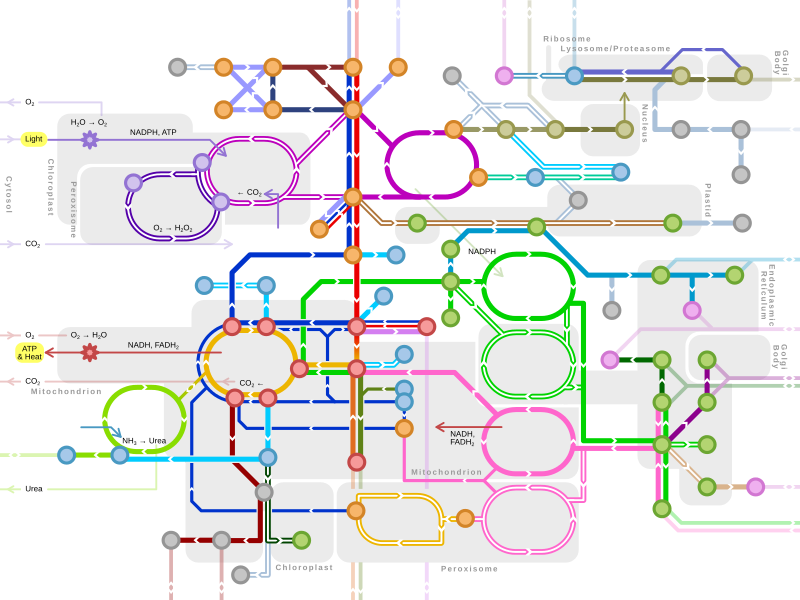
Protein metabolism
Protein metabolism denotes the various
The steps of protein synthesis include transcription, translation, and post translational modifications. During transcription,
Dietary proteins are first broken down to individual amino acids by various enzymes and hydrochloric acid present in the gastrointestinal tract. These amino acids are absorbed into the bloodstream to be transported to the liver and onward to the rest of the body. Absorbed amino acids are typically used to create functional proteins, but may also be used to create energy.[3] They can also be converted into glucose.[4] This glucose can then be converted to triglycerides and stored in fat cells.[5]
Proteins can be broken down by enzymes known as peptidases or can break down as a result of denaturation. Proteins can denature in environmental conditions the protein is not made for.[6]
Protein synthesis
Protein anabolism is the process by which proteins are formed from amino acids. It relies on five processes: amino acid synthesis, transcription, translation, post translational modifications, and protein folding. Proteins are made from amino acids. In humans, some amino acids can be synthesized using already existing intermediates. These amino acids are known as non-essential amino acids. Essential amino acids require intermediates not present in the human body. These intermediates must be ingested, mostly from eating other organisms.[6]
Amino Acid Synthesis
| Amino Acid | R-group ‡
|
Pathway* |
| Glycine | H- | hydroxymethyltransferase )
|
| Alanine | CH3- | aminotransferase )
|
| Valine§ | (CH3)2-CH- | Hydroxyethyl-TPP + Pyruvate → α-acetolactate → Valine
|
| Leucine§ | (CH3)2-CH-CH2- | Hydroxyethyl-TPP + Pyruvate → α-ketobutyrate → Leucine
|
| Isoleucine§ | CH3-CH2-CH(CH3)- | Hydroxyethyl-TPP + Pyruvate → α-acetolactate → Isoleucine
|
| Methionine§ | CH3-S-(CH2)2- | Homocysteine → Methionine (methionine synthase) |
| Proline | -(CH2)3- | Glutamic Acid → Glutamate-5-semialdehyde → Proline (γ-glutamyl kinase) |
| Phenylalanine§ | Ph-CH2- | Chorismate → Phenylalanine
|
| Tryptophan§ | Ph-NH-CH=C-CH2- | Phosphoenolpyruvate → 2-keto-3-deoxy arabino heptulosonate-7-phosphate → Chorismate → Tryptophan
|
| Tyrosine | HO-Ph-CH2- | Phenylalanine → Tyrosine (phenylalanine hydroxylase) |
| Serine | HO-CH2- | aminotransferase) → Serine (phosphoserine phosphatase )
|
| Threonine§ | CH3-CH(OH)- | Aspartate → β-aspartate-semialdehyde → Homoserine → Threonine |
| Cysteine | HS-CH2- | α-ketobutyrate → Cysteine
|
| Asparagine | H2N-CO-CH2- | Aspartic Acid → Asparagine (asparagine synthetase) |
| Glutamine | H2N-CO-(CH2)2- | Glutamic Acid → Glutamine (glutamine synthetase) |
| Arginine | +H2N=C(NH2)-NH-(CH2)3- | Glutamate → Glutamate-5-semialdehyde (γ-glutamyl kinase) → Arginine |
| Histidine§ | NH-CH=N-CH=C-CH2- | Glucose → Glucose-6-phosphate → Ribose-5-phosphate → Histidine |
| Lysine§ | +H3N-(CH2)4- | Aspartate → β-aspartate-semialdehyde → Homoserine + lysine |
| Aspartic Acid | −OOC-CH2- | aminotransferase )
|
| Glutamic Acid | −OOC-(CH2)2- | aminotransferase )
|
| ‡Shown at physiological conditions.
*Complexes that are italicized are enzymes. §Cannot be synthesized in humans. | ||
Polypeptide synthesis
Transcription

In
Transcription is regulated in the cell via transcription factors. Transcription factors are proteins that bind to regulatory sequences in the DNA strand such as promoter regions or operator regions. Proteins bound to these regions can either directly halt or allow RNA polymerase to read the DNA strand or can signal other proteins to halt or allow RNA polymerase reading.[11]
Translation

During translation, ribosomes convert a sequence of mRNA (messenger RNA) to an amino acid sequence. Each 3-base-pair-long segment of mRNA is a codon which corresponds to one amino acid or stop signal.[12] Amino acids can have multiple codons that correspond to them. Ribosomes do not directly attach amino acids to mRNA codons. They must utilize tRNAs (transfer RNAs) as well. Transfer RNAs can bind to amino acids and contain an anticodon which can hydrogen bind to an mRNA codon.[13] The process of bind an amino acid to a tRNA is known as tRNA charging. Here, the enzyme aminoacyl-tRNA-synthetase catalyzes two reactions. In the first one, it attaches an AMP molecule (cleaved from ATP) to the amino acid. The second reaction cleaves the aminoacyl-AMP producing the energy to join the amino acid to the tRNA molecule.[14]
Ribosomes have two subunits, one large and one small. These subunits surround the mRNA strand. The larger subunit contains three binding sites: A (aminoacyl), P (peptidyl), and E (exit). After translational initiation (which is different in prokaryotes and eukaryotes), the ribosome enters the elongation period which follows a repetitive cycle. First a tRNA with the correct amino acid enters the A site. The ribosome transfers the peptide from the tRNA in the P site to the new amino acid on the tRNA in the A site. The tRNA from the P site will be shifted into the E site where it will be ejected. This continually occurs until the ribosome reaches a stop codon or receives a signal to stop.[13] A peptide bond forms between the amino acid attached to the tRNA in the P site and the amino acid attached to a tRNA in the A site. The formation of a peptide bond requires an input of energy. The two reacting molecules are the alpha amino group of one amino acid and the alpha carboxyl group of the other amino acids. A by-product of this bond formation is the release of water (the amino group donates a proton while the carboxyl group donates a hydroxyl).[2]
Translation can be
Post-translational Modifications

Once the
Protein folding
A polypeptide chain in the cell does not have to stay linear; it can become branched or fold in on itself. Polypeptide chains fold in a particular manner depending on the solution they are in. The fact that all amino acids contain R groups with different properties is the main reason proteins fold. In a hydrophilic environment such as cytosol, the hydrophobic amino acids will concentrate at the core of the protein, while the hydrophilic amino acids will be on the exterior. This is entropically favorable since water molecules can move much more freely around hydrophilic amino acids than hydrophobic amino acids. In a hydrophobic environment, the hydrophilic amino acids will concentrate at the core of the protein, while the hydrophobic amino acids will be on the exterior. Since the new interactions between the hydrophilic amino acids are stronger than hydrophobic-hydrophilic interactions, this is enthalpically favorable.[18] Once a polypeptide chain is fully folded, it is called a protein. Often many subunits will combine to make a fully functional protein although physiological proteins do exist that contain only one polypeptide chain. Proteins may also incorporate other molecules such as the heme group in hemoglobin, a protein responsible for carrying oxygen in the blood.[19]