IDH3B
| IDH3B | |||
|---|---|---|---|
| Identifiers | |||
Gene ontology | |||
| Molecular function | |||
| Cellular component | |||
| Biological process | |||
| Sources:Amigo / QuickGO | |||
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt |
| ||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) |
| ||||||||
| Location (UCSC) | Chr 20: 2.66 – 2.66 Mb | Chr 2: 130.12 – 130.13 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Isocitrate dehydrogenase [NAD] subunit beta, mitochondrial is an enzyme that in humans is encoded by the IDH3B gene.[5][6]
heterotetramer that is composed of two alpha subunits, one beta subunit, and one gamma subunit. The protein encoded by this gene is the beta subunit of one isozyme of NAD(+)-dependent isocitrate dehydrogenase. Three alternatively spliced transcript variants encoding different isoforms have been described for this gene. [provided by RefSeq, Jul 2008][6]
Structure
IDH3 is one of three isocitrate dehydrogenase isozymes, the other two being
substrate isocitrate, all three subunits participate in the catalytic reaction.[10][11] Moreover, studies of the enzyme in pig heart reveal that the αβ and αγ dimers constitute two binding sites for each of its ligands, including isocitrate, Mn2+, and NAD, in one IDH3 tetramer.[9][10]
Isoforms
The IDH3B gene contains 12
alternatively spliced isoforms: IDH3β1 (349 residues) and IDH3β2 (354 residues).[13][14] These isoforms are tissue-specific and possess optimal pHs matching those of their target tissues. IDH3β1, with an optimal pH of 8.0, is expressed in brain and kidney, whereas IDH3β2, with an optimal pH of 7.6, is expressed in heart and skeletal muscle.[14]
Function
As an isocitrate dehydrogenase, IDH3 catalyzes the reversible oxidative decarboxylation of isocitrate to yield
Clinical Significance
Homozygous loss-of-function mutations of the IDH3B gene has been linked to retinitis pigmentosa, the neurodegeneration of rods and cones in the retina resulting in blindness.[12][13][17]
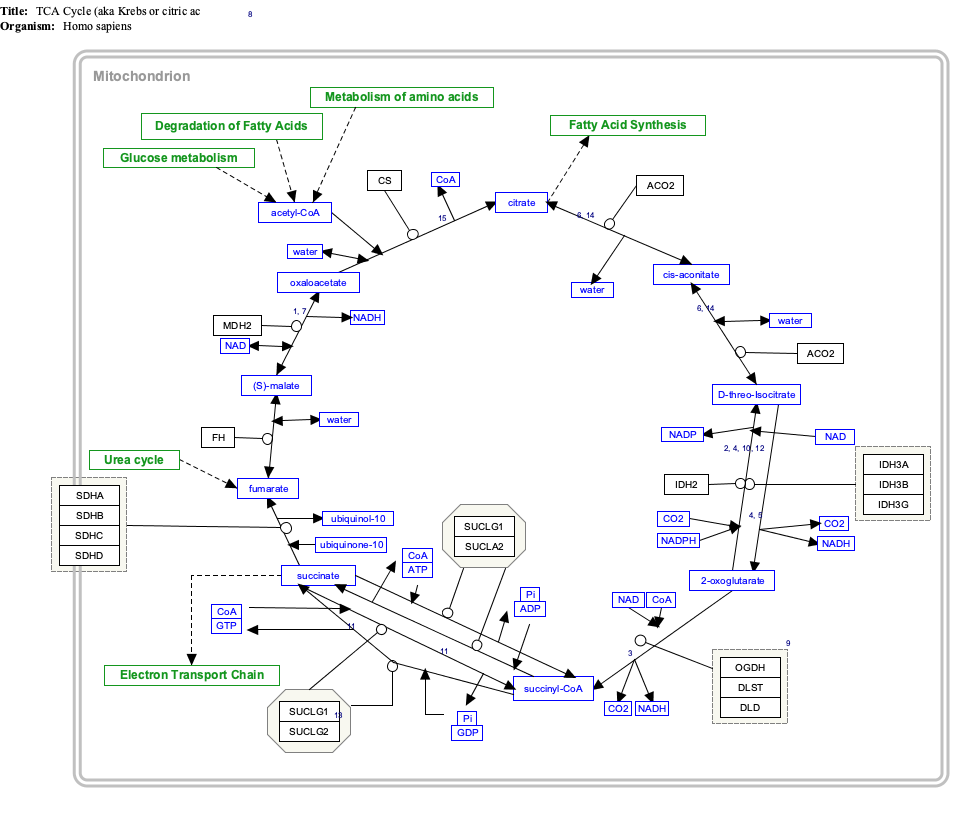
Interactive pathway map
Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
TCACycle_WP78 edit
- ^ The interactive pathway map can be edited at WikiPathways: "TCACycle_WP78".
See also
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000101365 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000027406 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- S2CID 85139434.
- ^ a b "Entrez Gene: IDH3B isocitrate dehydrogenase 3 (NAD+) beta".
- PMID 25678837.
- ^ PMID 25531325.
- ^ PMID 17432878.
- ^ PMID 16737955.
- ^ PMID 14555658.
- ^ PMID 20435888.
- ^ PMID 18806796.
- ^ PMID 10601238.
- ^ PMID 8833160.
- PMID 26782057.
- )
Further reading
- Kim YO, Koh HJ, Kim SH, et al. (2000). "Identification and functional characterization of a novel, tissue-specific NAD(+)-dependent isocitrate dehydrogenase beta subunit isoform". J. Biol. Chem. 274 (52): 36866–75. PMID 10601238.
- Weiss C, Zeng Y, Huang J, et al. (2000). "Bovine NAD+-dependent isocitrate dehydrogenase: alternative splicing and tissue-dependent expression of subunit 1". Biochemistry. 39 (7): 1807–16. PMID 10677231.
- Deloukas P, Matthews LH, Ashurst J, et al. (2002). "The DNA sequence and comparative analysis of human chromosome 20". Nature. 414 (6866): 865–71. PMID 11780052.
- Strausberg RL, Feingold EA, Grouse LH, et al. (2003). "Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences". Proc. Natl. Acad. Sci. U.S.A. 99 (26): 16899–903. PMID 12477932.
- Ota T, Suzuki Y, Nishikawa T, et al. (2004). "Complete sequencing and characterization of 21,243 full-length human cDNAs". Nat. Genet. 36 (1): 40–5. PMID 14702039.
- Gerhard DS, Wagner L, Feingold EA, et al. (2004). "The status, quality, and expansion of the NIH full-length cDNA project: the Mammalian Gene Collection (MGC)". Genome Res. 14 (10B): 2121–7. PMID 15489334.
- Kil IS, Park JW (2005). "Regulation of mitochondrial NADP+-dependent isocitrate dehydrogenase activity by glutathionylation". J. Biol. Chem. 280 (11): 10846–54. PMID 15653693.
- Rual JF, Venkatesan K, Hao T, et al. (2005). "Towards a proteome-scale map of the human protein-protein interaction network". Nature. 437 (7062): 1173–8. S2CID 4427026.
- Lim J, Hao T, Shaw C, et al. (2006). "A protein-protein interaction network for human inherited ataxias and disorders of Purkinje cell degeneration". Cell. 125 (4): 801–14. S2CID 13709685.
- Soundar S, O'hagan M, Fomulu KS, Colman RF (2006). "Identification of Mn2+-binding aspartates from alpha, beta, and gamma subunits of human NAD-dependent isocitrate dehydrogenase". J. Biol. Chem. 281 (30): 21073–81. PMID 16737955.
- Bzymek KP, Colman RF (2007). "Role of alpha-Asp181, beta-Asp192, and gamma-Asp190 in the distinctive subunits of human NAD-specific isocitrate dehydrogenase". Biochemistry. 46 (18): 5391–7. PMID 17432878.