User:Estevezj/sandbox/Genomics
Genomics is a discipline in
History
Etymology
While the word 'genome' (from the German Genom, attributed to Hans Winkler) was in use in English as early as 1926,
Early sequencing efforts
Following
DNA sequencing technology developed
In addition to his seminal work on the amino acid sequence of insulin,
Complete genomes
The advent of these technologies resulted in a rapid intensification in the scope and speed of completion of

Most of the microorganisms whose genomes have been completely sequenced are problematic
A rough draft of the human genome was completed by the Human Genome Project in early 2001, creating much fanfare.[34] This project, completed in 2003, sequenced the entire genome for one specific person, and by 2007 this sequence was declared "finished" (less than one error in 20,000 bases and all chromosomes assembled).[34] In the years since then, the genomes of many other individual people have been sequenced, partly under the auspices of the 1000 Genomes Project, which announced the sequencing of 1,092 genomes in October 2012.[35] Completion of this project was made possible by the development of dramatically more efficient sequencing technologies and required the commitment of significant bioinformatics resources from large international collaboration.[36] The continued analysis of human genomic data has profound political and social repercussions for human societies.[37]
Next-generation sequencing
| Latin Name | Common Name | Genome Size |
|---|---|---|
| Eukaryotes | ||
| Lilium longiflorum | Easter lily | 90,000,000 Kb |
Homo sapiens
|
Human | 3,200,000 Kb |
| Oryza sativa | Rice | 420,000 Kb |
| Drosophila melanogaster | Fruit fly | 137,000 Kb |
| Arabidopsis thaliana | Mustard cress | 115,428 Kb |
| Caenorhabditis elegans | Roundworm | 97,000 Kb |
| Saccharomyces cerevisiae | Yeast | 12,069 Kb |
| Eubacteria | ||
| Haemophilus influenzae | Pfeiffer's bacillus | 1,830 Kb |
| Escherichia coli | Human colon bacterium | 4,639 Kb |
| Helicobacter pylori | Stomach ulcer bacterium | 1,667 Kb |
| Mycobacterium tuberculosis | Tuberculosis | 4,411 Kb |
| Yersinia pestis | Plague | 4,653 Kb |
| Archaea | ||
| Halobacterium | Salt-tolerant archaean | 2,014 Kb |
| Methanobacterium thermoautotrophicum | Methane-producing archaean | 1,751 Kb |
Genome analysis
After an organism has been selected, genome projects involve three components: the sequencing of DNA, the assembly of that sequence to create a representation of the original chromosome, and the annotation and analysis of that representation.[3]
Sequencing
Historically, sequencing was done in sequencing centers, centralized facilities (ranging from large independent institutions such as Joint Genome Institute which sequence dozens of terabases a year, to local molecular biology core facilities) which contain research laboratories with the costly instrumentation and technical support necessary. As sequencing technology continues to improve, however, a new generation of effective fast turnaround benchtop sequencers has come within reach of the average academic laboratory.[41][42] On the whole, genome sequencing approaches fall into two broad categories, shotgun and high-throughput (aka next-generation) sequencing.[3]
Shotgun sequencing

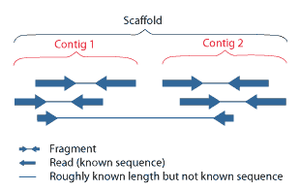
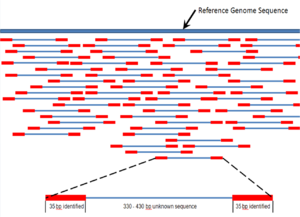
Shotgun sequencing (Sanger sequencing is used interchangably) is a sequencing method designed for analysis of DNA sequences longer than 1000 base pairs, up to and including entire chromosomes.[43] It is named by analogy with the rapidly-expanding, quasi-random firing pattern of a shotgun. Since the chain termination method of DNA sequencing can only be used for fairly short strands (100 to 1000 basepairs), longer DNA sequences must be broken into random small segments which are then sequenced to obtain reads. Multiple overlapping reads for the target DNA are obtained by performing several rounds of this fragmentation and sequencing. Computer programs then use the overlapping ends of different reads to assemble them into a continuous sequence.[43][44]
The technology underlying shotgun sequencing is the classical chain-termination method, which is based on the selective incorporation of chain-terminating
High-throughput sequencing
The high demand for low-cost sequencing has driven the development of high-throughput sequencing (or next-generation sequencing [NGS]) technologies that parallelize the sequencing process, producing thousands or millions of sequences at once.[48][49] High-throughput sequencing technologies are intended to lower the cost of DNA sequencing beyond what is possible with standard dye-terminator methods. In ultra-high-throughput sequencing as many as 500,000 sequencing-by-synthesis operations may be run in parallel.[50][51]
While NGS methods' high throughput are associated with the sequencing of large eukaryotic genomes, their scalability gives them applications in the sequencing of smaller prokaryotic genomes.
454 pyrosequencing
Illumina (Solexa) sequencing

Decoupling the enzymatic reaction and the image capture allows for optimal throughput and theoretically unlimited sequencing capacity. With an optimal configuration, the ultimately reachable instrument throughput is thus dictated solely by the analogic-to-digital conversion rate of the camera, multiplied by the number of cameras and divided by the number of pixels per DNA colony required for visualizing them optimally (approximately 10 pixels/colony). In 2012, with cameras operating at more than 10 MHz A/D conversion rates and available optics, fluidics and enzymatics, throughput can be multiples of 1 million nucleotides/second, corresponding roughly to 1 human genome equivalent at 1x coverage per hour per instrument, and 1 human genome re-sequenced (at approx. 30x) per day per instrument (equipped with a single camera). The camera takes images of the
Single cell genomics
Assembly
When are genomes finished?
- genome standards [58]
- Coverage?
Challenges
Algorithms
Scaffolding
De-novo vs. mapping assembly
Finishing
Annotation
The DNA sequence alone is of little value without additional analysis.
- identifying portions of the genome that do not code for proteins
- identifying elements on the genome, a process called gene prediction, and
- attaching biological information to these elements.
Automatic annotation tools try to perform these steps in silico, as opposed to manual annotation (a.k.a. curation) which involves human expertise and potential experimental verification.[64] Ideally, these approaches co-exist and complement each other in the same annotation pipeline (also see below).
Traditionally, the basic level of annotation is using
Sequencing pipelines and databases
The need for reproducibility and efficient management of large amount of data associated with genome projects mean that computational pipelines have important applications in genomics.[66]
Research areas
Functional genomics
Functional genomics is a field of
A major branch of genomics is still concerned with sequencing the genomes of various organisms, but the knowledge of full genomes has created the possibility for the field of functional genomics, mainly concerned with patterns of gene expression during various conditions. The most important tools here are microarrays and bioinformatics.
Evolutionary genomics
Structural genomics
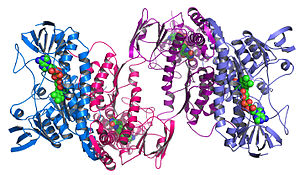
Structural genomics seeks to describe the
Epigenomics
Epigenomics is the study of the complete set of
Metagenomics
Study systems
Viruses and bacteriophages
Microbes
Cyanobacteria
At present there are 24
Animals
Human genomics
Applications of genomics
Genomics has provided applications in many fields, including
Genomic medicine
Synthetic biology and bioengineering
Social aspects
'Astrological' genomics
Race and genomics
'Omics
See also
- Computational genomics
- Epigenomics
- Whole genome sequencing
- Functional genomics
- Genomics of domestication
- Immunomics
- Metagenomics
- Personal genomics
- Proteomics
- Psychogenomics
References
- ^ National Human Genome Research Institute (2010-11-08). "A Brief Guide to Genomics". Genome.gov. Retrieved 2011-12-03.
- ISBN 9780321724120.
- ^ ISBN 9780470085851.
- ^ National Human Genome Research Institute (2010-11-08). "FAQ About Genetic and Genomic Science". Genome.gov. Retrieved 2011-12-03.
- ^ "Genome, n.". Oxford English Dictionary (Third ed.). Oxford University Press. 2008. Retrieved 2012-12-01.(subscription required)
- PMID 18166670.
- PMID 12798815. Retrieved 2012-06-18.
- PMID 14299636.)
{{cite journal}}: Check date values in:|date=(help)CS1 maint: multiple names: authors list (link - S2CID 40989800.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - PMID 5330357.)
{{cite journal}}: CS1 maint: date and year (link) CS1 maint: multiple names: authors list (link - S2CID 4153893.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - S2CID 4289674.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - )
- )
- ^ Sanger, F. (1980), Nobel lecture: Determination of nucleotide sequences in DNA (PDF), Nobelprize.org, retrieved 2010-10-18
- ^ S2CID 4206886.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - PMID 14651855. Retrieved 2012-12-20.
- PMID 271968.
- PMID 265521.
- ^ a b Darden, Lindley (2010). "Molecular Biology". In Edward N. Zalta (ed.) (ed.). The Stanford Encyclopedia of Philosophy (Fall 2010 ed.). Retrieved 2012-12-20.
{{cite book}}:|editor=has generic name (help); Unknown parameter|coauthors=ignored (|author=suggested) (help) - S2CID 4355527.(subscription required)
- PMID 16453699.
- S2CID 4311952.
- S2CID 4271784.
- PMID 7542800.)
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link - S2CID 16763139.(subscription required)
- ^ "Complete genomes: Viruses". NCBI. 2011-11-17. Retrieved 2011-11-18.
- ^ "Genome Project Statistics". Entrez Genome Project. 2011-10-07. Retrieved 2011-11-18.
- ISSN 0362-4331. Retrieved 2012-12-21.}
- PMID 20033048.
- ^ "Human gene number slashed". BBC. 2004-10-20. Retrieved 2012-12-21.
- S2CID 21797344.
- ^ National Human Genome Research Institute (2004-07-14). "Dog Genome Assembled: Canine Genome Now Available to Research Community Worldwide". Genome.gov. Retrieved 2012-01-20.
- ^ ISBN 9780465043330.
- PMID 23128226.
- S2CID 5889828.
- ^ ISBN 9780226172958-0226172953.).
{{cite book}}: Check|isbn=value: length (help); Unknown parameter|coauthors=ignored (|author=suggested) (help) Cite error: The named reference "barnes2008" was defined multiple times with different content (see the help page - ISSN 1099-274X.
- ISBN 0028656067.)
{{cite encyclopedia}}:|editor=has generic name (help); Unknown parameter|coauthors=ignored (|author=suggested) (help - ISBN 9781429233248.
- ^ a b Monya Baker (2012-09-14). "Benchtop sequencers ship off" (Blog). Nature News Blog. Retrieved 2012-12-22.
- PMID 22827831.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - ^ )
- PMID 6269069.
- PMID 1100841.)
{{cite journal}}: CS1 maint: date and year (link - ^ PMID 23251337.
- ^ a b Illumina, Inc. (2012-02-28). An Introduction to Next-Generation Sequencing Technology (PDF). San Diego, California, USA: Illumina, Inc. p. 12. Retrieved 2012-12-28.
- S2CID 25688677.)
{{cite journal}}: CS1 maint: date and year (link - PMID 16468433.)
{{cite journal}}: CS1 maint: date and year (link - PMID 18832462.
- PMID 19679224.
- ^ Kawashima, Eric H. (2005-05-12), Method of nucleic acid amplification, retrieved 2012-12-22
{{citation}}: Unknown parameter|coauthors=ignored (|author=suggested) (help) - PMID 18576944.
- ^ Gilbert, David (2009-04-29). "DNA of Uncultured Organisms Sequenced Using Novel Single-Cell Approach" (Press Release). DOE Joint Genome Institute. Retrieved 2012-12-23.
- PMID 19390573.
- PMID 1631067.
- ^ Stein, Richard A. (2009-06-01). "Single-Cell Genomics Clarifies Big Picture". Genetic Engineering & Biotechnology News. Vol. 29, no. 11. Retrieved 2012-12-23.
- PMID 19815760.
- PMID 18262676.
- ISBN 0470849746, 9780470849743, 047001153X, 9780470011539. Retrieved 2012-12-23.)
{{cite book}}:|editor=has generic name (help); Check|isbn=value: invalid character (help)CS1 maint: multiple names: editors list (link - PMID 21559467.
- S2CID 4648109.
- S2CID 12044602.
- S2CID 20412451.
- PMID 23203987.
- PMID 18720577.
- PMID 17349043.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - PMID 10739263.
- S2CID 5656447. Retrieved 2012-12-07.
- ^ ISBN 9780393070057.
- S2CID 6780101.
- ^
Hugenholtz, Philip; Goebel, Brett M.; Pace, Norman R. (1 September 1998). "Impact of Culture-Independent Studies on the Emerging Phylogenetic View of Bacterial Diversity". J. Bacteriol. 180 (18): 4765–74. PMID 9733676.
- ^
Eisen, JA (2007). "Environmental Shotgun Sequencing: Its Potential and Challenges for Studying the Hidden World of Microbes". PLOS Biology. 5 (3): e82. PMID 17355177.)
{{cite journal}}: CS1 maint: unflagged free DOI (link - ^
Marco, D, ed. (2010). Metagenomics: Theory, Methods and Applications. Caister Academic Press. ISBN 978-1-904455-54-7.
- ^
Marco, D, ed. (2011). Metagenomics: Current Innovations and Future Trends. ISBN 978-1-904455-87-5.
- PMID 12794192.
- ISBN 978-1-904455-14-1.
- S2CID 13318164.
- ^ Office of Science (2005-08). Genomics:GTL Roadmap (PDF). US Department of Energy.
{{cite conference}}: Check date values in:|date=(help) - ^ Microbiology in the 21st Century: Where Are We and Where Are We Going?. American Academy of Microbiology. 2004.
- ISBN 978-1-904455-15-8.
- ProQuest 220135424. Retrieved 2012-12-07.
- ProQuest 923268241. Retrieved 2012-12-07.
- ProQuest 914354372.)
{{cite journal}}: CS1 maint: numeric names: authors list (link - ProQuest 890081959. Retrieved 2012-12-07.
- ProQuest 907232641. Retrieved 2012-12-07.
- ISBN 9780465021758-0465021751.)
{{cite book}}: Check|isbn=value: length (help); Unknown parameter|coauthors=ignored (|author=suggested) (help - PMID 22265990.
- ISSN 0268-540X.
- )
- ProQuest 220163870. Retrieved 2012-12-07.
External links
- Annual Review of Genomics and Human Genetics
- BMC Genomics: A BMC journal on Genomics
- Genomics journal
- NHGRI: US government's genome institute
- JCVI Comprehensive Microbial Resource
- KoreaGenome.org: The first Korean Genome published and the sequence is available freely.
- GenomicsNetwork: Looks at the development and use of the science and technologies of genomics.
- Institute for Genome Sciences: Genomics research.
- MIT OpenCourseWare HST.512 Genomic Medicine A free, self-study course in genomic medicine. Resources include audio lectures and selected lecture notes.