BRAF (gene)
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) |
| ||||||||
| Location (UCSC) | Chr 7: 140.72 – 140.92 Mb | Chr 6: 39.58 – 39.7 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
BRAF is a human
The B-Raf protein is involved in sending
Certain other inherited BRAF mutations cause birth defects.
Drugs that treat cancers driven by BRAF mutations have been developed. Two of these drugs,
Function
Structure
B-Raf is a 766-
CR1
Conserved region 1 (CR1)
CR2
Conserved region 2 (CR2) provides a flexible linker that connects CR1 and CR3 and acts as a hinge.[citation needed]
CR3
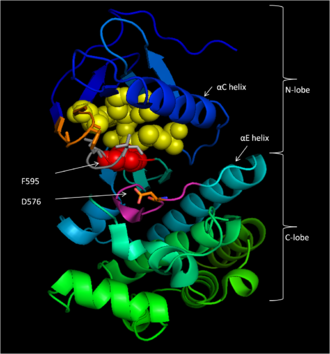
Conserved region 3 (CR3), residues 457–717,
Subregions
P-Loop
The P-loop of B-Raf (residues 464–471) stabilizes the non-transferable phosphate groups of ATP during enzyme ATP-binding. Specifically, S467, F468, and G469 backbone amides hydrogen-bond to the β-phosphate of ATP to anchor the molecule. B-Raf functional motifs have been determined by analyzing the homology of PKA analyzed by Hanks and Hunter to the B-Raf kinase domain.[16]
Nucleotide-binding pocket
V471, C532, W531, T529, L514, and A481 form a hydrophobic pocket within which the adenine of ATP is anchored through Van der Waals attractions upon ATP binding.[16][18]
Catalytic loop
Residues 574–581 compose a section of the kinase domain responsible for supporting the transfer of the γ-phosphate of ATP to B-Raf's protein substrate. In particular, D576 acts as a proton acceptor to activate the nucleophilic hydroxyl oxygen on substrate serine or threonine residues, allowing the phosphate transfer reaction to occur mediated by base-catalysis.[16]
DFG motif
Activation loop
Residues 596–600 form strong hydrophobic interactions with the P-loop in the inactive conformation of the kinase, locking the kinase in its inactive state until the
Enzymology
B-Raf is a
Activation
Relieving CR1 autoinhibition
The kinase (CR3) domain of human
CR3 domain activation
After the autoinhibitory CR1 regulatory domain is released, B-Raf's CR3
Mechanism of catalysis

To effectively catalyze protein phosphorylation via the bimolecular substitution of serine and threonine residues with ADP as a leaving group, B-Raf must first bind ATP and then stabilize the transition state as the γ-phosphate of ATP is transferred.[16]
ATP binding
B-Raf binds ATP by anchoring the adenine nucleotide in a
Phosphorylation
Once ATP is bound to the B-Raf kinase domain, D576 of the catalytic loop activates a substrate hydroxyl group, increasing its nucleophilicity to kinetically drive the phosphorylation reaction while other catalytic loop residues stabilize the transition state (Figure 2). N581 chelates the divalent magnesium cation associated with ATP to help orient the molecule for optimal substitution. K578 neutralizes the negative charge on the γ-phosphate group of ATP so that the activated ser/thr substrate residue will not experience as much electron-electron repulsion when attacking the phosphate. After the phosphate group is transferred, ADP and the new phosphoprotein are released.[16]
Inhibitors
Since constitutively active B-Raf mutants commonly cause cancer (see Clinical Significance) by excessively signaling cells to grow, inhibitors of B-Raf have been developed for both the inactive and active conformations of the kinase domain as cancer therapeutic candidates.[17][18][19]
Sorafenib
BAY43-9006 (
The distal
The trifluoromethyl phenyl moiety cements the thermodynamic favorability of the inactive conformation when the kinase domain is bound to BAY43-9006 by sterically blocking the hydrophobic pocket between the αC and αE helices that the DFG motif and activation loop would inhabit upon shifting to their locations in the active conformation of the protein.[17]
Vemurafenib

PLX4032 (Vemurafenib) is a V600 mutant B-Raf inhibitor approved by the FDA for the treatment of late-stage melanoma.[13] Unlike BAY43-9006, which inhibits the inactive form of the kinase domain, Vemurafenib inhibits the active "DFG-in" form of the kinase,[18][19] firmly anchoring itself in the ATP-binding site. By inhibiting only the active form of the kinase, Vemurafenib selectively inhibits the proliferation of cells with unregulated B-Raf, normally those that cause cancer.
Since Vemurafenib only differs from its precursor, PLX4720, in a
Clinical significance
Mutations in the BRAF gene can cause disease in two ways. First, mutations can be inherited and cause birth defects. Second, mutations can appear later in life and cause cancer, as an oncogene.
Inherited mutations in this gene cause cardiofaciocutaneous syndrome, a disease characterized by heart defects, mental retardation and a distinctive facial appearance.[24]
Mutations in this gene have been found in cancers, including
The V600E mutation of the BRAF gene has been associated with
Mutants
More than 30 mutations of the BRAF gene associated with human cancers have been identified. The frequency of BRAF mutations varies widely in human cancers, from more than 80% in
Other mutations which have been found are R461I, I462S, G463E, G463V, G465A, G465E, G465V, G468A, G468E, G469R, N580S, E585K, D593V, F594L, G595R, L596V, T598I, V599D, V599E, V599K, V599R, V600K, A727V, etc. and most of these mutations are clustered to two regions: the glycine-rich P loop of the N lobe and the activation segment and flanking regions.[17] These mutations change the activation segment from inactive state to active state, for example in the previous cited paper it has been reported that the aliphatic side chain of Val599 interacts with the phenyl ring of Phe467 in the P loop. Replacing the medium-sized hydrophobic Val side chain with a larger and charged residue as found in human cancer(Glu, Asp, Lys, or Arg) would be expected to destabilize the interactions that maintain the DFG motif in an inactive conformation, so flipping the activation segment into the active position. Depending on the type of mutation the kinase activity towards MEK may also vary. Most of the mutants stimulate enhanced B-Raf kinase activity toward MEK. However, a few mutants act through a different mechanism because although their activity toward MEK is reduced, they adopt a conformation that activates wild-type C-RAF, which then signals to ERK.
BRAF-V600E
- BRAFV600E mutation confers a poor prognosis in metastatic colorectal cancer. Upon targeted inhibition of BRAF and/or EGFR in BRAF V600E colorectal cancer, SRC kinases become activated in a systematic manner. Remarkably, concurrent targeting of SRC alongside BRAF and EGFR has demonstrated increased treatment effectiveness. The compensatory activation of SRC kinases is mediated by an autocrine prostaglandin E2 loop, which can be blocked using cyclooxygenase-2 (COX2) inhibitors. Notably, the simultaneous targeting of COX2 with BRAF and EGFR has shown significant potential in preclinical models, leading to a sustained suppression of tumor growth.[40]
- BRAF V600E is a determinant of sensitivity to proteasome inhibitors. Vulnerability to proteasome inhibitors is dependent on persistent BRAF signaling, because BRAF-V600E blockade by PLX4720 reversed sensitivity to carfilzomib in BRAF-mutant colorectal cancer cells. Proteasome inhibition might represent a valuable targeting strategy in BRAF V600E-mutant colorectal tumors.[41]
BRAF inhibitors
As mentioned above, some pharmaceutical firms are developing specific inhibitors of mutated B-raf protein for anticancer use because BRAF is a well-understood, high yield target.[18][42] Vemurafenib (RG7204 or PLX4032) was licensed by the US Food and Drug Administration as Zelboraf for the treatment of metastatic melanoma in August 2011 based on Phase III clinical data. Improved survival was seen, as well as a response rate to treatment of 53%, compared to 7–12% with the former best chemotherapeutic treatment, dacarbazine.[43] In clinical trials, B-Raf increased metastatic melanoma patient chance of survival. In spite of the drug's high efficacy, 20% of tumors still develop resistance to the treatment. In mice, 20% of tumors become resistant after 56 days.[44] While the mechanisms of this resistance are still disputed, some hypotheses include the overexpression of B-Raf to compensate for high concentrations of Vemurafenib[44] and upstream upregulation of growth signaling.[45]
More general
.panRAF inhibitors
Belvarafenib is classified as a panRAF inhibitor. A panRAF inhibitor blocks the catalytic function of both proteins in the dimer.[46]
Interactions
BRAF (gene) has been shown to
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000157764 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000002413 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- PMID 2284096.
- PMID 1565476.
- S2CID 3071547.
- ^ "FDA Approves Zelboraf (Vemurafenib) and Companion Diagnostic for BRAF Mutation-Positive Metastatic Melanoma, a Deadly Form of Skin Cancer" (Press release). Genentech. Retrieved 2011-08-17.
- S2CID 19634793.
- ^ a b "Entrez Gene: BRAF".
- PMID 7855890.
- ^ PMID 9689060.
- ^ S2CID 9337155.
- ^ a b c d "Serine/threonine protein kinase B-rAF". Retrieved 4 Mar 2013.
- ^ PMID 9069260.
- ^ S2CID 21377422.
- ^ S2CID 126161.
- ^ PMID 18287029.
- ^ PMID 20823850.
- PMID 3291115.
- .
- PMID 1956325.
- PMID 10205168.
- PMID 16825433.
- S2CID 204843221.
- PMID 19125127.
- PMID 12970315.
- S2CID 32051681.
- PMID 16403224.
- PMID 17065421.
- PMID 14734469.
- PMID 15277467.
- PMID 14679157.
- PMID 15126572.
- PMID 18682506.
- S2CID 34922416.
- PMID 21663470. *Lay summary in: "Research on hairy-cell leukaemia shows the promise of new DNA-scanning technologies". Cancer Research UK. June 11, 2011.
- PMID 24374844.
- PMID 24413733.*Lay summary in: Leah Eisenstadt (30 January 2014). "Single driver mutation found in rare brain tumor". BROAD Institute.
- PMID 36759733.
- S2CID 17012966.
- PMID 17145850.
- PMID 21639808.
- ^ PMID 23302800.
- PMID 21107323.
- PMID 35582307.
- PMID 10869359.
- PMID 11325826.
- PMID 9154803.
- PMID 7706312.
- PMID 17353931.
- PMID 10931830.
Further reading
- Garnett MJ, Marais R (October 2004). "Guilty as charged: B-RAF is a human oncogene". Cancer Cell. 6 (4): 313–319. PMID 15488754.
- Quiros RM, Ding HG, Gattuso P, Prinz RA, Xu X (June 2005). "Evidence that one subset of anaplastic thyroid carcinomas are derived from papillary carcinomas due to BRAF and p53 mutations". Cancer. 103 (11): 2261–2268. S2CID 29665029.
- Karbowniczek M, Henske EP (November 2005). "The role of tuberin in cellular differentiation: are B-Raf and MAPK involved?". Annals of the New York Academy of Sciences. 1059 (1): 168–173. S2CID 39146204.
- Ciampi R, Nikiforov YE (March 2007). "RET/PTC rearrangements and BRAF mutations in thyroid tumorigenesis". Endocrinology. 148 (3): 936–941. PMID 16946010.
- Espinosa AV, Porchia L, Ringel MD (January 2007). "Targeting BRAF in thyroid cancer". British Journal of Cancer. 96 (1): 16–20. PMID 17179987.
- Allanson JE, Roberts AE (8 August 2019). "Noonan Syndrome". In Pagon RA, Bird TD, Dolan CR, et al. (eds.). GeneReviews [Internet]. Seattle WA: University of Washington, Seattle. PMID 20301303.
- Rauen KA (3 March 2016) [18 January 2007]. "Cardiofaciocutaneous Syndrome". In Pagon RA, Bird TD, Dolan CR (eds.). GeneReviews [Internet]. Seattle WA: University of Washington, Seattle. PMID 20301365.
- Gelb BD, Tartaglia M (14 May 2015) [30 November 2007]. "LEOPARD Syndrome". In Pagon RA, Bird TD, Dolan CR (eds.). GeneReviews [Internet]. Seattle WA: University of Washington, Seattle. PMID 20301557.
External links
- "BRAF gene". NCI Dictionary of Cancer Terms. Retrieved 2007-11-25.
- Finding faults in BRAF Archived 2018-08-19 at the Wayback Machine — Cancer Research UK blog post about the discovery of cancer-causing BRAF mutations (incl video)
- Human BRAF genome location and BRAF gene details page in the UCSC Genome Browser.
![]() This article incorporates
This article incorporates