Isocitrate dehydrogenase
| Isocitrate dehydrogenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| isocitrate dehydrogenase (NAD+) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Monomeric isocitrate dehydrogenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
Isocitrate dehydrogenase (IDH) (
Structure

The NAD-IDH is composed of 3 subunits, is allosterically regulated, and requires an integrated Mg2+ or Mn2+ ion. The closest homologue that has a known structure is the E. coli NADP-dependent IDH, which has only 2 subunits and a 13% identity and 29% similarity based on the amino acid sequences, making it dissimilar to human IDH and not suitable for close comparison. All the known NADP-IDHs are homodimers.
Most isocitrate dehydrogenases are dimers, to be specific, homodimers (two identical monomer subunits forming one dimeric unit). In comparing C. glutamicum and E. coli,[4] monomer and dimer, respectively, both enzymes were found to "efficiently catalyze identical reactions." However, C. glutamicum was recorded as having ten times as much activity than E. coli and seven times more affinitive/specific for NADP. C. glutamicum favored NADP+ over NAD+. In terms of stability with response to temperature, both enzymes had a similar Tm or melting temperature at about 55 °C to 60 °C. However, the monomer C. glutamicum showed a more consistent stability at higher temperatures, which was expected. The dimer E. coli showed stability at a higher temperature than normal due to the interactions between the two monomeric subunits.
The structure of Mycobacterium tuberculosis (Mtb) ICDH-1 bound with NADPH and Mn(2+) bound has been solved by X-ray crystallography. It is a homodimer in which each subunit has a Rossmann fold, and a common top domain of interlocking β sheets. Mtb ICDH-1 is most structurally similar to the R132H mutant human ICDH found in CNS WHO grade 4 astrocytomas, formerly classified[5] as glioblastomas. Similar to human R132H ICDH, Mtb ICDH-1 also catalyzes the formation of α-hydroxyglutarate.[6]
Regulation
The IDH step of the citric acid cycle is often (but not always) an irreversible reaction due to its large negative change in free energy. It must therefore be carefully regulated to avoid depletion of isocitrate (and therefore an accumulation of alpha-ketoglutarate). The reaction is stimulated by the simple mechanisms of substrate availability (isocitrate,
Catalytic mechanisms
Isocitrate dehydrogenase catalyzes the chemical reactions:
- Isocitrate + NAD+ 2-oxoglutarate + CO2 + NADH + H+
- Isocitrate + NADP+ 2-oxoglutarate + CO2 + NADPH + H+[9][10][11]
The overall free energy for this reaction is -8.4 kJ/mol.[12]

Steps
Within the
.In the provided figure, the first box shows the overall isocitrate dehydrogenase reaction. The necessary reactants for this enzyme mechanism are isocitrate,
The second box in the figure illustrates step 1 of the reaction, which is the oxidation of the
The third box illustrates step 2, which is the decarboxylation of
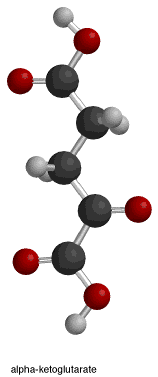
The fourth and final box illustrates step 3, which is the saturation of the alpha-beta unsaturated double bond that formed in the previous step. The negatively charged oxygen (attached to the alpha-carbon) donates its electrons, reforming the ketone double bond and pushing another lone pair (the one that forms the double bond between the alpha and beta carbons) "off" the molecule. This lone pair, in turn, picks up a proton from the nearby tyrosine.[14] This reaction results in the formation of alpha-ketoglutarate, NADH + H+/NADPH + H+, and CO2.
Detailed mechanism
Two
The decarboxylation of oxalosuccinate (below center) is a key step in the formation of alpha-ketoglutarate. In this reaction, the lone pair on the adjacent Tyrosine hydroxyl abstracts the proton off the carboxyl group.[14] This carboxyl group is also referred to as the beta subunit in the isocitrate molecule. The deprotonation of the carboxyl group causes the lone pair of electrons to move down making carbon dioxide and separating from oxalosuccinate. The electrons continue to move towards the alpha carbon pushing the double bond electrons (making the ketone) up to abstract a proton off an adjacent lysine residue. An alpha-beta unsaturated double bond results between carbon 2 and three. As you can see in the picture, the green ion represents either Mg2+ or Mn2+, which is a cofactor necessary for this reaction to occur. The metal-ion forms a little complex through ionic interactions with the oxygen atoms on the fourth and fifth carbons (also known as the gamma subunit of isocitrate).
After the carbon dioxide is split from the oxalosuccinate in the decarboxylation step (below right), the
 |
 |
 |
The isocitrate dehydrogenase enzyme as stated above produces alpha-ketoglutarate, carbon dioxide, and NADH + H+/NADPH + H+. There are three changes that occurred throughout the reaction. The oxidation of Carbon 2, the decarboxylation (loss of carbon dioxide) off Carbon 3, and the formation of a ketone group with a stereochemical change from sp3 to sp2.[14]
 |
 |
 |
 |
Active site
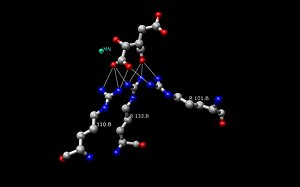
The Isocitrate Dehydrogenase (IDH) enzyme structure in Escherichia coli was the first IDH ortholog structure to be elucidated and understood.[14] Since then, the Escherichia coli IDH structure has been used by most researchers to make comparisons to other isocitrate dehydrogenase enzymes. There is much detailed knowledge about this bacterial enzyme, and it has been found that most isocitrate dehydrogenases are similar in structure and therefore also in function. This similarity of structure and function gives a reason to believe that the structures are conserved as well as the amino acids.[11] Therefore, the active sites amongst most prokaryotic isocitrate dehydrogenase enzymes should be conserved as well, which is observed throughout many studies done on prokaryotic enzymes. Eukaryotic isocitrate dehydrogenase enzymes on the other hand, have not been fully discovered yet. Each dimer of IDH has two active sites.[14] Each active site binds a NAD+/NADP+ molecule and a divalent metal ion (Mg2+,Mn2+). In general, each active site has a conserved sequence of amino acids for each specific binding site. In Desulfotalea psychrophila (DpIDH)[14] and porcine (PcIDH)[3] there are three substrates bound to the active site.
- Isocitrate binds within the active site to a conserved sequence of about eight amino acids through hydrogen bonds. These acids include (may vary in residue but with similar properties) tyrosine, serine, asparagine, arginine, arginine, arginine, tyrosine, and lysine. Their positions on the backbone vary but they are all within a close range (i.e. Arg131 DpIDH and Arg133 PcIDH, Tyr138 DpIDH and Tyr140 PcIDH).[14]
- The metal ion (Mg2+, Mn2+) binds to three conserved amino acids through hydrogen bonds. These amino acids include three Aspartate residues.[14]
- NAD+ and NADP+ bind within the active site within four regions with similar properties amongst IDH enzymes. These regions vary but are around [250–260], [280–290], [300–330], and [365–380]. Again regions vary but the proximity of regions are conserved.[14]
Clinical significance

Specific mutations in the isocitrate dehydrogenase gene IDH1 have been found in several tumor types,[16] notably brain tumors including astrocytoma and oligodendroglioma.[5] Patients whose tumor had an IDH1 mutation had longer survival compared to patients whose tumor had an IDH1 wild type.[17][18] Furthermore, mutations of IDH2 and IDH1 were found in up to 20% of cytogenetically normal acute myeloid leukemia (AML).[19][20] These mutations are known to produce D-2-hydroxyglutarate from alpha-ketoglutarate.[21] D-2-hydroxyglutarate accumulates to very high concentrations which inhibits the function of enzymes that are dependent on alpha-ketoglutarate.[22] This leads to a hypermethylated state of DNA and histones, which results in different gene expression that can activate oncogenes and inactivate tumor-suppressor genes. Ultimately, this may lead to the types of cancer described above.[23] Somatic mosaic mutations of this gene have also been found associated to Ollier disease and Maffucci syndrome.[24] However, recent studies have also shown that D-2-hydroxyglutarate may be converted back into alpha-ketoglutarate either enzymatically or non-enzymatically.[25][26] Further studies are required to fully understand the roles of IDH1 mutation (and D-2-hydroxyglutarate) in cancer. Recent research highlighted cancer-causing mutations in isocitrate dehydrogenase which may cause accumulation of the metabolite D-2-hydroxyglutarate (D-2HG). Notarangelo et al. showed that such high concentrations of D-2HG could act as a direct inhibitor of lactate dehydrogenase in mouse T cells. Inhibition of this metabolic enzyme altered glucose metabolism in the T cells and inhibited their proliferation, cytokine production, and ability to kill target cells.[27]
Isozymes
The following is a list of human isocitrate dehydrogenase isozymes:
NADP+ dependent
Each NADP+-dependent isozyme functions as a homodimer:
|
| |||||||||||||||||||||||||||
NAD+ dependent
The isocitrate dehydrogenase 3 isozyme is a heterotetramer that is composed of two alpha subunits, one beta subunit, and one gamma subunit:
|
|
| |||||||||||||||||||||||||||||||||||||||||||||
See also
- Oxidoreductase
- Myelodysplastic syndrome#IDH1 and IDH2 mutations
- Oncometabolism
- Isocitrate/isopropylmalate dehydrogenase family
References
- PMID 10623532.
- PMID 10557241.
- ^ PMID 12207025.
- ^ PMID 11185559.
- ^ PMID 34185076.
- PMID 23409873.
- .
- S2CID 234361097.
- ^ ISBN 0-9747077-1-6.
- ^ ISBN 0-7167-4339-6.
- ^ PMID 12855708.
- ISBN 978-1-133-10629-6.
- PMID 18203822.
- ^ PMID 17632124.
- S2CID 36093146.
- S2CID 245547597.
- S2CID 7323032.
- PMID 24510240.
- ^
Ward PS, Patel J, Wise DR, Abdel-Wahab O, Bennett BD, Coller HA, et al. (March 2010). "The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate". Cancer Cell. 17 (3): 225–234. PMID 20171147.
- PMID 25601757.
- PMID 20559394.
- PMID 21460794.
- PMID 24880135.
- S2CID 5592593.
- PMID 24594748.
- PMID 22343896.
- S2CID 252623286.
External links
- Isocitrate dehydrogenase: RCSB PDB Molecule of the Month Archived 2013-12-24 at the Wayback Machine
- Overview of all the structural information available in the PDB for UniProt: O75874 (Isocitrate dehydrogenase [NADP] cytoplasmic) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: P48735 (Isocitrate dehydrogenase [NADP], mitochondrial) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: P50213 (Isocitrate dehydrogenase [NAD] subunit alpha, mitochondrial) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: O43837 (Isocitrate dehydrogenase [NAD] subunit beta, mitochondrial) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: P51553 (Isocitrate dehydrogenase [NAD] subunit gamma, mitochondrial) at the PDBe-KB.

