Fumarase
| FH | |||
|---|---|---|---|
Gene ontology | |||
| Molecular function | |||
| Cellular component | |||
| Biological process | |||
| Sources:Amigo / QuickGO | |||
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 1: 241.5 – 241.52 Mb | Chr 1: 175.43 – 175.45 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
| Fumarase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Fumarase (or fumarate hydratase) is an
This enzyme participates in 2 metabolic pathways: citric acid cycle and reductive citric acid cycle (CO2 fixation), and is also important in renal cell carcinoma. Mutations in this gene have been associated with the development of leiomyomas in the skin and uterus in combination with renal cell carcinoma (HLRCC syndrome).
Nomenclature
This enzyme belongs to the family of lyases, specifically the hydro-lyases, which cleave carbon-oxygen bonds. The systematic name of this enzyme class is (S)-malate hydro-lyase (fumarate-forming). Other names in common use include:
- fumarase
- L-malate hydro-lyase
- (S)-malate hydro-lyase
Structure
Gene

In humans, the FH gene is localized to the chromosomal position 1q42.3-q43. The FH gene contains 10 exons.
Protein
Crystal structures of fumarase C from
Subtypes
There are two classes of fumarases, class I and class II.[7] Classification depends on the arrangement of their relative subunits, their metal ion requirement, and their thermal stability. Class I fumarases are change state or become inactive when subjected to heat or radiation, are sensitive to superoxide anion, are iron (Fe2+) dependent, and are dimeric proteins with each subunit consisting of around 120 kD. Class II fumarases, found in prokaryotes as well as in eukaryotes, are tetrameric enzymes with subunits of 200 kD that contain three distinct segments of significantly homologous amino acids. They are also iron-independent and thermally stable. Prokaryotes are known to have three different forms of fumarase: Fumarase A, Fumarase B, and Fumarase C. Fumarase A and Fumarase B from Escherichia coli are classified as class I, whereas Fumarase C is a part of the class II fumarases.[8]
Function
Mechanism

Figure 1 depicts the fumarase reaction mechanism. Two residues catalyze proton transfer and the ionization state of these residues is in part defined by two forms of the enzyme, E1 and E2. In E1, the groups exist in an internally neutralized AH/B: state, while in E2, they occur in a zwitterionic A−/BH+ state. E1 binds fumarate and facilitates its transformation into malate, and E2 binds malate and facilitates its transformation into fumarate. The two forms must undergo isomerization with each catalytic turnover.[9]

Despite its biological significance, the reaction mechanism of fumarase is not completely understood. The reaction itself can be monitored in either direction; however, it is the formation of fumarate from S-malate in particular that is less understood due to the high
Biochemical pathway
The function of fumarase in the
Other substrates
The main substrates for fumarase are malate and fumarate. However, the enzyme can also catalyze the dehydration of D-
Clinical significance
Fumarase deficiency is characterized by polyhydramnios and fetal brain abnormalities. In the newborn period, findings include severe neurologic abnormalities, poor feeding, failure to thrive, and hypotonia. Fumarase deficiency is suspected in infants with multiple severe neurologic abnormalities in the absence of an acute metabolic crisis. Inactivity of both cytosolic and mitochondrial forms of fumarase are potential causes. Isolated, increased concentration of fumaric acid on urine organic acid analysis is highly suggestive of fumarase deficiency. Molecular genetic testing for fumarase deficiency is currently available.[7]
Fumarase is prevalent in both fetal and adult tissues. A large percentage of the enzyme is expressed in the
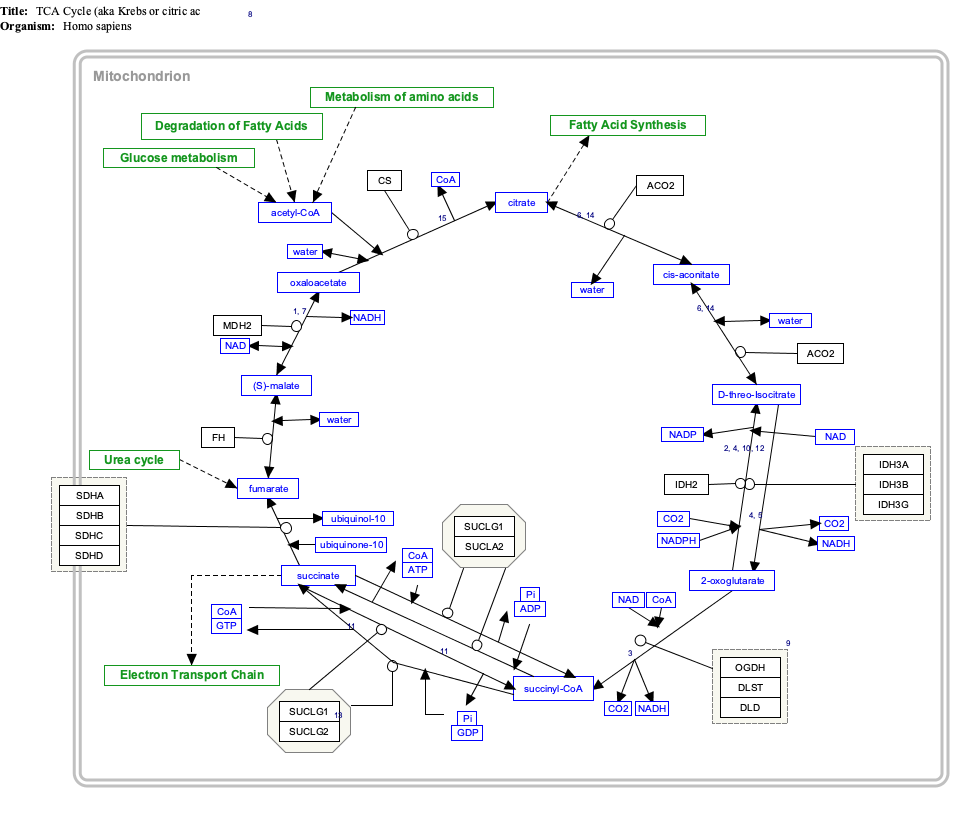
Interactive pathway map
Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "TCACycle_WP78".
See also
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000091483 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000026526 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ FH (fumarate hydratase)
- ^ PMID 16204892.
- ^ a b c Lynch AM, Morton CC (2006-07-01). "FH (fumarate hydratase)". Atlas of Genetics and Cytogenetics in Oncology and Haematology.
- ^ PMID 12021453.
- ^ ISBN 978-0-19-512258-9.
- ^ ISBN 978-0-9747077-1-6.
- ^ ISBN 978-0-7167-0070-8.
- PMID 21929734.
- ^ PMID 23405168.
External links
- Fumarase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Structure of Fumarate
- Structure of S-Malate
- Link to Breakdown of Citric Acid Cycle
- Video of Fumarate → (S)L-Malate