotherapy agent, inhibits mammalian DHFR. However, resistance has developed against some drugs, as a result of mutational changes in DHFR itself.[26]
DHFR mutations cause a rare autosomal recessive inborn error of folate metabolism that results in megaloblastic anemia, pancytopenia and severe cerebral folate deficiency which can be corrected by folinic acid supplementation .[27]
Therapeutic applications
Since folate is needed by rapidly dividing cells to make thymine, this effect may be used to therapeutic advantage.
DHFR can be targeted in the treatment of cancer. DHFR is responsible for the levels of tetrahydrofolate in a cell, and the inhibition of DHFR can limit the growth and proliferation of cells that are characteristic of cancer.
Trimethoprim has shown to have activity against a variety of
Bacteria also need DHFR to grow and multiply and hence inhibitors selective for bacterial DHFR have found application as antibacterial agents.[30]
Classes of small-molecules employed as inhibitors of dihydrofolate reductase include diaminoquinazoline & diaminopyrroloquinazoline,[36] diaminopyrimidine, diaminopteridine and diaminotriazines.[37]
Potential anthrax treatment

Dihydrofolate reductase from Bacillus anthracis (BaDHFR) a validated drug target in the treatment of the infectious disease, anthrax. BaDHFR is less sensitive to trimethoprim analogs than is dihydrofolate reductase from other species such as Escherichia coli, Staphylococcus aureus, and Streptococcus pneumoniae. A structural alignment of dihydrofolate reductase from all four species shows that only BaDHFR has the combination phenylalanine and tyrosine in positions 96 and 102, respectively.
BaDHFR's resistance to trimethoprim analogs is due to these two residues (F96 and Y102), which also confer improved kinetics and catalytic efficiency.[38] Current research uses active site mutants in BaDHFR to guide lead optimization for new antifolate inhibitors.[38]
As a research tool
DHFR has been used as a tool to detect
CHO cells
DHFR lacking
Interactions
Dihydrofolate reductase has been shown to interact with GroEL[39] and Mdm2.[40]
Interactive pathway map
Click on genes, proteins and metabolites below to link to respective articles.[§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "FluoropyrimidineActivity_WP1601".
References
- PMID 6961421.
- PMID 6323448.
- PMID 6504041.
- PMID 500653.
- PMID 17920.
- PMID 6815179.
- ^ PMID 11502178.
- PMID 6815178.
- ^ a b "Entrez Gene: DHFR dihydrofolate reductase".
- ^ PMID 15139807.
- PMID 6933469.
- PMID 19666465.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - ^ ISSN 0006-2960.
- ISSN 0006-2960.
- ^ PMID 12021443.)
{{cite journal}}: CS1 maint: PMC format (link - ISSN 0006-2960.
- ^ ISSN 0006-2960.
- PMID 12359872.)
{{cite journal}}: CS1 maint: PMC format (link - ^ ISSN 0006-2960.
- ^ ISSN 0002-7863.
- ^ ISSN 0006-2960.
- ^ PMID 25453083.)
{{cite journal}}: CS1 maint: PMC format (link - ^ PMID 25453098.)
{{cite journal}}: CS1 maint: PMC format (link - ^ PMID 21138249.)
{{cite journal}}: CS1 maint: PMC format (link - ISSN 0006-2960.
- PMID 2601715.
- PMID 21310276.
- PMID 10623528.
- PMID 3125607.
- ^ PMID 16359642.
- PMID 7583655.
- PMID 8762155.
- PMID 12084458.
- PMID 19706381.
- PMID 8508427.
- PMID 25703118.
- PMID 26414808.
- ^ PMID 20882962.
- PMID 8559246.
- PMID 18451149.
Further reading
- Joska TM, Anderson AC (October 2006). "Structure-activity relationships of Bacillus cereus and Bacillus anthracis dihydrofolate reductase: toward the identification of new potent drug leads". Antimicrobial Agents and Chemotherapy. 50 (10): 3435–43. PMID 17005826.
- Chan DC, Fu H, Forsch RA, Queener SF, Rosowsky A (June 2005). "Design, synthesis, and antifolate activity of new analogues of piritrexim and other diaminopyrimidine dihydrofolate reductase inhibitors with omega-carboxyalkoxy or omega-carboxy-1-alkynyl substitution in the side chain". Journal of Medicinal Chemistry. 48 (13): 4420–31. PMID 15974594.
- Banerjee D, Mayer-Kuckuk P, Capiaux G, Budak-Alpdogan T, Gorlick R, Bertino JR (July 2002). "Novel aspects of resistance to drugs targeted to dihydrofolate reductase and thymidylate synthase". Biochimica et Biophysica Acta. 1587 (2–3): 164–73. PMID 12084458.
- Stockman BJ, Nirmala NR, Wagner G, Delcamp TJ, DeYarman MT, Freisheim JH (January 1992). "Sequence-specific 1H and 15N resonance assignments for human dihydrofolate reductase in solution". Biochemistry. 31 (1): 218–29. PMID 1731871.
- Beltzer JP, Spiess M (December 1991). "In vitro binding of the asialoglycoprotein receptor to the beta adaptin of plasma membrane coated vesicles". The EMBO Journal. 10 (12): 3735–42. PMID 1935897.
- Davies JF, Delcamp TJ, Prendergast NJ, Ashford VA, Freisheim JH, Kraut J (October 1990). "Crystal structures of recombinant human dihydrofolate reductase complexed with folate and 5-deazafolate". Biochemistry. 29 (40): 9467–79. PMID 2248959.
- Will CL, Dolnick BJ (December 1989). "5-Fluorouracil inhibits dihydrofolate reductase precursor mRNA processing and/or nuclear mRNA stability in methotrexate-resistant KB cells". The Journal of Biological Chemistry. 264 (35): 21413–21. PMID 2592384.
- Masters JN, Attardi G (March 1985). "Discrete human dihydrofolate reductase gene transcripts present in polysomal RNA map with their 5' ends several hundred nucleotides upstream of the main mRNA start site". Molecular and Cellular Biology. 5 (3): 493–500. PMID 2859520.
- Miszta H, Dabrowski Z, Lanotte M (November 1988). "In vitro patterns of enzymic tetrahydrofolate dehydrogenase (EC 1.5.1.3) expression in bone marrow stromal cells". Leukemia. 2 (11): 754–9. PMID 3185016.
- Oefner C, D'Arcy A, Winkler FK (June 1988). "Crystal structure of human dihydrofolate reductase complexed with folate". European Journal of Biochemistry / FEBS. 174 (2): 377–85. PMID 3383852.
- Yang JK, Masters JN, Attardi G (June 1984). "Human dihydrofolate reductase gene organization. Extensive conservation of the G + C-rich 5' non-coding sequence and strong intron size divergence from homologous mammalian genes". Journal of Molecular Biology. 176 (2): 169–87. PMID 6235374.
- Masters JN, Yang JK, Cellini A, Attardi G (June 1983). "A human dihydrofolate reductase pseudogene and its relationship to the multiple forms of specific messenger RNA". Journal of Molecular Biology. 167 (1): 23–36. PMID 6306253.
- Chen MJ, Shimada T, Moulton AD, Cline A, Humphries RK, Maizel J, Nienhuis AW (March 1984). "The functional human dihydrofolate reductase gene". The Journal of Biological Chemistry. 259 (6): 3933–43. PMID 6323448.
- Funanage VL, Myoda TT, Moses PA, Cowell HR (October 1984). "Assignment of the human dihydrofolate reductase gene to the q11----q22 region of chromosome 5". Molecular and Cellular Biology. 4 (10): 2010–6. PMID 6504041.
- Masters JN, Attardi G (1983). "The nucleotide sequence of the cDNA coding for the human dihydrofolic acid reductase". Gene. 21 (1–2): 59–63. PMID 6687716.
- Morandi C, Masters JN, Mottes M, Attardi G (April 1982). "Multiple forms of human dihydrofolate reductase messenger RNA. Cloning and expression in Escherichia coli of their DNA coding sequence". Journal of Molecular Biology. 156 (3): 583–607. PMID 6750132.
- Bonifaci N, Sitia R, Rubartelli A (September 1995). "Nuclear translocation of an exogenous fusion protein containing HIV Tat requires unfolding". AIDS. 9 (9): 995–1000. PMID 8527095.
- Mayhew M, da Silva AC, Martin J, Erdjument-Bromage H, Tempst P, Hartl FU (February 1996). "Protein folding in the central cavity of the GroEL-GroES chaperonin complex". Nature. 379 (6564): 420–6. PMID 8559246.
- Gross M, Robinson CV, Mayhew M, Hartl FU, Radford SE (December 1996). "Significant hydrogen exchange protection in GroEL-bound DHFR is maintained during iterative rounds of substrate cycling". Protein Science. 5 (12): 2506–13. PMID 8976559.
- Schleiff E, Shore GC, Goping IS (March 1997). "Human mitochondrial import receptor, Tom20p. Use of glutathione to reveal specific interactions between Tom20-glutathione S-transferase and mitochondrial precursor proteins". FEBS Letters. 404 (2–3): 314–8. PMID 9119086.
- Cody V, Galitsky N, Luft JR, Pangborn W, Rosowsky A, Blakley RL (November 1997). "Comparison of two independent crystal structures of human dihydrofolate reductase ternary complexes reduced with nicotinamide adenine dinucleotide phosphate and the very tight-binding inhibitor PT523". Biochemistry. 36 (45): 13897–903. PMID 9374868.
- Vanguri VK, Wang S, Godyna S, Ranganathan S, Liau G (April 2000). "Thrombospondin-1 binds to polyhistidine with high affinity and specificity". The Biochemical Journal. 347 (Pt 2): 469–73. PMID 10749676.
External links











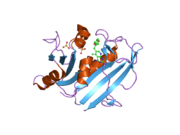
![1kms: HUMAN DIHYDROFOLATE REDUCTASE COMPLEXED WITH NADPH AND 6-([5-QUINOLYLAMINO]METHYL)-2,4-DIAMINO-5-METHYLPYRIDO[2,3-D]PYRIMIDINE (SRI-9439), A LIPOPHILIC ANTIFOLATE](http://upload.wikimedia.org/wikipedia/commons/thumb/c/cd/PDB_1kms_EBI.jpg/180px-PDB_1kms_EBI.jpg)
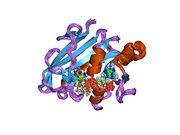
![1kmv: HUMAN DIHYDROFOLATE REDUCTASE COMPLEXED WITH NADPH AND (Z)-6-(2-[2,5-DIMETHOXYPHENYL]ETHEN-1-YL)-2,4-DIAMINO-5-METHYLPYRIDO[2,3-D]PYRIMIDINE (SRI-9662), A LIPOPHILIC ANTIFOLATE](http://upload.wikimedia.org/wikipedia/commons/thumb/6/62/PDB_1kmv_EBI.jpg/180px-PDB_1kmv_EBI.jpg)
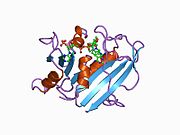
![1mvs: Analysis of Two Polymorphic Forms of a Pyrido[2,3-d]pyrimidine N9-C10 Reverse-Bridge Antifolate Binary Complex with Human Dihydrofolate Reductase](http://upload.wikimedia.org/wikipedia/commons/thumb/0/02/PDB_1mvs_EBI.jpg/180px-PDB_1mvs_EBI.jpg)
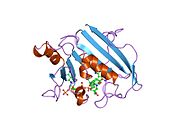
![1mvt: Analysis of Two Polymorphic Forms of a Pyrido[2,3-d]pyrimidine N9-C10 Reverse-Bridge Antifolate Binary Complex with Human Dihydrofolate Reductase](http://upload.wikimedia.org/wikipedia/commons/thumb/c/c3/PDB_1mvt_EBI.jpg/180px-PDB_1mvt_EBI.jpg)