Human Y-chromosome DNA haplogroup

| Part of a series on |
| Genetic genealogy |
|---|
| Concepts |
| Related topics |
|
In human genetics, a human Y-chromosome DNA haplogroup is a haplogroup defined by mutations in the non-recombining portions of DNA from the male-specific Y chromosome (called Y-DNA). Many people within a haplogroup share similar numbers of short tandem repeats (STRs) and types of mutations called single-nucleotide polymorphisms (SNPs).[2]
The human Y-chromosome accumulates roughly two mutations per generation.[3] Y-DNA haplogroups represent major branches of the Y-chromosome phylogenetic tree that share hundreds or even thousands of mutations unique to each haplogroup.
The Y-chromosomal most recent common ancestor (Y-MRCA, informally known as
Naming convention
This section has multiple issues. Please help improve it or discuss these issues on the talk page. (Learn how and when to remove these template messages)
|
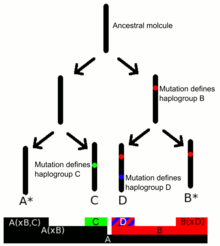
Y-DNA haplogroups are defined by the presence of a series of Y-DNA SNP markers. Subclades are defined by a terminal SNP, the SNP furthest down in the Y-chromosome phylogenetic tree.[7][8] The Y Chromosome Consortium (YCC) developed a system of naming major Y-DNA haplogroups with the capital letters A through T, with further subclades named using numbers and lower case letters (YCC longhand nomenclature). YCC shorthand nomenclature names Y-DNA haplogroups and their subclades with the first letter of the major Y-DNA haplogroup followed by a dash and the name of the defining terminal SNP.[9]
Y-DNA haplogroup nomenclature is changing over time to accommodate the increasing number of SNPs being discovered and tested, and the resulting expansion of the Y-chromosome phylogenetic tree. This change in nomenclature has resulted in inconsistent nomenclature being used in different sources.[2] This inconsistency, and increasingly cumbersome longhand nomenclature, has prompted a move toward using the simpler shorthand nomenclature.
Phylogenetic structure
- Phylogenetic tree of Y-DNA haplogroups [10]
Major Y-DNA haplogroups
Haplogroups A and B
Haplogroup A is the NRY (
- Haplogroup A
- Haplogroup A00
- Haplogroup A0(formerly also A1b)
- Haplogroup A1(also A1a-T)
- Haplogroup A1a (M31)
- Haplogroup A1b (also A2-T; P108, V221)
- Haplogroup A1b1a1 (also A2; M14)
- Haplogroup A1b1b (also A3; M32)
- Haplogroup BT(M91, M42, M94, M139, M299)
- Haplogroup B(M60)
- Haplogroup CT
Haplogroup CT (P143)
The defining mutations separating CT (all haplogroups except for A and B) are M168 and M294. The site of origin is likely in Africa. Its age has been estimated at approximately 88,000 years old,[11][12] and more recently at around 100,000[13] or 101,000 years old.[14]
Haplogroup C (M130)
-
- Haplogroup C1(F3393/Z1426)
- Haplogroup C1a (CTS11043)
- Haplogroup C1a1 (M8, M105, M131) Found with low frequency in Japan
- Haplogroup C1a2 (V20) Found with low frequency in Europe, Armenians, Algeria, and Nepal
- Haplogroup C1b (F1370, Z16480)
- Haplogroup C1b1 (AM00694/K281)
- Haplogroup C1b1a (B66/Z16458)
- China
- Haplogroup C1b1a2 (B65)
- Haplogroup C1b1a2a (B67) Found among Lebbo' people in Borneo, Indonesia
- Haplogroup C1b1a2b (F725) Found among Han Chinese (Guangdong, Hunan, and Shaanxi), Dai people (Yunnan), Murut people (Brunei), Malay people (Singapore), and Aeta people (Philippines)
- Haplogroup C1b1a3 (Z16582) Found with low frequency in Saudi Arabia and Iraq
- Haplogroup C1b1b (B68) Found among Dusun people (Brunei)
- Haplogroup C1b1a (B66/Z16458)
- Haplogroup C1b2 (C-Z16582)
- Haplogroup C1b3 (B477/Z31885)
- Haplogroup C1b3a (M38) Found in Indonesia, New Guinea, Melanesia, Micronesia, and Polynesia
- Haplogroup C1b3b (M347, P309) Found among the indigenous peoples in Australia
- Haplogroup C1b1 (AM00694/K281)
- Haplogroup C1a (CTS11043)
- Na-Dené-speaking peoples
Haplogroup D (CTS3946)
- Haplogroup D(CTS3946)
- China (especially Tibet), the Andaman Islands
- Haplogroup D1a (CTS11577)
- Haplogroup D1a1 (Z27276, Z27283, Z29263)
- Haplogroup D1a1a (M15) Found mainly in Hmong-Mienpeoples
- Haplogroup D1a1b (P99) Found mainly in Naxi, and Turkic peoples
- Haplogroup D1a1a (M15) Found mainly in
- Haplogroup D1a2 (M55, M57, M64.1, M179, P12, P37.1, P41.1 (M359.1), 12f2.2) Found mainly in Japan
- Onge, Jarawa)
- Haplogroup D1a1 (Z27276, Z27283, Z29263)
- Haplogroup D1b (L1366, L1378, M226.2) Found in Mactan Island, Philippines
- Haplogroup D1a (CTS11577)
- Haplogroup D2 (A5580.2) Found in Nigeria, Saudi Arabia and Syria
Haplogroup E (M96)
- Haplogroup E (M40, M96) Found in Africa and parts of the Middle Eastand Europe
- Haplogroup E1(P147)
- Haplogroup E1a(M33, M132) formerly E1
- Haplogroup E1b(P177)
- Haplogroup E1b1(P2, DYS391p); formerly E3
- Haplogroup E1b1a(V38)
- Haplogroup E1b1a1 (M2) Found in Africa, especially among Niger–Congo-speaking populations.; formerly E3a
- -speaking populations.; formerly E3*
- Haplogroup E1b1b(M215)
- Haplogroup E1b1b1 (M35) Found in Horn of Africa, North Africa, the Middle East, and Europe (especially in areas near the Mediterranean and the Balkans); formerly E3b
- Haplogroup E2(M75)
Haplogroup F (M89)
The groups descending from haplogroup F are found in some 90% of the world's population, but almost exclusively outside of sub-Saharan Africa.
F xG,H,I,J,K is rare in modern populations and peaks in
Haplogroup G (M201)
It is found in many ethnic groups in Eurasia; most common in the
G-M201 is also found in small numbers in northwestern China and India, Bangladesh, Pakistan, Sri Lanka, Malaysia, and North Africa.
- Haplogroup G1
- Haplogroup G2
- Haplogroup G2a
- Haplogroup G2a1
- Haplogroup G2a2
- Haplogroup G2a3
- Haplogroup G2a3a
- Haplogroup G2a3b
- Haplogroup G2a3b1
- Haplogroup G2b
- Haplogroup G2c(formerly Haplogroup G5)
- Haplogroup G2a
Haplogroup H (M69)
Haplogroup H (M69) probably emerged in Southern Central Asia, South Asia or West Asia, about 48,000 years BP, and remains largely prevalent there in the forms of H1 (M69) and H3 (Z5857). Its sub-clades are also found in lower frequencies in Iran, Central Asia, across the middle-east, and the Arabian peninsula.
However, H2 (P96) is present in Europe since the Neolithic and H1a1 (M82) spread westward in the
This section needs expansion. You can help by adding to it. (September 2016) |
Haplogroup I (M170)
Haplogroup I (M170, M258) is found mainly in Europe and the Caucasus.
- Haplogroup I1Nordid/Nordic Europids (M253) Found mainly in northern Europe
- Haplogroup I2Dinarid/Dinaric Europids (P215) Found mainly in Balkans, southeast Europe and Sardinia save for I2B1 (m223) which is found at a moderate frequency in Western, Central, and Northern Europe.
Haplogroup J (M304)
- Haplogroup J* (J-M304*) is rare outside the island of Socotra.
- Haplogroup J1 Semitid/Bedouinid Arabids (M267) are associated with Northeast Caucasian peoples in Dagestan and Semitic languages speaking people in the Middle East, Ethiopia, and North Africaand also found in Mediterranean Europe in smaller frequencies much like haplogroup T.
- Haplogroup J2 Syrid/Nahrainid Arabids (M172) is found mainly in the Semitic-speaking peoples, Anatolia, Greece, the Balkans, Italy, Iran, the Caucasus, South Asia, and Central Asia.
Haplogroup K (M9)
Haplogroup K (M9) is spread all over Eurasia, Oceania and among Native Americans.
K(xLT,K2a,K2b) – that is, K*, K2c, K2d or K2e – is found mainly in
Haplogroups L and T (K1)
Haplogroup L (M20) is found in South Asia, Central Asia, South-West Asia, and the Mediterranean.
Haplogroup K2 (K-M526)
The only living males reported to carry the basal
This section needs expansion. You can help by adding to it. (September 2016) |
Haplogroups K2a, K2a1, NO & NO1
This section needs expansion. You can help by adding to it. (September 2016) |
Haplogroup N
Haplogroup N possibly originated in eastern Asia and spread both northward and westward into Siberia, being the most common group found in some Uralic-speaking peoples.
Haplogroup O
Haplogroup O (M175) is found with its highest frequency in East Asia and Southeast Asia, with lower frequencies in the South Pacific, Central Asia, South Asia, and islands in the Indian Ocean (e.g. Madagascar, the Comoros).
- Haplogroup O1 (F265/M1354, CTS2866, F75/M1297, F429/M1415, F465/M1422)
- Tai–Kadaipeoples
- Haplogroup O1b(P31, M268)
- Tai–Kadai-speaking peoples, Malays, and Indonesians
- Haplogroup O1b2 (SRY465, M176) Found in Japan, Korea, Manchuria, and Southeast Asia
- Austronesia including Polynesia
Haplogroups K2b1, M & S
No examples of the basal
Its primary subclades are two major haplogroups:
- Haplogroup S (B254) also known as K2b1a: found in the highlands of Papua New Guineaand;
- Haplogroup M (P256) also known as K2b1b: found in New Guineaand Melanesia.
This section needs expansion. You can help by adding to it. (September 2016) |
Haplogroup P (K2b2)
Haplogroup P (P295) has two primary branches: P1 (P-M45) and the extremely rare P2 (P-B253).[21]
P*, P1* and P2 are found together only on the island of
P1 is also the parent node of two primary clades:
- Haplogroup Q(Q-M242) and;
- Haplogroup R (R-M207). These share the common marker M45 in addition to at least 18 other SNPs.
Haplogroup Q (MEH2, M242, P36) found in Siberia and the Americas Haplogroup R (M207, M306): found in Europe, West Asia, Central Asia, and South Asia
Haplogroup Q M242
Q is defined by the SNP M242. It is believed to have arisen in Central Asia approximately 32,000 years ago.[24][25] The subclades of Haplogroup Q with their defining mutation(s), according to the 2008 ISOGG tree[26] are provided below. ss4 bp, rs41352448, is not represented in the ISOGG 2008 tree because it is a value for an STR. This low frequency value has been found as a novel Q lineage (Q5) in Indian populations[27]
The 2008 ISOGG tree
- Q (M242)
- Q*
- Q1 (P36.2)
- Q1*
- Q1a (MEH2)
- Q1a*
- Q1a1 (M120, M265/N14) Found with low frequency among
- Q1a2 (M25, M143) Found at low to moderate frequency among some populations of Southwest Asia, Central Asia, and Siberia
- Q1a3 (M346)
- Q1a3* Found at low frequency in Pakistan, India, and Tibet
- Q1a3a (M3) Typical of indigenous peoples of the Americas
- Q1a3a*
- Q1a3a1 (M19) Found among some indigenous peoples of
- Q1a3a2 (M194)
- Q1a3a3 (M199, P106, P292)
- Q1a4 (P48)
- Q1a5 (P89)
- Q1a6 (M323) Found in a significant minority of Yemeni Jews
- Q1b (M378) Found at low frequency among samples of Sindhis
Haplogroup R (M207)

Haplogroup R is defined by the SNP M207. The bulk of Haplogroup R is represented in the descendant subclade R1 (M173), which originated on the Siberia. R1 has two descendant subclades: R1a and R1b.
R1a is associated with the proto-Indo-Iranian and Balto-Slavic peoples, and is now found primarily in Central Asia, South Asia, and Eastern Europe.
- Haplogroup R1(M173) Found throughout western Eurasia
- Haplogroup R1a(M420) Found in Central Asia, South Asia, and Central, Northern and Eastern Europe, Balkans
- Haplogroup R1b (M343) Found in Western Europe, West Asia, Central Asia, North Africa, and northern Cameroon
- Haplogroup R2(M124) Found in South Asia, Caucasus, Central Asia, and Eastern Europe
Chronological development of haplogroups
| Haplogroup | Possible time of origin | Possible place of origin | Possible |
|---|---|---|---|
| A00 | 235,900[5] or 275,000 years ago[34] | Africa[35] | 235,900 years ago |
BT |
130,700 years ago[5] | Africa | 88,000 years ago |
CT |
88,000[5] or 101-100,000 years ago[13][14] | Africa | 68,500 years ago |
E |
65,200,[5] 69,000,[36] or 73,000 years ago[37] | East Africa[38] or Asia[15] | 53,100 years ago |
F |
65,900 years ago[5] | Eurasia | 48,800 years ago |
G |
48,500 years ago[5] | Middle East | 26,200 years ago |
IJ |
47,200 years ago[5] | Middle East | 42,900 years ago |
K |
47,200 years ago[5] | Asia | 45,400 years ago |
| P | 45,400 years ago[5] | Asia | 31,900 years ago |
| J | 42,900 years ago[5][39] | Middle East | 31,600 years ago |
| I | 42,900 years ago[5] | Europe | 27,500 years ago |
E-M215 (E1b1b) |
42,300 years ago[5][40] | East Africa | 34,800 years ago |
E-V38 (E1b1a) |
42,300 years ago[5][40] | East Africa | 40,100 years ago |
| N | 36,800 years ago[5][41] | Asia | 22,100 years ago |
| E1b1b-M35 | 34,800 years ago[5][40] | East Africa | 24,100 years ago |
| R | 31,900 years ago[5] | Asia | 28,200 years ago |
| J-M267 (J1) | 31,600 years ago[5][39] | Middle East | 18,500 years ago |
| J-M172 (J2) | 31,600 years ago[5][39] | Middle East | 27,800 years ago[5][42] |
| R-M173 (R1) | 28,200 years ago[5] | Asia | 22,800 years ago |
| I-M253 (I1) | 27,500 years ago[5][43][44] | Europe | 4,600 years ago |
| I-M438 (I2) | 27,500 years ago[5][44] | Europe | 21,800 years ago |
| R-M420 (R1a) | 22,800 years ago[5][45] | Eurasia | 18,300 years ago |
| R-M343 (R1b) | 22,800 years ago[5][46] | Eurasia[47] | 20,400 years ago |
| I2-L460 (I2a) | 21,800 years ago[5][48] | Europe | 21,100 years ago |
| I2a-P37 | 21,100 years ago[5][43][49] | Europe | 18,500 years ago |
| E1b1b-M78 | 19,800 years ago[5][40][50] | Northeast Africa[50] | 13,400 years ago[5][50] |
| I2a-M423 | 18,500 years ago[5][49] | Europe | 13,500 years ago |
| I2a-M223 | 17,400 years ago[5] | Europe | 12,100 years ago |
| N1c-M178 | 14,200 years ago[5][41] | Asia | 11,900 years ago |
| R1a-M17 | 14,100 years ago[5][45][51] | Eastern Europe | 8,500 years ago |
| R1b-M269 | 13,300 years ago[5] | Eastern Europe | 6,400 years ago[52] |
| E1b1b-V12 | 11,800 years ago[5][50] | North Africa | 9,900 years ago |
| E-U175 (E1b1a8) | 9,200 years ago[5][40] | East Africa | 8,500 years ago |
| E1b1b-V13 | 8,100 years ago[5][50] | Southern Europe | 4,800 years ago |
| E-M191 (E1b1a7) | 7,400 years ago[5][40] | East Africa | 6,400 years ago |
| E-U174 (E1b1a-U174) | 6,400 years ago[5][40] | East Africa | 5,300 years ago |
| R1b-L151 | 5,800 years ago[5] | Eastern Europe | 4,800 years ago |
| R1a-Z280 | 5,000 years ago[5] | Eastern Europe | 4,600 years ago[53] |
| R1a-M458 | 4,700 years ago[5] | Eastern Europe | 4,700 years ago[53] |
See also
- List of Y-chromosome haplogroups in populations of the world
- Y-DNA haplogroups in populations of Europe
- Genetic history of Europe
- List of Y-DNA single-nucleotide polymorphisms
- List of Y-STR markers
- Human mitochondrial DNA haplogroup
- * (haplogroup)
- Molecular phylogeny
- Genetic genealogy
- Genealogical DNA test
- Conversion table for Y chromosome haplogroups
References
- PMID 32666166.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - ^ a b "Understanding Haplogroups: How are the haplogroups named?". Family Tree DNA. Archived from the original on 21 June 2012. Retrieved 31 March 2013.
- . Retrieved 18 September 2017. "one mutation in every 30 million base pairs"
- PMID 25770088. "we date the Y-chromosomal most recent common ancestor (MRCA) in Africa at 254 (95% CI 192–307) kya and detect a cluster of major non-African founder haplogroups in a narrow time interval at 47–52 kya, consistent with a rapid initial colonization model of Eurasia and Oceania after the out-of-Africa bottleneck. In contrast to demographic reconstructions based on mtDNA, we infer a second strong bottleneck in Y-chromosome lineages dating to the last 10 ky. We hypothesize that this bottleneck is caused by cultural changes affecting variance of reproductive success among males."
- ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai aj ak al am an ao ap "YFull YTree". YFull. Retrieved 15 September 2017.
- ^ "Something Weird Happened to Men 7,000 Years Ago, And We Finally Know Why". 31 May 2018.
Around 7000 years ago - all the way back in the Neolithic - something really peculiar happened to human genetic diversity. Over the next 2,000 years, and seen across Africa, Europe and Asia, the genetic diversity of the Y chromosome collapsed, becoming as though there was only one man for every 17 women.
- ^ "Understanding Results: Y-DNA Single Nucleotide Polymorphism (SNP): What is a Y-chromosome DNA (Y-DNA) haplogroup?". Family Tree DNA. Retrieved 31 March 2013.
Y-chromosome DNA (Y-DNA) haplogroups are the major branches on the human paternal family tree. Each haplogroup has many subbranches. These are subclades.
- ^ "myFTDNA 2.0 User Guide: Y-DNA: What is the Y-DNA – Matches page?". Family Tree DNA. Retrieved 31 March 2013.
A terminal SNP determines the terminal (final) subbranch on the Y-DNA Tree to which someone belongs.
- ^ "Understanding Results: Y-DNA Single Nucleotide Polymorphism (SNP): How are haplogroups and their subclades named?". Family Tree DNA. Retrieved 31 March 2013.
- ^ a b Copyright 2015 ISOGG. "ISOGG 2015 Y-DNA Haplogroup Tree Trunk". isogg.org.
{{cite web}}: CS1 maint: numeric names: authors list (link) - PMID 18076332.
- ^ PMID 18385274.
- ^ PMID 25770088.
- ^ PMID 31196864.
- ^ PMID 19920170.
- PMID 25078354.
- ^ This was, for instance, the case with the original subclade F3 (M96), which has since been renamed Haplogroup H2.
- PMID 12173029.
- PMID 16724001.
- PMID 19918998.
- ^ a b ISOGG, 2016, Y-DNA Haplogroup P and its Subclades – 2016 (20 June 2016).
- ^ PMID 25078354.
- PMID 24896152.
- PMID 18313026. Archived from the original(PDF) on 2009-03-25. Retrieved 2013-05-22.
Since the first studies, it has been found that extant Native American populations exhibit almost exclusively five "mtDNA haplogroups" (A–D and X)6 classified in the autochthonous haplogroups A2, B2, C1, D1, and X2a.7 Haplogroups A–D are found all over the New World and are frequent in Asia, supporting a northeastern Asian origin of these lineages
- PMID 14595095.
- ^ "Y-DNA Haplogroup Tree 2010". International Society of Genetic Genealogy. Retrieved 1 July 2010.
- PMID 18021436.
- PMID 25468874.
- PMID 24204668.
- S2CID 4301581.
- PMID 11526236.
- PMID 12900798.
- PMID 25988751.
- PMID 27058445.
- ^ "The father of all men is 340,000 years old".
- PMID 25770088.
- PMID 31196864.
- PMID 15069642.
- ^ PMID 15069642.
- ^ PMID 26108492.
- ^ PMID 23840409.
- PMID 25988751.
- ^ PMID 15162323. Archived from the original(PDF) on 2009-06-19. Retrieved 2016-05-04.
- ^ a b P.A. Underhill, N.M. Myres, S. Rootsi, C.T. Chow, A.A. Lin, R.P. Otillar, R. King, L.A. Zhivotovsky, O. Balanovsky, A. Pshenichnov, K.H. Ritchie, L.L. Cavalli-Sforza, T. Kivisild, R. Villems, S.R. Woodward, New Phylogenetic Relationships for Y-chromosome Haplogroup I: Reappraising its Phylogeography and Prehistory, in P. Mellars, K. Boyle, O. Bar-Yosef and C. Stringer (eds.), Rethinking the Human Evolution (2007), pp. 33–42.
- ^ PMID 19158816.
- ^ ftDNA
- ^ Myres2010
- PMID 26567969.
- ^ a b "Mesolithic Western Eurasian DNA". Ancestral Journeys. Archived from the original on 2016-04-07. Retrieved 2016-05-04.
- ^
- ^ Evatt, Danny (2013-11-01). "The Evatt Clan: A Worldwide Historical Review of the Evatt Family Surname".
- PMID 21720564.
- ^ PMID 19888303.
- ^ 2005 Y-chromosome Phylogenetic Tree, from FamilyTreeDNA.com
- ^ A Nomenclature system for the Tree of Human Y-Chromosomal Haplogroups, Genome.org
Further reading
- Mendez, Fernando; Krahn, Thomas; Schrack, Bonnie; Krahn, Astrid-Maria; Veeramah, Krishna; Woerner, August; Fomine, Forka Leypey Mathew; Bradman, Neil; Thomas, Mark; Karafet, Tatiana M.; Hammer, Michael F. (7 March 2013). "An African American paternal lineage adds an extremely ancient root to the human Y chromosome phylogenetic tree" (PDF). PMID 23453668. Archived from the original(PDF) on 24 September 2019. Retrieved 31 March 2013.
- "Y-Haplogroup A Phylogenetic Tree". March 2013. Archived from the original on 18 August 2018. Retrieved 30 March 2013. (chart highlighting new branches added to the A phylotree in March 2013)
External links
- ISOGG Y-DNA Haplogroup Tree
- Family Tree DNA Public Haplotree
- Chart of the speed of different Y chromosomal STR mutation rates
- Map of Y Haplogroups
- Atlas of the Human Journey, from the Genographic Project, National Geographic
- DNA Heritage's Y-haplogroup map Archived 2006-02-18 at the Wayback Machine
- Video tutorial on Discovering Paternal Ancestry with Y-Chromosomes
- Haplogroup Predictor
- Semino O, Passarino G, Oefner PJ, et al. (November 2000). "The genetic legacy of Paleolithic Homo sapiens sapiens in extant Europeans: a Y chromosome perspective". Science. 290 (5494): 1155–59. Paper that defined "Eu" haplogroups
- Y-DNA Haplogroup and Sub-clade Projects
- Kerchner's YDNA Haplogroup Descriptions, Projects & Links
- Y-DNA Testing Company STR Marker Comparison Chart
- Y-DNA Ethnographic and Genographic Atlas and Open-Source Data Compilation
- Y Chromosome Consortium
